Paddy J. Slator
MRI Parameter Mapping via Gaussian Mixture VAE: Breaking the Assumption of Independent Pixels
Nov 16, 2024



Abstract:We introduce and demonstrate a new paradigm for quantitative parameter mapping in MRI. Parameter mapping techniques, such as diffusion MRI and quantitative MRI, have the potential to robustly and repeatably measure biologically-relevant tissue maps that strongly relate to underlying microstructure. Quantitative maps are calculated by fitting a model to multiple images, e.g. with least-squares or machine learning. However, the overwhelming majority of model fitting techniques assume that each voxel is independent, ignoring any co-dependencies in the data. This makes model fitting sensitive to voxelwise measurement noise, hampering reliability and repeatability. We propose a self-supervised deep variational approach that breaks the assumption of independent pixels, leveraging redundancies in the data to effectively perform data-driven regularisation of quantitative maps. We demonstrate that our approach outperforms current model fitting techniques in dMRI simulations and real data. Especially with a Gaussian mixture prior, our model enables sharper quantitative maps, revealing finer anatomical details that are not presented in the baselines. Our approach can hence support the clinical adoption of parameter mapping methods such as dMRI and qMRI.
ssVERDICT: Self-Supervised VERDICT-MRI for Enhanced Prostate Tumour Characterisation
Sep 27, 2023
Abstract:Purpose: Demonstrating and assessing self-supervised machine learning fitting of the VERDICT (Vascular, Extracellular and Restricted DIffusion for Cytometry in Tumours) model for prostate. Methods: We derive a self-supervised neural network for fitting VERDICT (ssVERDICT) that estimates parameter maps without training data. We compare the performance of ssVERDICT to two established baseline methods for fitting diffusion MRI models: conventional nonlinear least squares (NLLS) and supervised deep learning. We do this quantitatively on simulated data, by comparing the Pearson's correlation coefficient, mean-squared error (MSE), bias, and variance with respect to the simulated ground truth. We also calculate in vivo parameter maps on a cohort of 20 prostate cancer patients and compare the methods' performance in discriminating benign from cancerous tissue via Wilcoxon's signed-rank test. Results: In simulations, ssVERDICT outperforms the baseline methods (NLLS and supervised DL) in estimating all the parameters from the VERDICT prostate model in terms of Pearson's correlation coefficient, bias, and MSE. In vivo, ssVERDICT shows stronger lesion conspicuity across all parameter maps, and improves discrimination between benign and cancerous tissue over the baseline methods. Conclusion: ssVERDICT significantly outperforms state-of-the-art methods for VERDICT model fitting, and shows for the first time, fitting of a complex three-compartment biophysical model with machine learning without the requirement of explicit training labels.
Fitting a Directional Microstructure Model to Diffusion-Relaxation MRI Data with Self-Supervised Machine Learning
Oct 05, 2022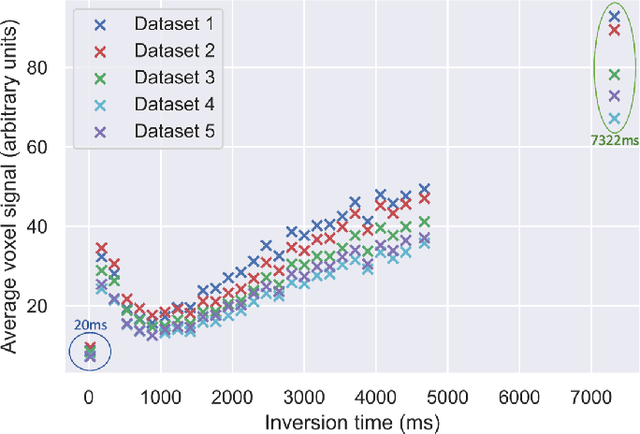
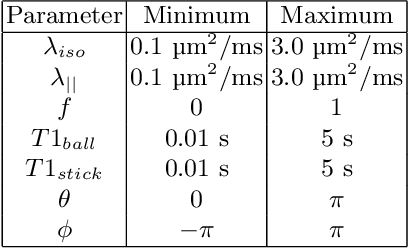
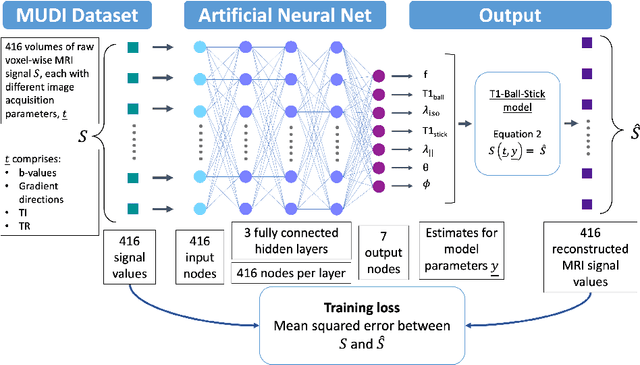
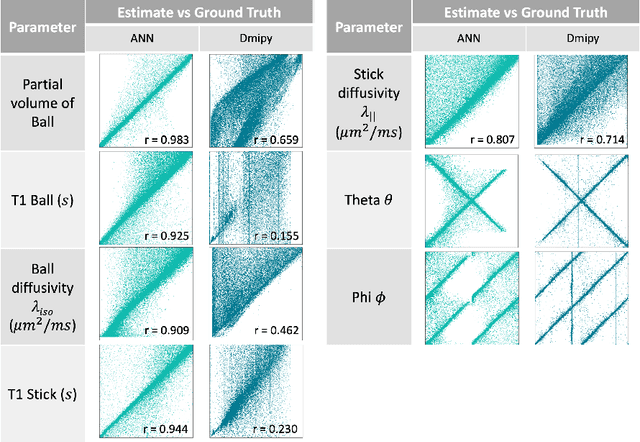
Abstract:Machine learning is a powerful approach for fitting microstructural models to diffusion MRI data. Early machine learning microstructure imaging implementations trained regressors to estimate model parameters in a supervised way, using synthetic training data with known ground truth. However, a drawback of this approach is that the choice of training data impacts fitted parameter values. Self-supervised learning is emerging as an attractive alternative to supervised learning in this context. Thus far, both supervised and self-supervised learning have typically been applied to isotropic models, such as intravoxel incoherent motion (IVIM), as opposed to models where the directionality of anisotropic structures is also estimated. In this paper, we demonstrate self-supervised machine learning model fitting for a directional microstructural model. In particular, we fit a combined T1-ball-stick model to the multidimensional diffusion (MUDI) challenge diffusion-relaxation dataset. Our self-supervised approach shows clear improvements in parameter estimation and computational time, for both simulated and in-vivo brain data, compared to standard non-linear least squares fitting. Code for the artificial neural net constructed for this study is available for public use from the following GitHub repository: https://github.com/jplte/deep-T1-ball-stick
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge