Min Cen
Replay Failures as Successes: Sample-Efficient Reinforcement Learning for Instruction Following
Dec 29, 2025Abstract:Reinforcement Learning (RL) has shown promise for aligning Large Language Models (LLMs) to follow instructions with various constraints. Despite the encouraging results, RL improvement inevitably relies on sampling successful, high-quality responses; however, the initial model often struggles to generate responses that satisfy all constraints due to its limited capabilities, yielding sparse or indistinguishable rewards that impede learning. In this work, we propose Hindsight instruction Replay (HiR), a novel sample-efficient RL framework for complex instruction following tasks, which employs a select-then-rewrite strategy to replay failed attempts as successes based on the constraints that have been satisfied in hindsight. We perform RL on these replayed samples as well as the original ones, theoretically framing the objective as dual-preference learning at both the instruction- and response-level to enable efficient optimization using only a binary reward signal. Extensive experiments demonstrate that the proposed HiR yields promising results across different instruction following tasks, while requiring less computational budget. Our code and dataset is available at https://github.com/sastpg/HIR.
Enhancing WSI-Based Survival Analysis with Report-Auxiliary Self-Distillation
Sep 19, 2025Abstract:Survival analysis based on Whole Slide Images (WSIs) is crucial for evaluating cancer prognosis, as they offer detailed microscopic information essential for predicting patient outcomes. However, traditional WSI-based survival analysis usually faces noisy features and limited data accessibility, hindering their ability to capture critical prognostic features effectively. Although pathology reports provide rich patient-specific information that could assist analysis, their potential to enhance WSI-based survival analysis remains largely unexplored. To this end, this paper proposes a novel Report-auxiliary self-distillation (Rasa) framework for WSI-based survival analysis. First, advanced large language models (LLMs) are utilized to extract fine-grained, WSI-relevant textual descriptions from original noisy pathology reports via a carefully designed task prompt. Next, a self-distillation-based pipeline is designed to filter out irrelevant or redundant WSI features for the student model under the guidance of the teacher model's textual knowledge. Finally, a risk-aware mix-up strategy is incorporated during the training of the student model to enhance both the quantity and diversity of the training data. Extensive experiments carried out on our collected data (CRC) and public data (TCGA-BRCA) demonstrate the superior effectiveness of Rasa against state-of-the-art methods. Our code is available at https://github.com/zhengwang9/Rasa.
GraphCoT-VLA: A 3D Spatial-Aware Reasoning Vision-Language-Action Model for Robotic Manipulation with Ambiguous Instructions
Aug 11, 2025Abstract:Vision-language-action models have emerged as a crucial paradigm in robotic manipulation. However, existing VLA models exhibit notable limitations in handling ambiguous language instructions and unknown environmental states. Furthermore, their perception is largely constrained to static two-dimensional observations, lacking the capability to model three-dimensional interactions between the robot and its environment. To address these challenges, this paper proposes GraphCoT-VLA, an efficient end-to-end model. To enhance the model's ability to interpret ambiguous instructions and improve task planning, we design a structured Chain-of-Thought reasoning module that integrates high-level task understanding and planning, failed task feedback, and low-level imaginative reasoning about future object positions and robot actions. Additionally, we construct a real-time updatable 3D Pose-Object graph, which captures the spatial configuration of robot joints and the topological relationships between objects in 3D space, enabling the model to better understand and manipulate their interactions. We further integrates a dropout hybrid reasoning strategy to achieve efficient control outputs. Experimental results across multiple real-world robotic tasks demonstrate that GraphCoT-VLA significantly outperforms existing methods in terms of task success rate and response speed, exhibiting strong generalization and robustness in open environments and under uncertain instructions.
Constraint Multi-class Positive and Unlabeled Learning for Distantly Supervised Named Entity Recognition
Apr 07, 2025



Abstract:Distantly supervised named entity recognition (DS-NER) has been proposed to exploit the automatically labeled training data by external knowledge bases instead of human annotations. However, it tends to suffer from a high false negative rate due to the inherent incompleteness. To address this issue, we present a novel approach called \textbf{C}onstraint \textbf{M}ulti-class \textbf{P}ositive and \textbf{U}nlabeled Learning (CMPU), which introduces a constraint factor on the risk estimator of multiple positive classes. It suggests that the constraint non-negative risk estimator is more robust against overfitting than previous PU learning methods with limited positive data. Solid theoretical analysis on CMPU is provided to prove the validity of our approach. Extensive experiments on two benchmark datasets that were labeled using diverse external knowledge sources serve to demonstrate the superior performance of CMPU in comparison to existing DS-NER methods.
Dynamic Entity-Masked Graph Diffusion Model for histopathological image Representation Learning
Dec 13, 2024



Abstract:Significant disparities between the features of natural images and those inherent to histopathological images make it challenging to directly apply and transfer pre-trained models from natural images to histopathology tasks. Moreover, the frequent lack of annotations in histopathology patch images has driven researchers to explore self-supervised learning methods like mask reconstruction for learning representations from large amounts of unlabeled data. Crucially, previous mask-based efforts in self-supervised learning have often overlooked the spatial interactions among entities, which are essential for constructing accurate representations of pathological entities. To address these challenges, constructing graphs of entities is a promising approach. In addition, the diffusion reconstruction strategy has recently shown superior performance through its random intensity noise addition technique to enhance the robust learned representation. Therefore, we introduce H-MGDM, a novel self-supervised Histopathology image representation learning method through the Dynamic Entity-Masked Graph Diffusion Model. Specifically, we propose to use complementary subgraphs as latent diffusion conditions and self-supervised targets respectively during pre-training. We note that the graph can embed entities' topological relationships and enhance representation. Dynamic conditions and targets can improve pathological fine reconstruction. Our model has conducted pretraining experiments on three large histopathological datasets. The advanced predictive performance and interpretability of H-MGDM are clearly evaluated on comprehensive downstream tasks such as classification and survival analysis on six datasets. Our code will be publicly available at https://github.com/centurion-crawler/H-MGDM.
DPSeq: A Novel and Efficient Digital Pathology Classifier for Predicting Cancer Biomarkers using Sequencer Architecture
May 03, 2023Abstract:In digital pathology tasks, transformers have achieved state-of-the-art results, surpassing convolutional neural networks (CNNs). However, transformers are usually complex and resource intensive. In this study, we developed a novel and efficient digital pathology classifier called DPSeq, to predict cancer biomarkers through fine-tuning a sequencer architecture integrating horizon and vertical bidirectional long short-term memory (BiLSTM) networks. Using hematoxylin and eosin (H&E)-stained histopathological images of colorectal cancer (CRC) from two international datasets: The Cancer Genome Atlas (TCGA) and Molecular and Cellular Oncology (MCO), the predictive performance of DPSeq was evaluated in series of experiments. DPSeq demonstrated exceptional performance for predicting key biomarkers in CRC (MSI status, Hypermutation, CIMP status, BRAF mutation, TP53 mutation and chromosomal instability [CING]), outperforming most published state-of-the-art classifiers in a within-cohort internal validation and a cross-cohort external validation. Additionally, under the same experimental conditions using the same set of training and testing datasets, DPSeq surpassed 4 CNN (ResNet18, ResNet50, MobileNetV2, and EfficientNet) and 2 transformer (ViT and Swin-T) models, achieving the highest AUROC and AUPRC values in predicting MSI status, BRAF mutation, and CIMP status. Furthermore, DPSeq required less time for both training and prediction due to its simple architecture. Therefore, DPSeq appears to be the preferred choice over transformer and CNN models for predicting cancer biomarkers.
Time to Embrace Natural Language Processing (NLP)-based Digital Pathology: Benchmarking NLP- and Convolutional Neural Network-based Deep Learning Pipelines
Feb 21, 2023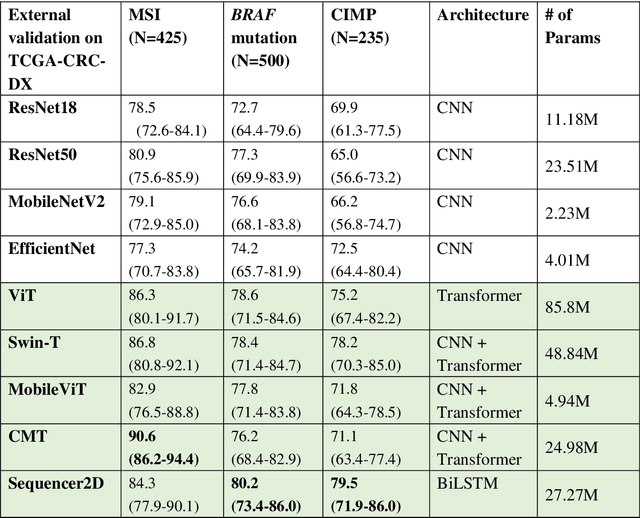
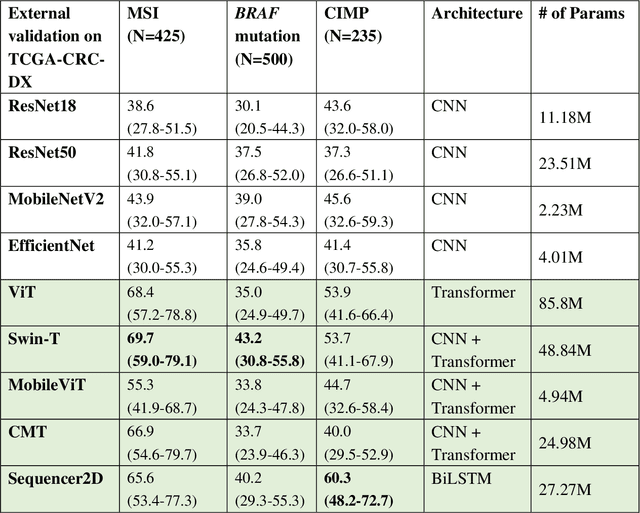
Abstract:NLP-based computer vision models, particularly vision transformers, have been shown to outperform CNN models in many imaging tasks. However, most digital pathology artificial-intelligence models are based on CNN architectures, probably owing to a lack of data regarding NLP models for pathology images. In this study, we developed digital pathology pipelines to benchmark the five most recently proposed NLP models (vision transformer (ViT), Swin Transformer, MobileViT, CMT, and Sequencer2D) and four popular CNN models (ResNet18, ResNet50, MobileNetV2, and EfficientNet) to predict biomarkers in colorectal cancer (microsatellite instability, CpG island methylator phenotype, and BRAF mutation). Hematoxylin and eosin-stained whole-slide images from Molecular and Cellular Oncology and The Cancer Genome Atlas were used as training and external validation datasets, respectively. Cross-study external validations revealed that the NLP-based models significantly outperformed the CNN-based models in biomarker prediction tasks, improving the overall prediction and precision up to approximately 10% and 26%, respectively. Notably, compared with existing models in the current literature using large training datasets, our NLP models achieved state-of-the-art predictions for all three biomarkers using a relatively small training dataset, suggesting that large training datasets are not a prerequisite for NLP models or transformers, and NLP may be more suitable for clinical studies in which small training datasets are commonly collected. The superior performance of Sequencer2D suggests that further research and innovation on both transformer and bidirectional long short-term memory architectures are warranted in the field of digital pathology. NLP models can replace classic CNN architectures and become the new workhorse backbone in the field of digital pathology.
A Novel Few-Shot Relation Extraction Pipeline Based on Adaptive Prototype Fusion
Oct 15, 2022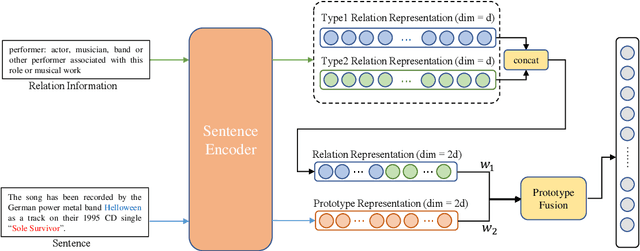

Abstract:Few-shot relation extraction (FSRE) aims at recognizing unseen relations by learning with merely a handful of annotated instances. To more effectively generalize to new relations, this paper proposes a novel pipeline for the FSRE task based on adaptive prototype fusion. Specifically, for each relation class, the pipeline fully explores the relation information by concatenating two types of embedding, and then elaborately combine the relation representation with the adaptive prototype fusion mechanism. The whole framework can be effectively and efficiently optimized in an end-to-end fashion. Experiments on the benchmark dataset FewRel 1.0 show a significant improvement of our method against state-of-the-art methods.
Colorectal cancer survival prediction using deep distribution based multiple-instance learning
Apr 24, 2022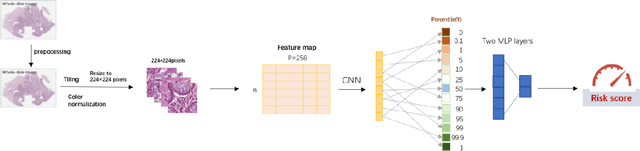
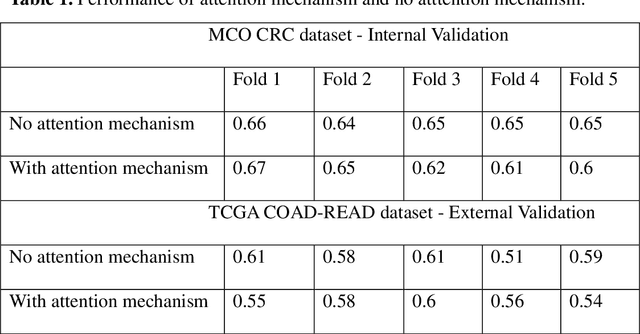
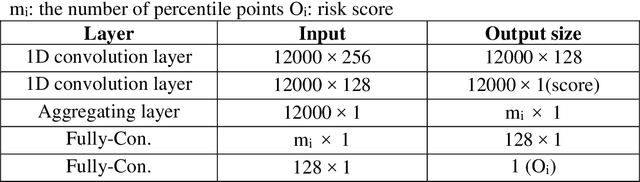
Abstract:Several deep learning algorithms have been developed to predict survival of cancer patients using whole slide images (WSIs).However, identification of image phenotypes within the WSIs that are relevant to patient survival and disease progression is difficult for both clinicians, and deep learning algorithms. Most deep learning based Multiple Instance Learning (MIL) algorithms for survival prediction use either top instances (e.g., maxpooling) or top/bottom instances (e.g., MesoNet) to identify image phenotypes. In this study, we hypothesize that wholistic information of the distribution of the patch scores within a WSI can predict the cancer survival better. We developed a distribution based multiple-instance survival learning algorithm (DeepDisMISL) to validate this hypothesis. We designed and executed experiments using two large international colorectal cancer WSIs datasets - MCO CRC and TCGA COAD-READ. Our results suggest that the more information about the distribution of the patch scores for a WSI, the better is the prediction performance. Including multiple neighborhood instances around each selected distribution location (e.g., percentiles) could further improve the prediction. DeepDisMISL demonstrated superior predictive ability compared to other recently published, state-of-the-art algorithms. Furthermore, our algorithm is interpretable and could assist in understanding the relationship between cancer morphological phenotypes and patients cancer survival risk.
Improving Feature Extraction from Histopathological Images Through A Fine-tuning ImageNet Model
Jan 03, 2022Abstract:Due to lack of annotated pathological images, transfer learning has been the predominant approach in the field of digital pathology.Pre-trained neural networks based on ImageNet database are often used to extract "off the shelf" features, achieving great success in predicting tissue types, molecular features, and clinical outcomes, etc. We hypothesize that fine-tuning the pre-trained models using histopathological images could further improve feature extraction, and downstream prediction performance.We used 100,000 annotated HE image patches for colorectal cancer (CRC) to finetune a pretrained Xception model via a twostep approach.The features extracted from finetuned Xception (FTX2048) model and Imagepretrained (IMGNET2048) model were compared through: (1) tissue classification for HE images from CRC, same image type that was used for finetuning; (2) prediction of immunerelated gene expression and (3) gene mutations for lung adenocarcinoma (LUAD).Fivefold cross validation was used for model performance evaluation. The extracted features from the finetuned FTX2048 exhibited significantly higher accuracy for predicting tisue types of CRC compared to the off the shelf feature directly from Xception based on ImageNet database. Particularly, FTX2048 markedly improved the accuracy for stroma from 87% to 94%. Similarly, features from FTX2048 boosted the prediction of transcriptomic expression of immunerelated genesin LUAD. For the genes that had signigicant relationships with image fetures, the features fgrom the finetuned model imprroved the prediction for the majority of the genes. Inaddition, fetures from FTX2048 improved prediction of mutation for 5 out of 9 most frequently mutated genes in LUAD.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge