Michael Ingrisch
Post-Training Network Compression for 3D Medical Image Segmentation: Reducing Computational Efforts via Tucker Decomposition
Apr 15, 2024Abstract:We address the computational barrier of deploying advanced deep learning segmentation models in clinical settings by studying the efficacy of network compression through tensor decomposition. We propose a post-training Tucker factorization that enables the decomposition of pre-existing models to reduce computational requirements without impeding segmentation accuracy. We applied Tucker decomposition to the convolutional kernels of the TotalSegmentator (TS) model, an nnU-Net model trained on a comprehensive dataset for automatic segmentation of 117 anatomical structures. Our approach reduced the floating-point operations (FLOPs) and memory required during inference, offering an adjustable trade-off between computational efficiency and segmentation quality. This study utilized the publicly available TS dataset, employing various downsampling factors to explore the relationship between model size, inference speed, and segmentation performance. The application of Tucker decomposition to the TS model substantially reduced the model parameters and FLOPs across various compression rates, with limited loss in segmentation accuracy. We removed up to 88% of the model's parameters with no significant performance changes in the majority of classes after fine-tuning. Practical benefits varied across different graphics processing unit (GPU) architectures, with more distinct speed-ups on less powerful hardware. Post-hoc network compression via Tucker decomposition presents a viable strategy for reducing the computational demand of medical image segmentation models without substantially sacrificing accuracy. This approach enables the broader adoption of advanced deep learning technologies in clinical practice, offering a way to navigate the constraints of hardware capabilities.
Unreading Race: Purging Protected Features from Chest X-ray Embeddings
Nov 02, 2023Abstract:Purpose: To analyze and remove protected feature effects in chest radiograph embeddings of deep learning models. Materials and Methods: An orthogonalization is utilized to remove the influence of protected features (e.g., age, sex, race) in chest radiograph embeddings, ensuring feature-independent results. To validate the efficacy of the approach, we retrospectively study the MIMIC and CheXpert datasets using three pre-trained models, namely a supervised contrastive, a self-supervised contrastive, and a baseline classifier model. Our statistical analysis involves comparing the original versus the orthogonalized embeddings by estimating protected feature influences and evaluating the ability to predict race, age, or sex using the two types of embeddings. Results: Our experiments reveal a significant influence of protected features on predictions of pathologies. Applying orthogonalization removes these feature effects. Apart from removing any influence on pathology classification, while maintaining competitive predictive performance, orthogonalized embeddings further make it infeasible to directly predict protected attributes and mitigate subgroup disparities. Conclusion: The presented work demonstrates the successful application and evaluation of the orthogonalization technique in the domain of chest X-ray classification.
Exploring the Impact of Image Resolution on Chest X-ray Classification Performance
Jun 09, 2023



Abstract:Deep learning models for image classification have often used a resolution of $224\times224$ pixels for computational reasons. This study investigates the effect of image resolution on chest X-ray classification performance, using the ChestX-ray14 dataset. The results show that a higher image resolution, specifically $1024\times1024$ pixels, has the best overall classification performance, with a slight decline in performance between $256\times256$ to $512\times512$ pixels for most of the pathological classes. Comparison of saliency map-generated bounding boxes revealed that commonly used resolutions are insufficient for finding most pathologies.
Automated Labeling of German Chest X-Ray Radiology Reports using Deep Learning
Jun 09, 2023



Abstract:Radiologists are in short supply globally, and deep learning models offer a promising solution to address this shortage as part of clinical decision-support systems. However, training such models often requires expensive and time-consuming manual labeling of large datasets. Automatic label extraction from radiology reports can reduce the time required to obtain labeled datasets, but this task is challenging due to semantically similar words and missing annotated data. In this work, we explore the potential of weak supervision of a deep learning-based label prediction model, using a rule-based labeler. We propose a deep learning-based CheXpert label prediction model, pre-trained on reports labeled by a rule-based German CheXpert model and fine-tuned on a small dataset of manually labeled reports. Our results demonstrate the effectiveness of our approach, which significantly outperformed the rule-based model on all three tasks. Our findings highlight the benefits of employing deep learning-based models even in scenarios with sparse data and the use of the rule-based labeler as a tool for weak supervision.
WindowNet: Learnable Windows for Chest X-ray Classification
Jun 09, 2023



Abstract:Chest X-ray (CXR) images are commonly compressed to a lower resolution and bit depth to reduce their size, potentially altering subtle diagnostic features. Radiologists use windowing operations to enhance image contrast, but the impact of such operations on CXR classification performance is unclear. In this study, we show that windowing can improve CXR classification performance, and propose WindowNet, a model that learns optimal window settings. We first investigate the impact of bit-depth on classification performance and find that a higher bit-depth (12-bit) leads to improved performance. We then evaluate different windowing settings and show that training with a distinct window generally improves pathology-wise classification performance. Finally, we propose and evaluate WindowNet, a model that learns optimal window settings, and show that it significantly improves performance compared to the baseline model without windowing.
German CheXpert Chest X-ray Radiology Report Labeler
Jun 05, 2023Abstract:This study aimed to develop an algorithm to automatically extract annotations for chest X-ray classification models from German thoracic radiology reports. An automatic label extraction model was designed based on the CheXpert architecture, and a web-based annotation interface was created for iterative improvements. Results showed that automated label extraction can reduce time spent on manual labeling and improve overall modeling performance. The model trained on automatically extracted labels performed competitively to manually labeled data and strongly outperformed the model trained on publicly available data.
Constrained Probabilistic Mask Learning for Task-specific Undersampled MRI Reconstruction
May 25, 2023



Abstract:Undersampling is a common method in Magnetic Resonance Imaging (MRI) to subsample the number of data points in k-space and thereby reduce acquisition times at the cost of decreased image quality. In this work, we directly learn the undersampling masks to derive task- and domain-specific patterns. To solve this discrete optimization challenge, we propose a general optimization routine called ProM: A fully probabilistic, differentiable, versatile, and model-free framework for mask optimization that enforces acceleration factors through a convex constraint. Analyzing knee, brain, and cardiac MRI datasets with our method, we discover that different anatomic regions reveal distinct optimal undersampling masks. Furthermore, ProM can create undersampling masks that maximize performance in downstream tasks like segmentation with networks trained on fully-sampled MRIs. Even with extreme acceleration factors, ProM yields reasonable performance while being more versatile than existing methods, paving the way for data-driven all-purpose mask generation.
Cascaded Latent Diffusion Models for High-Resolution Chest X-ray Synthesis
Mar 20, 2023Abstract:While recent advances in large-scale foundational models show promising results, their application to the medical domain has not yet been explored in detail. In this paper, we progress into the realms of large-scale modeling in medical synthesis by proposing Cheff - a foundational cascaded latent diffusion model, which generates highly-realistic chest radiographs providing state-of-the-art quality on a 1-megapixel scale. We further propose MaCheX, which is a unified interface for public chest datasets and forms the largest open collection of chest X-rays up to date. With Cheff conditioned on radiological reports, we further guide the synthesis process over text prompts and unveil the research area of report-to-chest-X-ray generation.
ChatGPT Makes Medicine Easy to Swallow: An Exploratory Case Study on Simplified Radiology Reports
Dec 30, 2022Abstract:The release of ChatGPT, a language model capable of generating text that appears human-like and authentic, has gained significant attention beyond the research community. We expect that the convincing performance of ChatGPT incentivizes users to apply it to a variety of downstream tasks, including prompting the model to simplify their own medical reports. To investigate this phenomenon, we conducted an exploratory case study. In a questionnaire, we asked 15 radiologists to assess the quality of radiology reports simplified by ChatGPT. Most radiologists agreed that the simplified reports were factually correct, complete, and not potentially harmful to the patient. Nevertheless, instances of incorrect statements, missed key medical findings, and potentially harmful passages were reported. While further studies are needed, the initial insights of this study indicate a great potential in using large language models like ChatGPT to improve patient-centered care in radiology and other medical domains.
A knee cannot have lung disease: out-of-distribution detection with in-distribution voting using the medical example of chest X-ray classification
Aug 01, 2022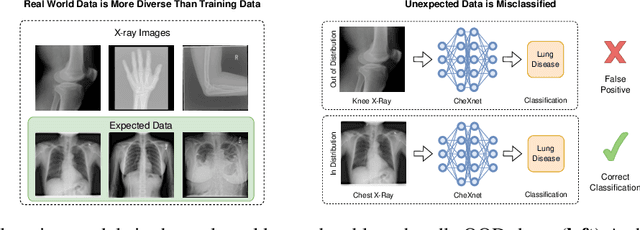
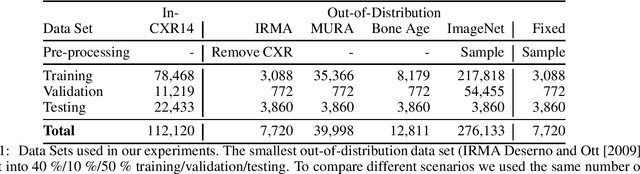
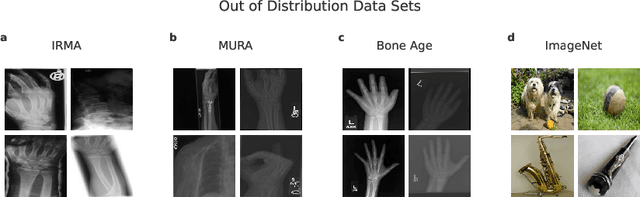
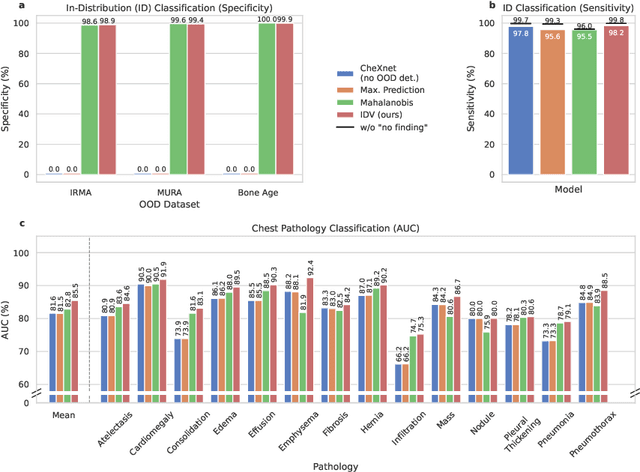
Abstract:Deep learning models are being applied to more and more use cases with astonishing success stories, but how do they perform in the real world? To test a model, a specific cleaned data set is assembled. However, when deployed in the real world, the model will face unexpected, out-of-distribution (OOD) data. In this work, we show that the so-called "radiologist-level" CheXnet model fails to recognize all OOD images and classifies them as having lung disease. To address this issue, we propose in-distribution voting, a novel method to classify out-of-distribution images for multi-label classification. Using independent class-wise in-distribution (ID) predictors trained on ID and OOD data we achieve, on average, 99 % ID classification specificity and 98 % sensitivity, improving the end-to-end performance significantly compared to previous works on the chest X-ray 14 data set. Our method surpasses other output-based OOD detectors even when trained solely with ImageNet as OOD data and tested with X-ray OOD images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge