Medha Sawhney
TaxaAdapter: Vision Taxonomy Models are Key to Fine-grained Image Generation over the Tree of Life
Mar 27, 2026Abstract:Accurately generating images across the Tree of Life is difficult: there are over 10M distinct species on Earth, many of which differ only by subtle visual traits. Despite the remarkable progress in text-to-image synthesis, existing models often fail to capture the fine-grained visual cues that define species identity, even when their outputs appear photo-realistic. To this end, we propose TaxaAdapter, a simple and lightweight approach that incorporates Vision Taxonomy Models (VTMs) such as BioCLIP to guide fine-grained species generation. Our method injects VTM embeddings into a frozen text-to-image diffusion model, improving species-level fidelity while preserving flexible text control over attributes such as pose, style, and background. Extensive experiments demonstrate that TaxaAdapter consistently improves morphology fidelity and species-identity accuracy over strong baselines, with a cleaner architecture and training recipe. To better evaluate these improvements, we also introduce a multimodal Large Language Model-based metric that summarizes trait-level descriptions from generated and real images, providing a more interpretable measure of morphological consistency. Beyond this, we observe that TaxaAdapter exhibits strong generalization capabilities, enabling species synthesis in challenging regimes such as few-shot species with only a handful of training images and even species unseen during training. Overall, our results highlight that VTMs are a key ingredient for scalable, fine-grained species generation.
Open World Scene Graph Generation using Vision Language Models
Jun 09, 2025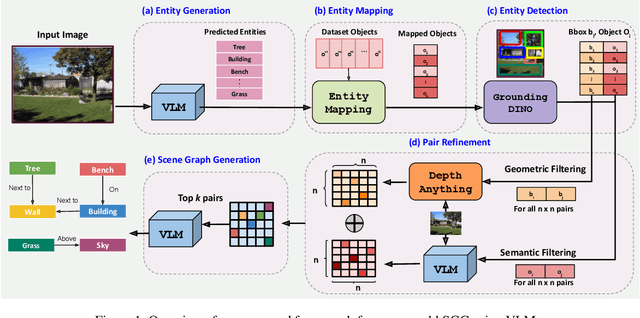
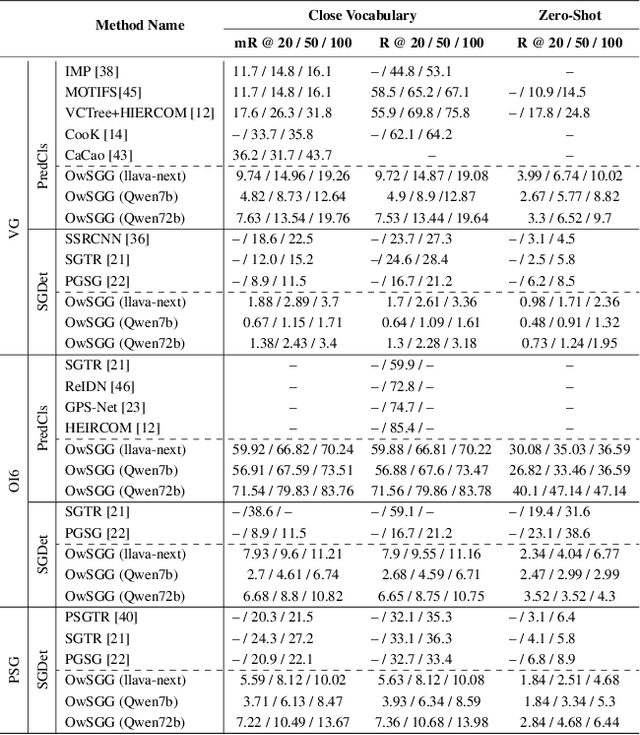
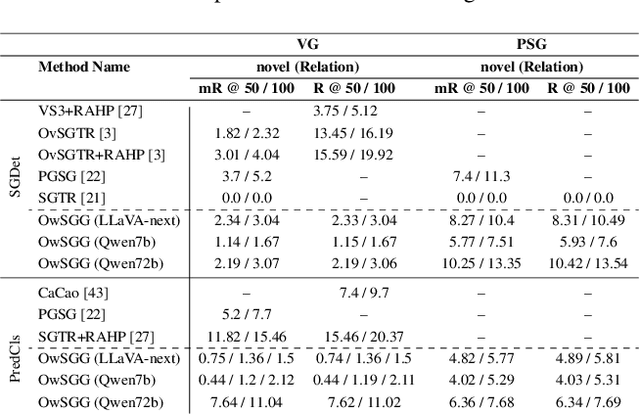
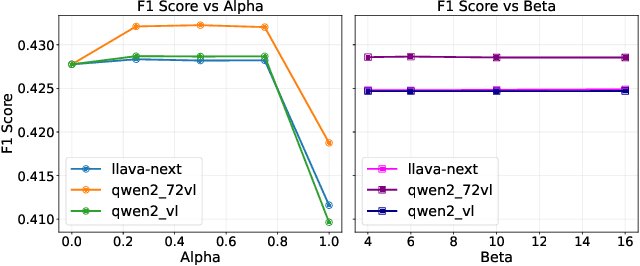
Abstract:Scene-Graph Generation (SGG) seeks to recognize objects in an image and distill their salient pairwise relationships. Most methods depend on dataset-specific supervision to learn the variety of interactions, restricting their usefulness in open-world settings, involving novel objects and/or relations. Even methods that leverage large Vision Language Models (VLMs) typically require benchmark-specific fine-tuning. We introduce Open-World SGG, a training-free, efficient, model-agnostic framework that taps directly into the pretrained knowledge of VLMs to produce scene graphs with zero additional learning. Casting SGG as a zero-shot structured-reasoning problem, our method combines multimodal prompting, embedding alignment, and a lightweight pair-refinement strategy, enabling inference over unseen object vocabularies and relation sets. To assess this setting, we formalize an Open-World evaluation protocol that measures performance when no SGG-specific data have been observed either in terms of objects and relations. Experiments on Visual Genome, Open Images V6, and the Panoptic Scene Graph (PSG) dataset demonstrate the capacity of pretrained VLMs to perform relational understanding without task-level training.
A Unified Framework for Forward and Inverse Problems in Subsurface Imaging using Latent Space Translations
Oct 15, 2024Abstract:In subsurface imaging, learning the mapping from velocity maps to seismic waveforms (forward problem) and waveforms to velocity (inverse problem) is important for several applications. While traditional techniques for solving forward and inverse problems are computationally prohibitive, there is a growing interest in leveraging recent advances in deep learning to learn the mapping between velocity maps and seismic waveform images directly from data. Despite the variety of architectures explored in previous works, several open questions still remain unanswered such as the effect of latent space sizes, the importance of manifold learning, the complexity of translation models, and the value of jointly solving forward and inverse problems. We propose a unified framework to systematically characterize prior research in this area termed the Generalized Forward-Inverse (GFI) framework, building on the assumption of manifolds and latent space translations. We show that GFI encompasses previous works in deep learning for subsurface imaging, which can be viewed as specific instantiations of GFI. We also propose two new model architectures within the framework of GFI: Latent U-Net and Invertible X-Net, leveraging the power of U-Nets for domain translation and the ability of IU-Nets to simultaneously learn forward and inverse translations, respectively. We show that our proposed models achieve state-of-the-art (SOTA) performance for forward and inverse problems on a wide range of synthetic datasets, and also investigate their zero-shot effectiveness on two real-world-like datasets.
VLM4Bio: A Benchmark Dataset to Evaluate Pretrained Vision-Language Models for Trait Discovery from Biological Images
Aug 28, 2024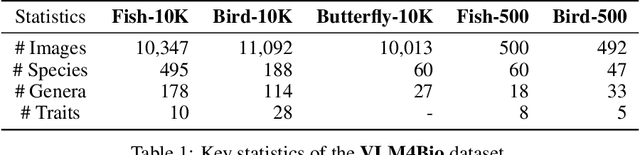
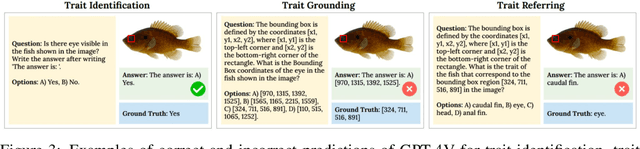
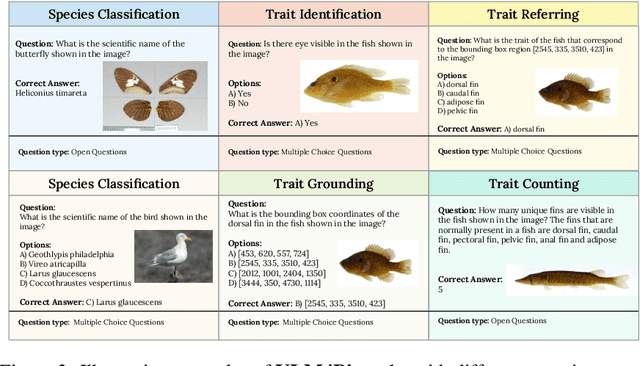


Abstract:Images are increasingly becoming the currency for documenting biodiversity on the planet, providing novel opportunities for accelerating scientific discoveries in the field of organismal biology, especially with the advent of large vision-language models (VLMs). We ask if pre-trained VLMs can aid scientists in answering a range of biologically relevant questions without any additional fine-tuning. In this paper, we evaluate the effectiveness of 12 state-of-the-art (SOTA) VLMs in the field of organismal biology using a novel dataset, VLM4Bio, consisting of 469K question-answer pairs involving 30K images from three groups of organisms: fishes, birds, and butterflies, covering five biologically relevant tasks. We also explore the effects of applying prompting techniques and tests for reasoning hallucination on the performance of VLMs, shedding new light on the capabilities of current SOTA VLMs in answering biologically relevant questions using images. The code and datasets for running all the analyses reported in this paper can be found at https://github.com/sammarfy/VLM4Bio.
MEMTRACK: A Deep Learning-Based Approach to Microrobot Tracking in Dense and Low-Contrast Environments
Oct 13, 2023



Abstract:Tracking microrobots is challenging, considering their minute size and high speed. As the field progresses towards developing microrobots for biomedical applications and conducting mechanistic studies in physiologically relevant media (e.g., collagen), this challenge is exacerbated by the dense surrounding environments with feature size and shape comparable to microrobots. Herein, we report Motion Enhanced Multi-level Tracker (MEMTrack), a robust pipeline for detecting and tracking microrobots using synthetic motion features, deep learning-based object detection, and a modified Simple Online and Real-time Tracking (SORT) algorithm with interpolation for tracking. Our object detection approach combines different models based on the object's motion pattern. We trained and validated our model using bacterial micro-motors in collagen (tissue phantom) and tested it in collagen and aqueous media. We demonstrate that MEMTrack accurately tracks even the most challenging bacteria missed by skilled human annotators, achieving precision and recall of 77% and 48% in collagen and 94% and 35% in liquid media, respectively. Moreover, we show that MEMTrack can quantify average bacteria speed with no statistically significant difference from the laboriously-produced manual tracking data. MEMTrack represents a significant contribution to microrobot localization and tracking, and opens the potential for vision-based deep learning approaches to microrobot control in dense and low-contrast settings. All source code for training and testing MEMTrack and reproducing the results of the paper have been made publicly available https://github.com/sawhney-medha/MEMTrack.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge