Matteo Salvador
Full-field surrogate modeling of cardiac function encoding geometric variability
Apr 29, 2025Abstract:Combining physics-based modeling with data-driven methods is critical to enabling the translation of computational methods to clinical use in cardiology. The use of rigorous differential equations combined with machine learning tools allows for model personalization with uncertainty quantification in time frames compatible with clinical practice. However, accurate and efficient surrogate models of cardiac function, built from physics-based numerical simulation, are still mostly geometry-specific and require retraining for different patients and pathological conditions. We propose a novel computational pipeline to embed cardiac anatomies into full-field surrogate models. We generate a dataset of electrophysiology simulations using a complex multi-scale mathematical model coupling partial and ordinary differential equations. We adopt Branched Latent Neural Maps (BLNMs) as an effective scientific machine learning method to encode activation maps extracted from physics-based numerical simulations into a neural network. Leveraging large deformation diffeomorphic metric mappings, we build a biventricular anatomical atlas and parametrize the anatomical variability of a small and challenging cohort of 13 pediatric patients affected by Tetralogy of Fallot. We propose a novel statistical shape modeling based z-score sampling approach to generate a new synthetic cohort of 52 biventricular geometries that are compatible with the original geometrical variability. This synthetic cohort acts as the training set for BLNMs. Our surrogate model demonstrates robustness and great generalization across the complex original patient cohort, achieving an average adimensional mean squared error of 0.0034. The Python implementation of our BLNM model is publicly available under MIT License at https://github.com/StanfordCBCL/BLNM.
Liquid Fourier Latent Dynamics Networks for fast GPU-based numerical simulations in computational cardiology
Aug 19, 2024



Abstract:Scientific Machine Learning (ML) is gaining momentum as a cost-effective alternative to physics-based numerical solvers in many engineering applications. In fact, scientific ML is currently being used to build accurate and efficient surrogate models starting from high-fidelity numerical simulations, effectively encoding the parameterized temporal dynamics underlying Ordinary Differential Equations (ODEs), or even the spatio-temporal behavior underlying Partial Differential Equations (PDEs), in appropriately designed neural networks. We propose an extension of Latent Dynamics Networks (LDNets), namely Liquid Fourier LDNets (LFLDNets), to create parameterized space-time surrogate models for multiscale and multiphysics sets of highly nonlinear differential equations on complex geometries. LFLDNets employ a neurologically-inspired, sparse, liquid neural network for temporal dynamics, relaxing the requirement of a numerical solver for time advancement and leading to superior performance in terms of tunable parameters, accuracy, efficiency and learned trajectories with respect to neural ODEs based on feedforward fully-connected neural networks. Furthermore, in our implementation of LFLDNets, we use a Fourier embedding with a tunable kernel in the reconstruction network to learn high-frequency functions better and faster than using space coordinates directly as input. We challenge LFLDNets in the framework of computational cardiology and evaluate their capabilities on two 3-dimensional test cases arising from multiscale cardiac electrophysiology and cardiovascular hemodynamics. This paper illustrates the capability to run Artificial Intelligence-based numerical simulations on single or multiple GPUs in a matter of minutes and represents a significant step forward in the development of physics-informed digital twins.
NeurAM: nonlinear dimensionality reduction for uncertainty quantification through neural active manifolds
Aug 07, 2024Abstract:We present a new approach for nonlinear dimensionality reduction, specifically designed for computationally expensive mathematical models. We leverage autoencoders to discover a one-dimensional neural active manifold (NeurAM) capturing the model output variability, plus a simultaneously learnt surrogate model with inputs on this manifold. The proposed dimensionality reduction framework can then be applied to perform outer loop many-query tasks, like sensitivity analysis and uncertainty propagation. In particular, we prove, both theoretically under idealized conditions, and numerically in challenging test cases, how NeurAM can be used to obtain multifidelity sampling estimators with reduced variance by sampling the models on the discovered low-dimensional and shared manifold among models. Several numerical examples illustrate the main features of the proposed dimensionality reduction strategy, and highlight its advantages with respect to existing approaches in the literature.
Branched Latent Neural Operators
Aug 04, 2023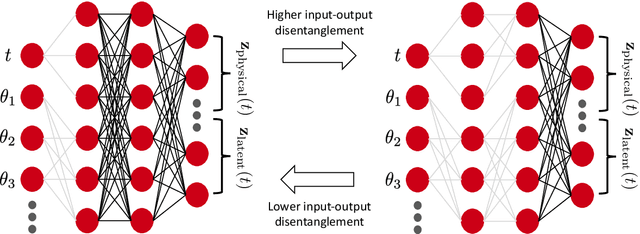



Abstract:We introduce Branched Latent Neural Operators (BLNOs) to learn input-output maps encoding complex physical processes. A BLNO is defined by a simple and compact feedforward partially-connected neural network that structurally disentangles inputs with different intrinsic roles, such as the time variable from model parameters of a differential equation, while transferring them into a generic field of interest. BLNOs leverage interpretable latent outputs to enhance the learned dynamics and break the curse of dimensionality by showing excellent generalization properties with small training datasets and short training times on a single processor. Indeed, their generalization error remains comparable regardless of the adopted discretization during the testing phase. Moreover, the partial connections, in place of a fully-connected structure, significantly reduce the number of tunable parameters. We show the capabilities of BLNOs in a challenging test case involving biophysically detailed electrophysiology simulations in a biventricular cardiac model of a pediatric patient with hypoplastic left heart syndrome. The model includes a purkinje network for fast conduction and a heart-torso geometry. Specifically, we trained BLNOs on 150 in silico generated 12-lead electrocardiograms (ECGs) while spanning 7 model parameters, covering cell-scale, organ-level and electrical dyssynchrony. Although the 12-lead ECGs manifest very fast dynamics with sharp gradients, after automatic hyperparameter tuning the optimal BLNO, trained in less than 3 hours on a single CPU, retains just 7 hidden layers and 19 neurons per layer. The mean square error is on the order of $10^{-4}$ on an independent test dataset comprised of 50 additional electrophysiology simulations. This paper provides a novel computational tool to build reliable and efficient reduced-order models for digital twinning in engineering applications.
Real-time whole-heart electromechanical simulations using Latent Neural Ordinary Differential Equations
Jun 08, 2023



Abstract:Cardiac digital twins provide a physics and physiology informed framework to deliver predictive and personalized medicine. However, high-fidelity multi-scale cardiac models remain a barrier to adoption due to their extensive computational costs and the high number of model evaluations needed for patient-specific personalization. Artificial Intelligence-based methods can make the creation of fast and accurate whole-heart digital twins feasible. In this work, we use Latent Neural Ordinary Differential Equations (LNODEs) to learn the temporal pressure-volume dynamics of a heart failure patient. Our surrogate model based on LNODEs is trained from 400 3D-0D whole-heart closed-loop electromechanical simulations while accounting for 43 model parameters, describing single cell through to whole organ and cardiovascular hemodynamics. The trained LNODEs provides a compact and efficient representation of the 3D-0D model in a latent space by means of a feedforward fully-connected Artificial Neural Network that retains 3 hidden layers with 13 neurons per layer and allows for 300x real-time numerical simulations of the cardiac function on a single processor of a standard laptop. This surrogate model is employed to perform global sensitivity analysis and robust parameter estimation with uncertainty quantification in 3 hours of computations, still on a single processor. We match pressure and volume time traces unseen by the LNODEs during the training phase and we calibrate 4 to 11 model parameters while also providing their posterior distribution. This paper introduces the most advanced surrogate model of cardiac function available in the literature and opens new important venues for parameter calibration in cardiac digital twins.
Latent Dynamics Networks (LDNets): learning the intrinsic dynamics of spatio-temporal processes
Apr 28, 2023



Abstract:Predicting the evolution of systems that exhibit spatio-temporal dynamics in response to external stimuli is a key enabling technology fostering scientific innovation. Traditional equations-based approaches leverage first principles to yield predictions through the numerical approximation of high-dimensional systems of differential equations, thus calling for large-scale parallel computing platforms and requiring large computational costs. Data-driven approaches, instead, enable the description of systems evolution in low-dimensional latent spaces, by leveraging dimensionality reduction and deep learning algorithms. We propose a novel architecture, named Latent Dynamics Network (LDNet), which is able to discover low-dimensional intrinsic dynamics of possibly non-Markovian dynamical systems, thus predicting the time evolution of space-dependent fields in response to external inputs. Unlike popular approaches, in which the latent representation of the solution manifold is learned by means of auto-encoders that map a high-dimensional discretization of the system state into itself, LDNets automatically discover a low-dimensional manifold while learning the latent dynamics, without ever operating in the high-dimensional space. Furthermore, LDNets are meshless algorithms that do not reconstruct the output on a predetermined grid of points, but rather at any point of the domain, thus enabling weight-sharing across query-points. These features make LDNets lightweight and easy-to-train, with excellent accuracy and generalization properties, even in time-extrapolation regimes. We validate our method on several test cases and we show that, for a challenging highly-nonlinear problem, LDNets outperform state-of-the-art methods in terms of accuracy (normalized error 5 times smaller), by employing a dramatically smaller number of trainable parameters (more than 10 times fewer).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge