Luxia Zhang
National Institute of Health Data Science, Peking University, Beijing, China, Renal Division, Department of Medicine, Peking University First Hospital, Beijing, China, Research Units of Diagnosis and Treatment of Immune-Mediated Kidney Diseases, Chinese Academy of Medical Sciences, Beijing, China, State Key Laboratory of Vascular Homeostasis and Remodeling, Peking University, Beijing, China, Center for Digital Health and Artificial Intelligence, Peking University First Hospital, Beijing, China
KidneyTalk-open: No-code Deployment of a Private Large Language Model with Medical Documentation-Enhanced Knowledge Database for Kidney Disease
Mar 06, 2025Abstract:Privacy-preserving medical decision support for kidney disease requires localized deployment of large language models (LLMs) while maintaining clinical reasoning capabilities. Current solutions face three challenges: 1) Cloud-based LLMs pose data security risks; 2) Local model deployment demands technical expertise; 3) General LLMs lack mechanisms to integrate medical knowledge. Retrieval-augmented systems also struggle with medical document processing and clinical usability. We developed KidneyTalk-open, a desktop system integrating three technical components: 1) No-code deployment of state-of-the-art (SOTA) open-source LLMs (such as DeepSeek-r1, Qwen2.5) via local inference engine; 2) Medical document processing pipeline combining context-aware chunking and intelligent filtering; 3) Adaptive Retrieval and Augmentation Pipeline (AddRep) employing agents collaboration for improving the recall rate of medical documents. A graphical interface was designed to enable clinicians to manage medical documents and conduct AI-powered consultations without technical expertise. Experimental validation on 1,455 challenging nephrology exam questions demonstrates AddRep's effectiveness: achieving 29.1% accuracy (+8.1% over baseline) with intelligent knowledge integration, while maintaining robustness through 4.9% rejection rate to suppress hallucinations. Comparative case studies with the mainstream products (AnythingLLM, Chatbox, GPT4ALL) demonstrate KidneyTalk-open's superior performance in real clinical query. KidneyTalk-open represents the first no-code medical LLM system enabling secure documentation-enhanced medical Q&A on desktop. Its designs establishes a new framework for privacy-sensitive clinical AI applications. The system significantly lowers technical barriers while improving evidence traceability, enabling more medical staff or patients to use SOTA open-source LLMs conveniently.
ChatSchema: A pipeline of extracting structured information with Large Multimodal Models based on schema
Jul 26, 2024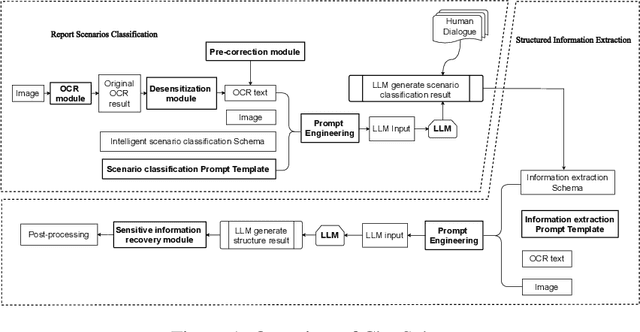

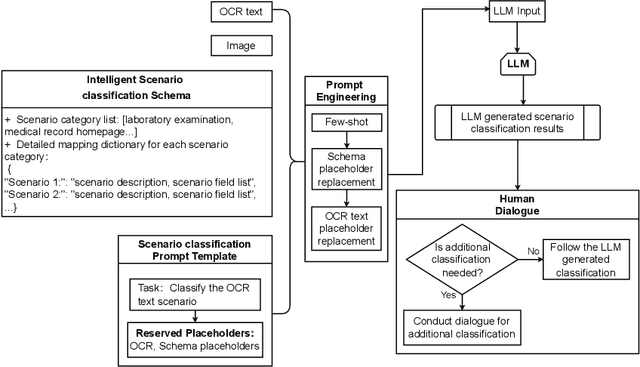

Abstract:Objective: This study introduces ChatSchema, an effective method for extracting and structuring information from unstructured data in medical paper reports using a combination of Large Multimodal Models (LMMs) and Optical Character Recognition (OCR) based on the schema. By integrating predefined schema, we intend to enable LMMs to directly extract and standardize information according to the schema specifications, facilitating further data entry. Method: Our approach involves a two-stage process, including classification and extraction for categorizing report scenarios and structuring information. We established and annotated a dataset to verify the effectiveness of ChatSchema, and evaluated key extraction using precision, recall, F1-score, and accuracy metrics. Based on key extraction, we further assessed value extraction. We conducted ablation studies on two LMMs to illustrate the improvement of structured information extraction with different input modals and methods. Result: We analyzed 100 medical reports from Peking University First Hospital and established a ground truth dataset with 2,945 key-value pairs. We evaluated ChatSchema using GPT-4o and Gemini 1.5 Pro and found a higher overall performance of GPT-4o. The results are as follows: For the result of key extraction, key-precision was 98.6%, key-recall was 98.5%, key-F1-score was 98.6%. For the result of value extraction based on correct key extraction, the overall accuracy was 97.2%, precision was 95.8%, recall was 95.8%, and F1-score was 95.8%. An ablation study demonstrated that ChatSchema achieved significantly higher overall accuracy and overall F1-score of key-value extraction, compared to the Baseline, with increases of 26.9% overall accuracy and 27.4% overall F1-score, respectively.
Iterative Bilinear Temporal-Spectral Fusion for Unsupervised Time-Series Representation Learning
Feb 16, 2022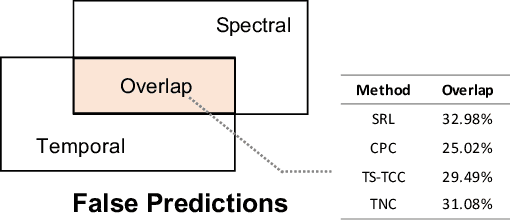

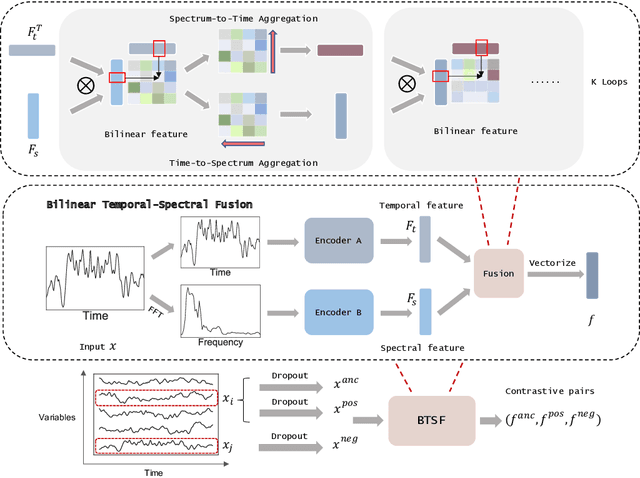

Abstract:Unsupervised/self-supervised time series representation learning is a challenging problem because of its complex dynamics and sparse annotations. Existing works mainly adopt the framework of contrastive learning with the time-based augmentation techniques to sample positives and negatives for contrastive training. Nevertheless, they mostly use segment-level augmentation derived from time slicing, which may bring about sampling bias and incorrect optimization with false negatives due to the loss of global context. Besides, they all pay no attention to incorporate the spectral information in feature representation. In this paper, we propose a unified framework, namely Bilinear Temporal-Spectral Fusion (BTSF). Specifically, we firstly utilize the instance-level augmentation with a simple dropout on the entire time series for maximally capturing long-term dependencies. We devise a novel iterative bilinear temporal-spectral fusion to explicitly encode the affinities of abundant time-frequency pairs, and iteratively refines representations in a fusion-and-squeeze manner with Spectrum-to-Time (S2T) and Time-to-Spectrum (T2S) Aggregation modules. We firstly conducts downstream evaluations on three major tasks for time series including classification, forecasting and anomaly detection. Experimental results shows that our BTSF consistently significantly outperforms the state-of-the-art methods.
Spectral Propagation Graph Network for Few-shot Time Series Classification
Feb 08, 2022



Abstract:Few-shot Time Series Classification (few-shot TSC) is a challenging problem in time series analysis. It is more difficult to classify when time series of the same class are not completely consistent in spectral domain or time series of different classes are partly consistent in spectral domain. To address this problem, we propose a novel method named Spectral Propagation Graph Network (SPGN) to explicitly model and propagate the spectrum-wise relations between different time series with graph network. To the best of our knowledge, SPGN is the first to utilize spectral comparisons in different intervals and involve spectral propagation across all time series with graph networks for few-shot TSC. SPGN first uses bandpass filter to expand time series in spectral domain for calculating spectrum-wise relations between time series. Equipped with graph networks, SPGN then integrates spectral relations with label information to make spectral propagation. The further study conveys the bi-directional effect between spectral relations acquisition and spectral propagation. We conduct extensive experiments on few-shot TSC benchmarks. SPGN outperforms state-of-the-art results by a large margin in $4\% \sim 13\%$. Moreover, SPGN surpasses them by around $12\%$ and $9\%$ under cross-domain and cross-way settings respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge