Lucas Mansilla
In-Context Reverse Classification Accuracy: Efficient Estimation of Segmentation Quality without Ground-Truth
Mar 06, 2025Abstract:Assessing the quality of automatic image segmentation is crucial in clinical practice, but often very challenging due to the limited availability of ground truth annotations. In this paper, we introduce In-Context Reverse Classification Accuracy (In-Context RCA), a novel framework for automatically estimating segmentation quality in the absence of ground-truth annotations. By leveraging recent in-context learning segmentation models and incorporating retrieval-augmentation techniques to select the most relevant reference images, our approach enables efficient quality estimation with minimal reference data. Validated across diverse medical imaging modalities, our method demonstrates robust performance and computational efficiency, offering a promising solution for automated quality control in clinical workflows, where fast and reliable segmentation assessment is essential. The code is available at https://github.com/mcosarinsky/In-Context-RCA.
Unsupervised bias discovery in medical image segmentation
Sep 01, 2023Abstract:It has recently been shown that deep learning models for anatomical segmentation in medical images can exhibit biases against certain sub-populations defined in terms of protected attributes like sex or ethnicity. In this context, auditing fairness of deep segmentation models becomes crucial. However, such audit process generally requires access to ground-truth segmentation masks for the target population, which may not always be available, especially when going from development to deployment. Here we propose a new method to anticipate model biases in biomedical image segmentation in the absence of ground-truth annotations. Our unsupervised bias discovery method leverages the reverse classification accuracy framework to estimate segmentation quality. Through numerical experiments in synthetic and realistic scenarios we show how our method is able to successfully anticipate fairness issues in the absence of ground-truth labels, constituting a novel and valuable tool in this field.
CheXmask: a large-scale dataset of anatomical segmentation masks for multi-center chest x-ray images
Jul 06, 2023Abstract:The development of successful artificial intelligence models for chest X-ray analysis relies on large, diverse datasets with high-quality annotations. While several databases of chest X-ray images have been released, most include disease diagnosis labels but lack detailed pixel-level anatomical segmentation labels. To address this gap, we introduce an extensive chest X-ray multi-center segmentation dataset with uniform and fine-grain anatomical annotations for images coming from six well-known publicly available databases: CANDID-PTX, ChestX-ray8, Chexpert, MIMIC-CXR-JPG, Padchest, and VinDr-CXR, resulting in 676,803 segmentation masks. Our methodology utilizes the HybridGNet model to ensure consistent and high-quality segmentations across all datasets. Rigorous validation, including expert physician evaluation and automatic quality control, was conducted to validate the resulting masks. Additionally, we provide individualized quality indices per mask and an overall quality estimation per dataset. This dataset serves as a valuable resource for the broader scientific community, streamlining the development and assessment of innovative methodologies in chest X-ray analysis. The CheXmask dataset is publicly available at: \url{https://physionet.org/content/chexmask-cxr-segmentation-data/}.
Improving anatomical plausibility in medical image segmentation via hybrid graph neural networks: applications to chest x-ray analysis
Apr 01, 2022

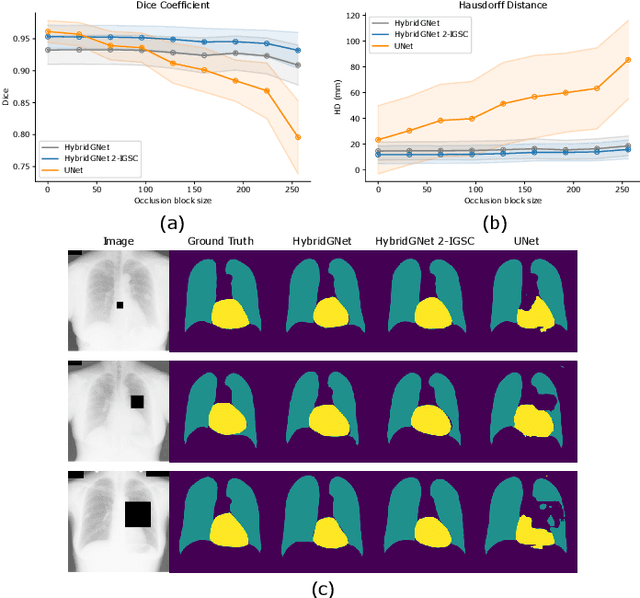

Abstract:Anatomical segmentation is a fundamental task in medical image computing, generally tackled with fully convolutional neural networks which produce dense segmentation masks. These models are often trained with loss functions such as cross-entropy or Dice, which assume pixels to be independent of each other, thus ignoring topological errors and anatomical inconsistencies. We address this limitation by moving from pixel-level to graph representations, which allow to naturally incorporate anatomical constraints by construction. To this end, we introduce HybridGNet, an encoder-decoder neural architecture that leverages standard convolutions for image feature encoding and graph convolutional neural networks (GCNNs) to decode plausible representations of anatomical structures. We also propose a novel image-to-graph skip connection layer which allows localized features to flow from standard convolutional blocks to GCNN blocks, and show that it improves segmentation accuracy. The proposed architecture is extensively evaluated in a variety of domain shift and image occlusion scenarios, and audited considering different types of demographic domain shift. Our comprehensive experimental setup compares HybridGNet with other landmark and pixel-based models for anatomical segmentation in chest x-ray images, and shows that it produces anatomically plausible results in challenging scenarios where other models tend to fail.
Domain Generalization via Gradient Surgery
Aug 03, 2021
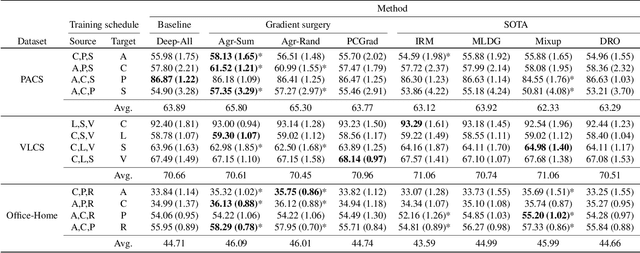

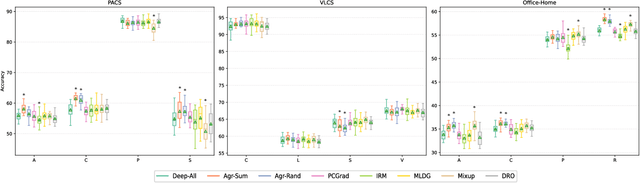
Abstract:In real-life applications, machine learning models often face scenarios where there is a change in data distribution between training and test domains. When the aim is to make predictions on distributions different from those seen at training, we incur in a domain generalization problem. Methods to address this issue learn a model using data from multiple source domains, and then apply this model to the unseen target domain. Our hypothesis is that when training with multiple domains, conflicting gradients within each mini-batch contain information specific to the individual domains which is irrelevant to the others, including the test domain. If left untouched, such disagreement may degrade generalization performance. In this work, we characterize the conflicting gradients emerging in domain shift scenarios and devise novel gradient agreement strategies based on gradient surgery to alleviate their effect. We validate our approach in image classification tasks with three multi-domain datasets, showing the value of the proposed agreement strategy in enhancing the generalization capability of deep learning models in domain shift scenarios.
Hybrid graph convolutional neural networks for landmark-based anatomical segmentation
Jun 17, 2021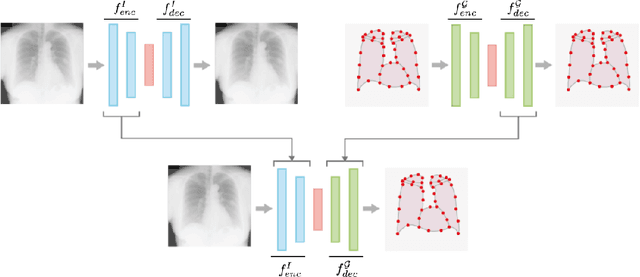
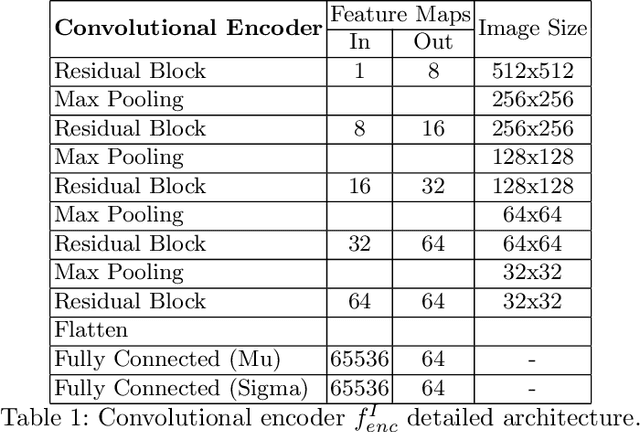
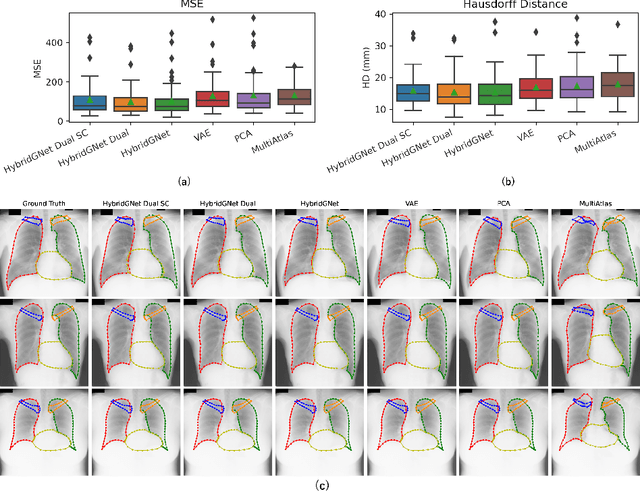

Abstract:In this work we address the problem of landmark-based segmentation for anatomical structures. We propose HybridGNet, an encoder-decoder neural architecture which combines standard convolutions for image feature encoding, with graph convolutional neural networks to decode plausible representations of anatomical structures. We benchmark the proposed architecture considering other standard landmark and pixel-based models for anatomical segmentation in chest x-ray images, and found that HybridGNet is more robust to image occlusions. We also show that it can be used to construct landmark-based segmentations from pixel level annotations. Our experimental results suggest that HybridGNet produces accurate and anatomically plausible landmark-based segmentations, by naturally incorporating shape constraints within the decoding process via spectral convolutions.
Learning Deformable Registration of Medical Images with Anatomical Constraints
Jan 22, 2020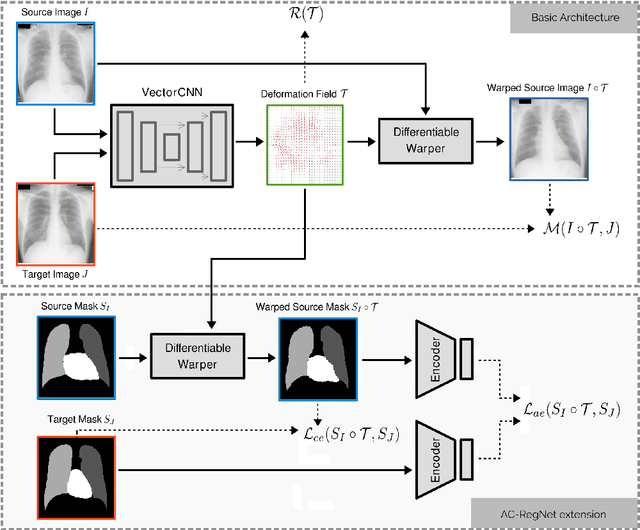
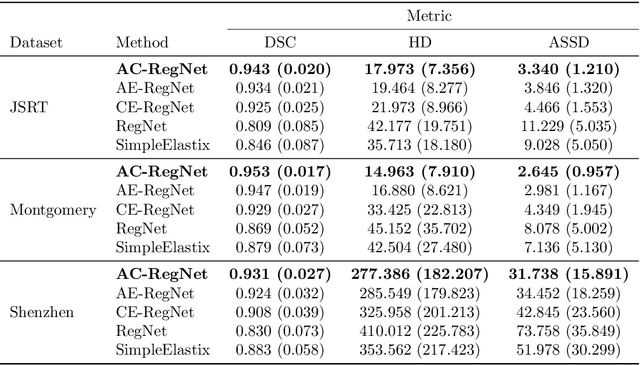
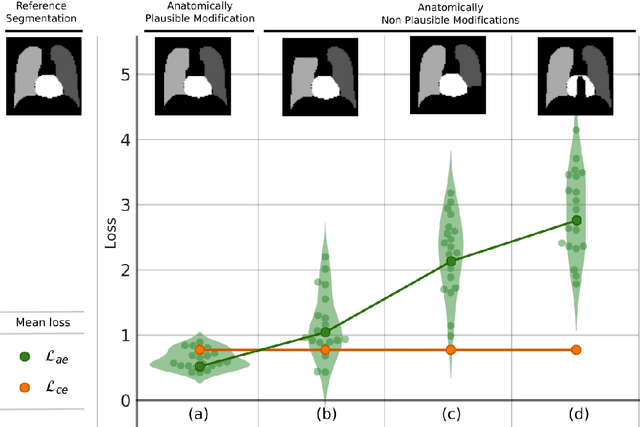
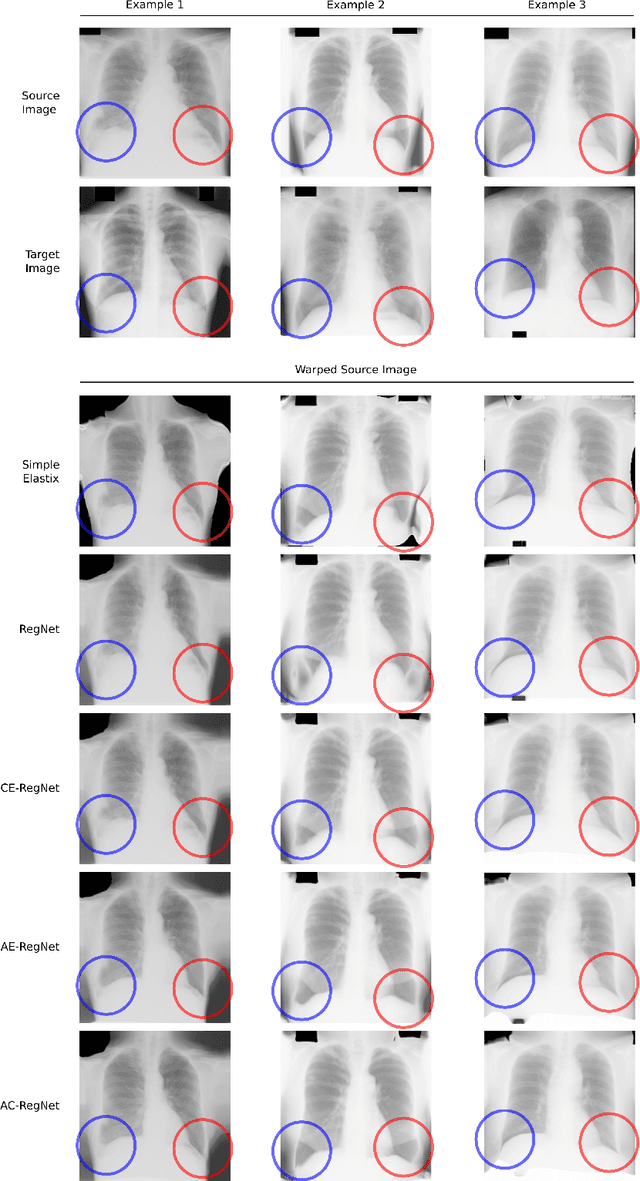
Abstract:Deformable image registration is a fundamental problem in the field of medical image analysis. During the last years, we have witnessed the advent of deep learning-based image registration methods which achieve state-of-the-art performance, and drastically reduce the required computational time. However, little work has been done regarding how can we encourage our models to produce not only accurate, but also anatomically plausible results, which is still an open question in the field. In this work, we argue that incorporating anatomical priors in the form of global constraints into the learning process of these models, will further improve their performance and boost the realism of the warped images after registration. We learn global non-linear representations of image anatomy using segmentation masks, and employ them to constraint the registration process. The proposed AC-RegNet architecture is evaluated in the context of chest X-ray image registration using three different datasets, where the high anatomical variability makes the task extremely challenging. Our experiments show that the proposed anatomically constrained registration model produces more realistic and accurate results than state-of-the-art methods, demonstrating the potential of this approach.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge