Kui Su
Mamba Learns in Context: Structure-Aware Domain Generalization for Multi-Task Point Cloud Understanding
Mar 21, 2026Abstract:While recent Transformer and Mamba architectures have advanced point cloud representation learning, they are typically developed for single-task or single-domain settings. Directly applying them to multi-task domain generalization (DG) leads to degraded performance. Transformers effectively model global dependencies but suffer from quadratic attention cost and lack explicit structural ordering, whereas Mamba offers linear-time recurrence yet often depends on coordinate-driven serialization, which is sensitive to viewpoint changes and missing regions, causing structural drift and unstable sequential modeling. In this paper, we propose Structure-Aware Domain Generalization (SADG), a Mamba-based In-Context Learning framework that preserves structural hierarchy across domains and tasks. We design structure-aware serialization (SAS) that generates transformation-invariant sequences using centroid-based topology and geodesic curvature continuity. We further devise hierarchical domain-aware modeling (HDM) that stabilizes cross-domain reasoning by consolidating intra-domain structure and fusing inter-domain relations. At test time, we introduce a lightweight spectral graph alignment (SGA) that shifts target features toward source prototypes in the spectral domain without updating model parameters, ensuring structure-preserving test-time feature shifting. In addition, we introduce MP3DObject, a real-scan object dataset for multi-task DG evaluation. Comprehensive experiments demonstrate that the proposed approach improves structural fidelity and consistently outperforms state-of-the-art methods across multiple tasks including reconstruction, denoising, and registration.
Tissue-Contrastive Semi-Masked Autoencoders for Segmentation Pretraining on Chest CT
Jul 12, 2024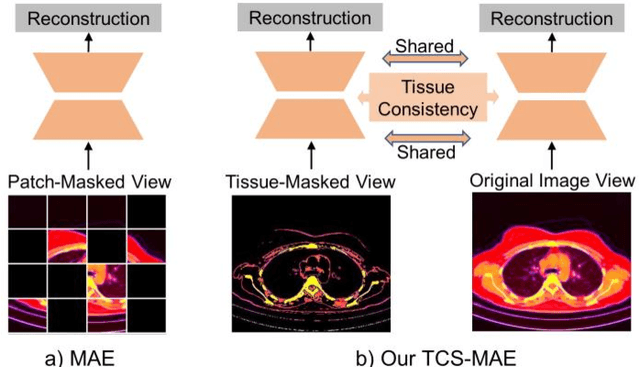
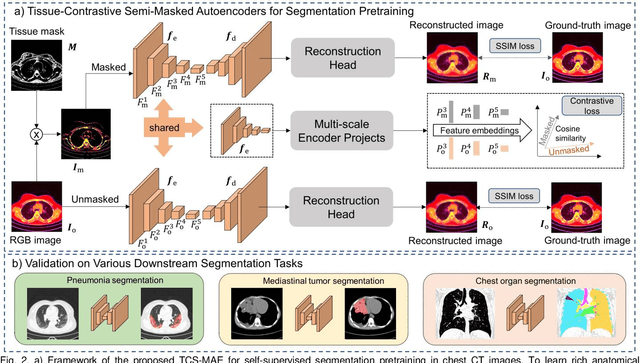
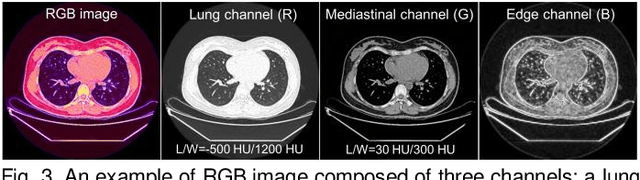
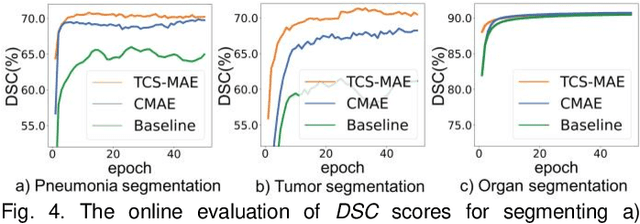
Abstract:Existing Masked Image Modeling (MIM) depends on a spatial patch-based masking-reconstruction strategy to perceive objects'features from unlabeled images, which may face two limitations when applied to chest CT: 1) inefficient feature learning due to complex anatomical details presented in CT images, and 2) suboptimal knowledge transfer owing to input disparity between upstream and downstream models. To address these issues, we propose a new MIM method named Tissue-Contrastive Semi-Masked Autoencoder (TCS-MAE) for modeling chest CT images. Our method has two novel designs: 1) a tissue-based masking-reconstruction strategy to capture more fine-grained anatomical features, and 2) a dual-AE architecture with contrastive learning between the masked and original image views to bridge the gap of the upstream and downstream models. To validate our method, we systematically investigate representative contrastive, generative, and hybrid self-supervised learning methods on top of tasks involving segmenting pneumonia, mediastinal tumors, and various organs. The results demonstrate that, compared to existing methods, our TCS-MAE more effectively learns tissue-aware representations, thereby significantly enhancing segmentation performance across all tasks.
Supervised Contrastive Learning for Fine-grained Chromosome Recognition
Dec 12, 2023Abstract:Chromosome recognition is an essential task in karyotyping, which plays a vital role in birth defect diagnosis and biomedical research. However, existing classification methods face significant challenges due to the inter-class similarity and intra-class variation of chromosomes. To address this issue, we propose a supervised contrastive learning strategy that is tailored to train model-agnostic deep networks for reliable chromosome classification. This method enables extracting fine-grained chromosomal embeddings in latent space. These embeddings effectively expand inter-class boundaries and reduce intra-class variations, enhancing their distinctiveness in predicting chromosome types. On top of two large-scale chromosome datasets, we comprehensively validate the power of our contrastive learning strategy in boosting cutting-edge deep networks such as Transformers and ResNets. Extensive results demonstrate that it can significantly improve models' generalization performance, with an accuracy improvement up to +4.5%. Codes and pretrained models will be released upon acceptance of this work.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge