Keith Cheng
Selective Synthetic Augmentation with HistoGAN for Improved Histopathology Image Classification
Nov 10, 2021
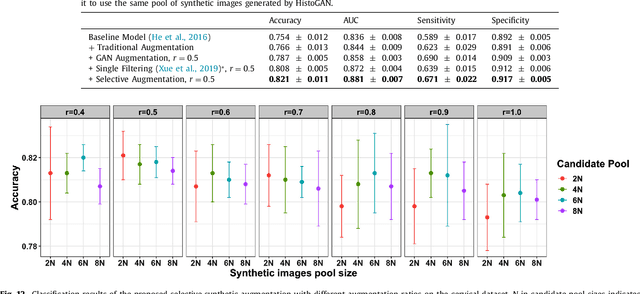

Abstract:Histopathological analysis is the present gold standard for precancerous lesion diagnosis. The goal of automated histopathological classification from digital images requires supervised training, which requires a large number of expert annotations that can be expensive and time-consuming to collect. Meanwhile, accurate classification of image patches cropped from whole-slide images is essential for standard sliding window based histopathology slide classification methods. To mitigate these issues, we propose a carefully designed conditional GAN model, namely HistoGAN, for synthesizing realistic histopathology image patches conditioned on class labels. We also investigate a novel synthetic augmentation framework that selectively adds new synthetic image patches generated by our proposed HistoGAN, rather than expanding directly the training set with synthetic images. By selecting synthetic images based on the confidence of their assigned labels and their feature similarity to real labeled images, our framework provides quality assurance to synthetic augmentation. Our models are evaluated on two datasets: a cervical histopathology image dataset with limited annotations, and another dataset of lymph node histopathology images with metastatic cancer. Here, we show that leveraging HistoGAN generated images with selective augmentation results in significant and consistent improvements of classification performance (6.7% and 2.8% higher accuracy, respectively) for cervical histopathology and metastatic cancer datasets.
* Elsevier Medical Image Analysis Best Paper Award runner up. arXiv admin note: substantial text overlap with arXiv:1912.03837
A Multi-attribute Controllable Generative Model for Histopathology Image Synthesis
Nov 10, 2021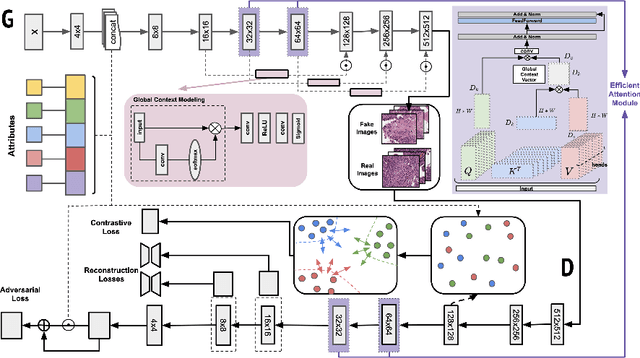
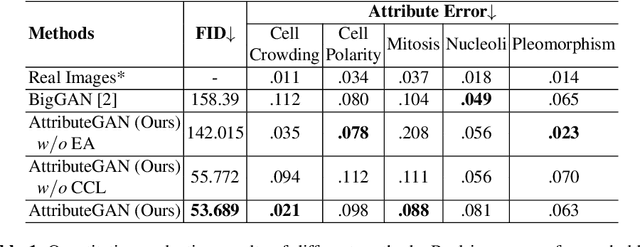
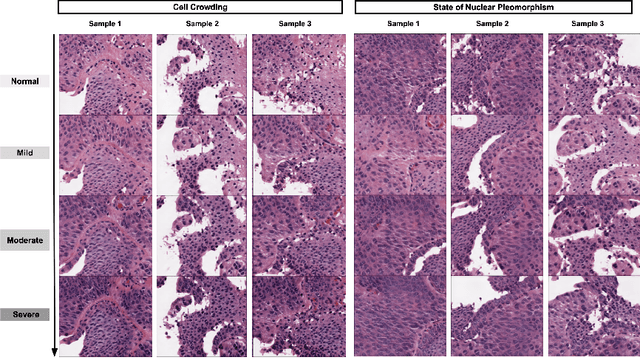
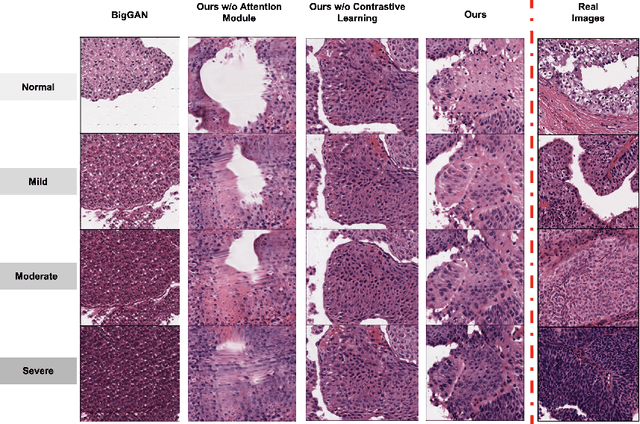
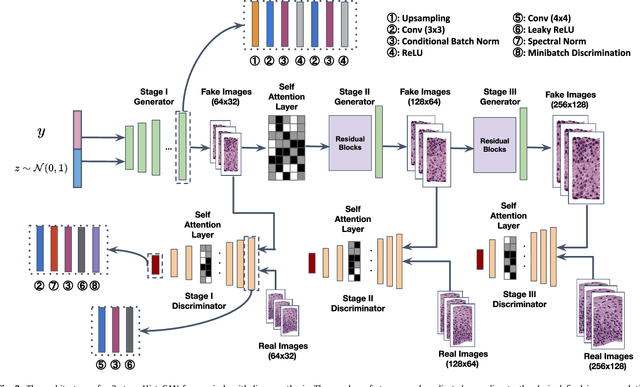
Abstract:Generative models have been applied in the medical imaging domain for various image recognition and synthesis tasks. However, a more controllable and interpretable image synthesis model is still lacking yet necessary for important applications such as assisting in medical training. In this work, we leverage the efficient self-attention and contrastive learning modules and build upon state-of-the-art generative adversarial networks (GANs) to achieve an attribute-aware image synthesis model, termed AttributeGAN, which can generate high-quality histopathology images based on multi-attribute inputs. In comparison to existing single-attribute conditional generative models, our proposed model better reflects input attributes and enables smoother interpolation among attribute values. We conduct experiments on a histopathology dataset containing stained H&E images of urothelial carcinoma and demonstrate the effectiveness of our proposed model via comprehensive quantitative and qualitative comparisons with state-of-the-art models as well as different variants of our model. Code is available at https://github.com/karenyyy/MICCAI2021AttributeGAN.
Synthetic Sample Selection via Reinforcement Learning
Aug 26, 2020



Abstract:Synthesizing realistic medical images provides a feasible solution to the shortage of training data in deep learning based medical image recognition systems. However, the quality control of synthetic images for data augmentation purposes is under-investigated, and some of the generated images are not realistic and may contain misleading features that distort data distribution when mixed with real images. Thus, the effectiveness of those synthetic images in medical image recognition systems cannot be guaranteed when they are being added randomly without quality assurance. In this work, we propose a reinforcement learning (RL) based synthetic sample selection method that learns to choose synthetic images containing reliable and informative features. A transformer based controller is trained via proximal policy optimization (PPO) using the validation classification accuracy as the reward. The selected images are mixed with the original training data for improved training of image recognition systems. To validate our method, we take the pathology image recognition as an example and conduct extensive experiments on two histopathology image datasets. In experiments on a cervical dataset and a lymph node dataset, the image classification performance is improved by 8.1% and 2.3%, respectively, when utilizing high-quality synthetic images selected by our RL framework. Our proposed synthetic sample selection method is general and has great potential to boost the performance of various medical image recognition systems given limited annotation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge