Keelan O'Riordan
Deep Learning-based Detection of Bacterial Swarm Motion Using a Single Image
Oct 19, 2024
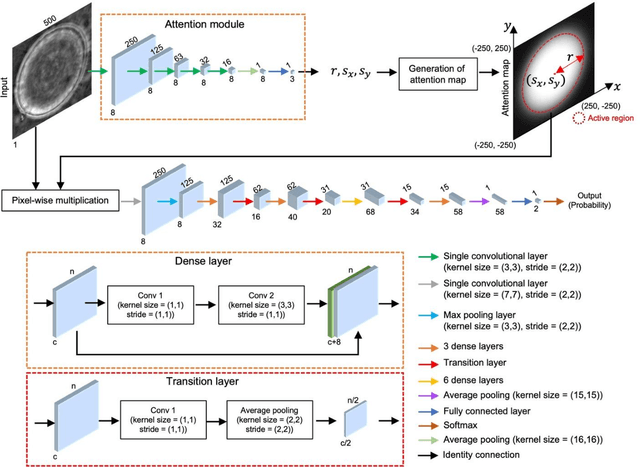
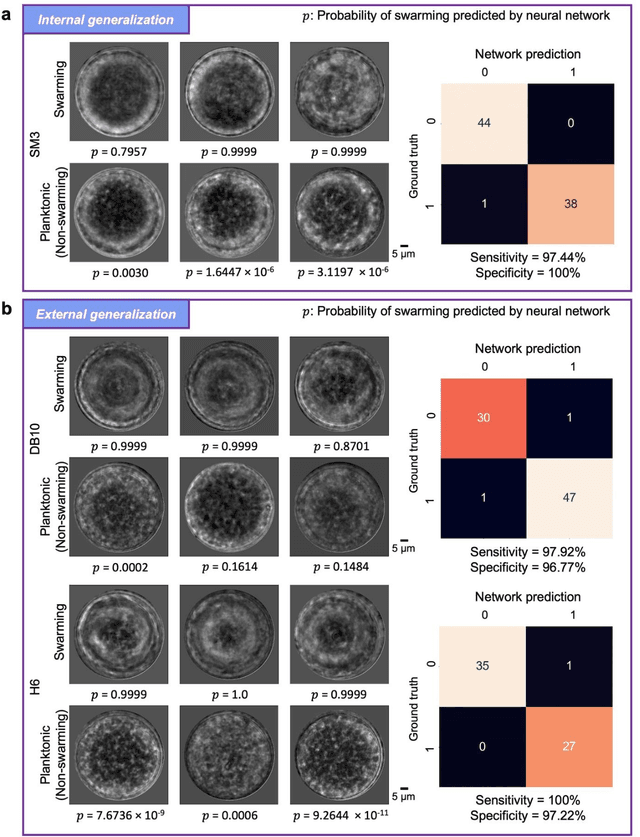
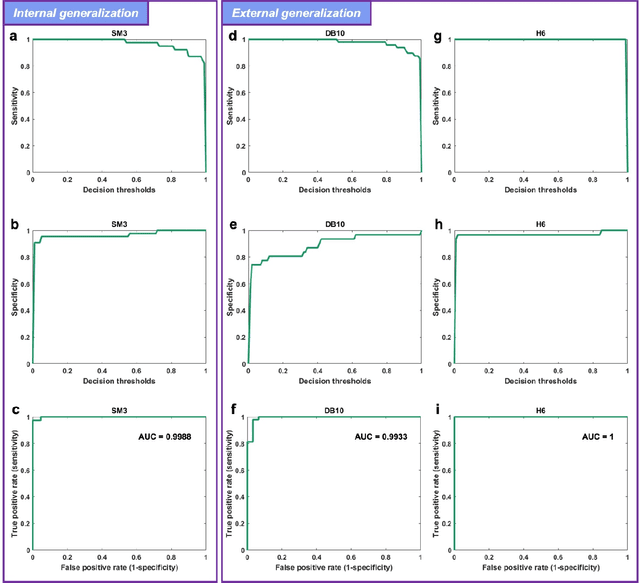
Abstract:Distinguishing between swarming and swimming, the two principal forms of bacterial movement, holds significant conceptual and clinical relevance. This is because bacteria that exhibit swarming capabilities often possess unique properties crucial to the pathogenesis of infectious diseases and may also have therapeutic potential. Here, we report a deep learning-based swarming classifier that rapidly and autonomously predicts swarming probability using a single blurry image. Compared with traditional video-based, manually-processed approaches, our method is particularly suited for high-throughput environments and provides objective, quantitative assessments of swarming probability. The swarming classifier demonstrated in our work was trained on Enterobacter sp. SM3 and showed good performance when blindly tested on new swarming (positive) and swimming (negative) test images of SM3, achieving a sensitivity of 97.44% and a specificity of 100%. Furthermore, this classifier demonstrated robust external generalization capabilities when applied to unseen bacterial species, such as Serratia marcescens DB10 and Citrobacter koseri H6. It blindly achieved a sensitivity of 97.92% and a specificity of 96.77% for DB10, and a sensitivity of 100% and a specificity of 97.22% for H6. This competitive performance indicates the potential to adapt our approach for diagnostic applications through portable devices or even smartphones. This adaptation would facilitate rapid, objective, on-site screening for bacterial swarming motility, potentially enhancing the early detection and treatment assessment of various diseases, including inflammatory bowel diseases (IBD) and urinary tract infections (UTI).
Deep Learning-enabled Detection and Classification of Bacterial Colonies using a Thin Film Transistor (TFT) Image Sensor
May 07, 2022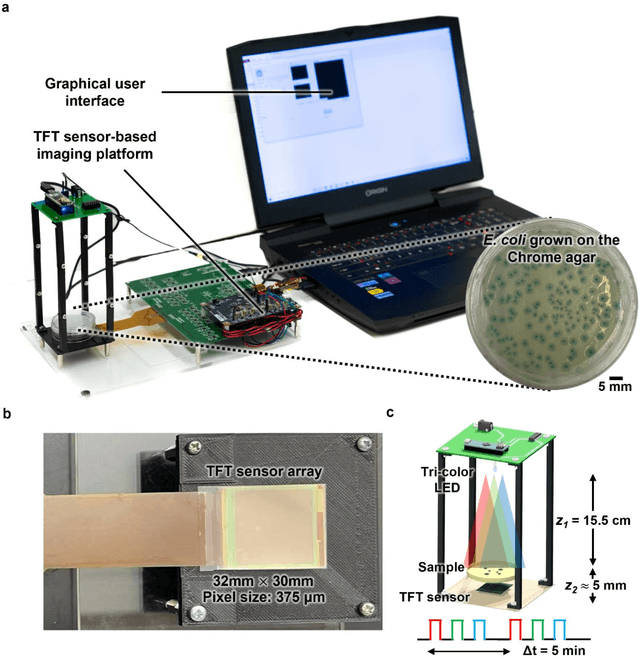
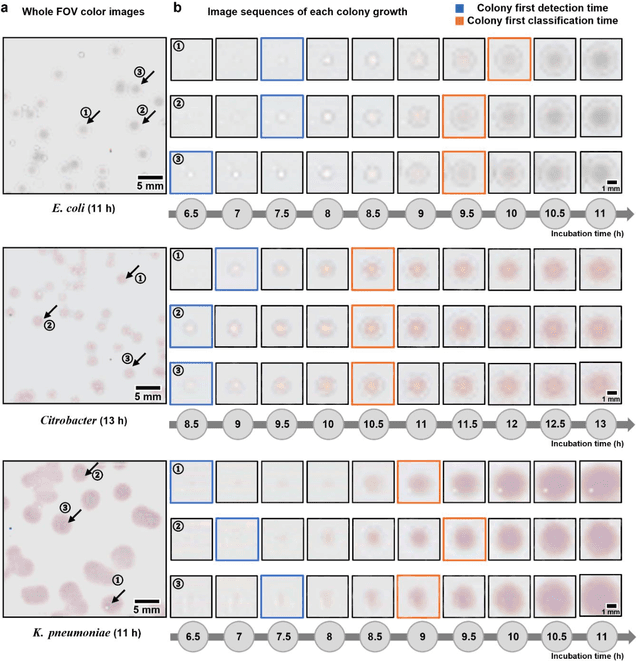
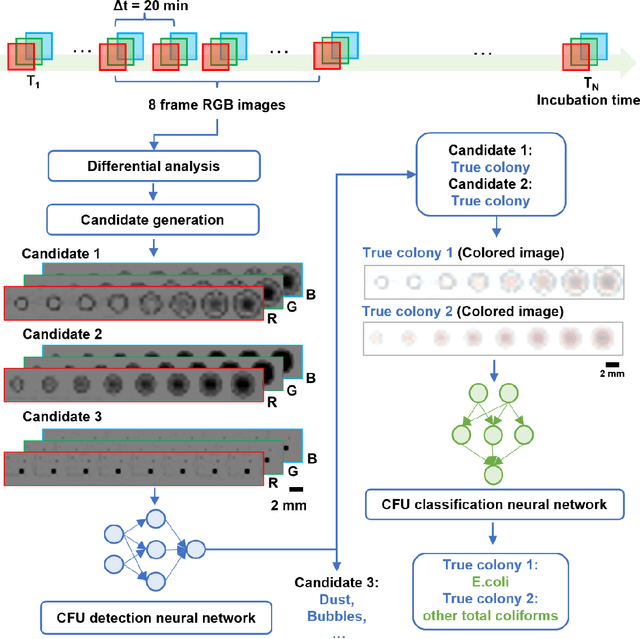

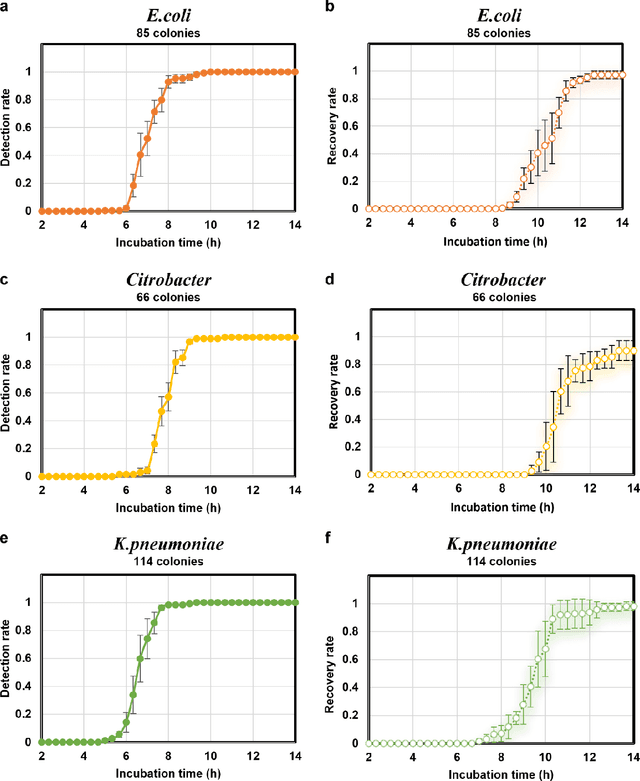
Abstract:Early detection and identification of pathogenic bacteria such as Escherichia coli (E. coli) is an essential task for public health. The conventional culture-based methods for bacterial colony detection usually take >24 hours to get the final read-out. Here, we demonstrate a bacterial colony-forming-unit (CFU) detection system exploiting a thin-film-transistor (TFT)-based image sensor array that saves ~12 hours compared to the Environmental Protection Agency (EPA)-approved methods. To demonstrate the efficacy of this CFU detection system, a lensfree imaging modality was built using the TFT image sensor with a sample field-of-view of ~10 cm^2. Time-lapse images of bacterial colonies cultured on chromogenic agar plates were automatically collected at 5-minute intervals. Two deep neural networks were used to detect and count the growing colonies and identify their species. When blindly tested with 265 colonies of E. coli and other coliform bacteria (i.e., Citrobacter and Klebsiella pneumoniae), our system reached an average CFU detection rate of 97.3% at 9 hours of incubation and an average recovery rate of 91.6% at ~12 hours. This TFT-based sensor can be applied to various microbiological detection methods. Due to the large scalability, ultra-large field-of-view, and low cost of the TFT-based image sensors, this platform can be integrated with each agar plate to be tested and disposed of after the automated CFU count. The imaging field-of-view of this platform can be cost-effectively increased to >100 cm^2 to provide a massive throughput for CFU detection using, e.g., roll-to-roll manufacturing of TFTs as used in the flexible display industry.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge