Kasra Borazjani
Bringing Multi-Modal Multi-Task Federated Foundation Models to Education Domain: Prospects and Challenges
Sep 09, 2025Abstract:Multi-modal multi-task (M3T) foundation models (FMs) have recently shown transformative potential in artificial intelligence, with emerging applications in education. However, their deployment in real-world educational settings is hindered by privacy regulations, data silos, and limited domain-specific data availability. We introduce M3T Federated Foundation Models (FedFMs) for education: a paradigm that integrates federated learning (FL) with M3T FMs to enable collaborative, privacy-preserving training across decentralized institutions while accommodating diverse modalities and tasks. Subsequently, this position paper aims to unveil M3T FedFMs as a promising yet underexplored approach to the education community, explore its potentials, and reveal its related future research directions. We outline how M3T FedFMs can advance three critical pillars of next-generation intelligent education systems: (i) privacy preservation, by keeping sensitive multi-modal student and institutional data local; (ii) personalization, through modular architectures enabling tailored models for students, instructors, and institutions; and (iii) equity and inclusivity, by facilitating participation from underrepresented and resource-constrained entities. We finally identify various open research challenges, including studying of (i) inter-institution heterogeneous privacy regulations, (ii) the non-uniformity of data modalities' characteristics, (iii) the unlearning approaches for M3T FedFMs, (iv) the continual learning frameworks for M3T FedFMs, and (v) M3T FedFM model interpretability, which must be collectively addressed for practical deployment.
Hierarchical Federated Foundation Models over Wireless Networks for Multi-Modal Multi-Task Intelligence: Integration of Edge Learning with D2D/P2P-Enabled Fog Learning Architectures
Sep 03, 2025


Abstract:The rise of foundation models (FMs) has reshaped the landscape of machine learning. As these models continued to grow, leveraging geo-distributed data from wireless devices has become increasingly critical, giving rise to federated foundation models (FFMs). More recently, FMs have evolved into multi-modal multi-task (M3T) FMs (e.g., GPT-4) capable of processing diverse modalities across multiple tasks, which motivates a new underexplored paradigm: M3T FFMs. In this paper, we unveil an unexplored variation of M3T FFMs by proposing hierarchical federated foundation models (HF-FMs), which in turn expose two overlooked heterogeneity dimensions to fog/edge networks that have a direct impact on these emerging models: (i) heterogeneity in collected modalities and (ii) heterogeneity in executed tasks across fog/edge nodes. HF-FMs strategically align the modular structure of M3T FMs, comprising modality encoders, prompts, mixture-of-experts (MoEs), adapters, and task heads, with the hierarchical nature of fog/edge infrastructures. Moreover, HF-FMs enable the optional usage of device-to-device (D2D) communications, enabling horizontal module relaying and localized cooperative training among nodes when feasible. Through delving into the architectural design of HF-FMs, we highlight their unique capabilities along with a series of tailored future research directions. Finally, to demonstrate their potential, we prototype HF-FMs in a wireless network setting and release the open-source code for the development of HF-FMs with the goal of fostering exploration in this untapped field (GitHub: https://github.com/payamsiabd/M3T-FFM).
Multi-Modal Multi-Task (M3T) Federated Foundation Models for Embodied AI: Potentials and Challenges for Edge Integration
May 16, 2025Abstract:As embodied AI systems become increasingly multi-modal, personalized, and interactive, they must learn effectively from diverse sensory inputs, adapt continually to user preferences, and operate safely under resource and privacy constraints. These challenges expose a pressing need for machine learning models capable of swift, context-aware adaptation while balancing model generalization and personalization. Here, two methods emerge as suitable candidates, each offering parts of these capabilities: Foundation Models (FMs) provide a pathway toward generalization across tasks and modalities, whereas Federated Learning (FL) offers the infrastructure for distributed, privacy-preserving model updates and user-level model personalization. However, when used in isolation, each of these approaches falls short of meeting the complex and diverse capability requirements of real-world embodied environments. In this vision paper, we introduce Federated Foundation Models (FFMs) for embodied AI, a new paradigm that unifies the strengths of multi-modal multi-task (M3T) FMs with the privacy-preserving distributed nature of FL, enabling intelligent systems at the wireless edge. We collect critical deployment dimensions of FFMs in embodied AI ecosystems under a unified framework, which we name "EMBODY": Embodiment heterogeneity, Modality richness and imbalance, Bandwidth and compute constraints, On-device continual learning, Distributed control and autonomy, and Yielding safety, privacy, and personalization. For each, we identify concrete challenges and envision actionable research directions. We also present an evaluation framework for deploying FFMs in embodied AI systems, along with the associated trade-offs.
Redefining non-IID Data in Federated Learning for Computer Vision Tasks: Migrating from Labels to Embeddings for Task-Specific Data Distributions
Mar 17, 2025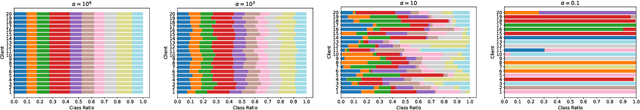
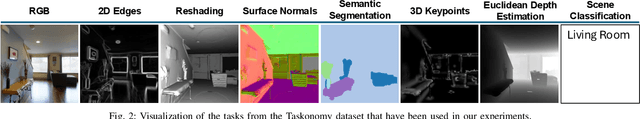

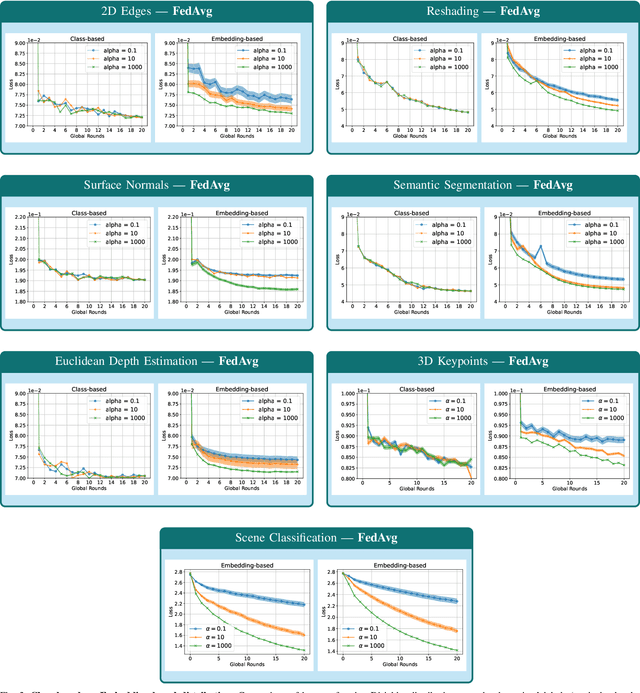
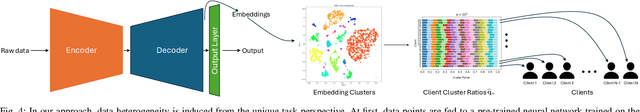
Abstract:Federated Learning (FL) represents a paradigm shift in distributed machine learning (ML), enabling clients to train models collaboratively while keeping their raw data private. This paradigm shift from traditional centralized ML introduces challenges due to the non-iid (non-independent and identically distributed) nature of data across clients, significantly impacting FL's performance. Existing literature, predominantly model data heterogeneity by imposing label distribution skew across clients. In this paper, we show that label distribution skew fails to fully capture the real-world data heterogeneity among clients in computer vision tasks beyond classification. Subsequently, we demonstrate that current approaches overestimate FL's performance by relying on label/class distribution skew, exposing an overlooked gap in the literature. By utilizing pre-trained deep neural networks to extract task-specific data embeddings, we define task-specific data heterogeneity through the lens of each vision task and introduce a new level of data heterogeneity called embedding-based data heterogeneity. Our methodology involves clustering data points based on embeddings and distributing them among clients using the Dirichlet distribution. Through extensive experiments, we evaluate the performance of different FL methods under our revamped notion of data heterogeneity, introducing new benchmark performance measures to the literature. We further unveil a series of open research directions that can be pursued.
Multi-Modal Federated Learning for Cancer Staging over Non-IID Datasets with Unbalanced Modalities
Jan 07, 2024Abstract:The use of machine learning (ML) for cancer staging through medical image analysis has gained substantial interest across medical disciplines. When accompanied by the innovative federated learning (FL) framework, ML techniques can further overcome privacy concerns related to patient data exposure. Given the frequent presence of diverse data modalities within patient records, leveraging FL in a multi-modal learning framework holds considerable promise for cancer staging. However, existing works on multi-modal FL often presume that all data-collecting institutions have access to all data modalities. This oversimplified approach neglects institutions that have access to only a portion of data modalities within the system. In this work, we introduce a novel FL architecture designed to accommodate not only the heterogeneity of data samples, but also the inherent heterogeneity/non-uniformity of data modalities across institutions. We shed light on the challenges associated with varying convergence speeds observed across different data modalities within our FL system. Subsequently, we propose a solution to tackle these challenges by devising a distributed gradient blending and proximity-aware client weighting strategy tailored for multi-modal FL. To show the superiority of our method, we conduct experiments using The Cancer Genome Atlas program (TCGA) datalake considering different cancer types and three modalities of data: mRNA sequences, histopathological image data, and clinical information.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge