Karin Stacke
Correlated Feature Aggregation by Region Helps Distinguish Aggressive from Indolent Clear Cell Renal Cell Carcinoma Subtypes on CT
Sep 29, 2022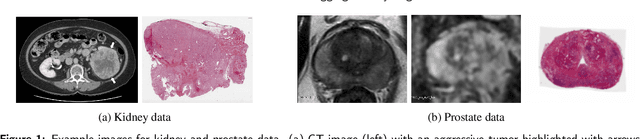
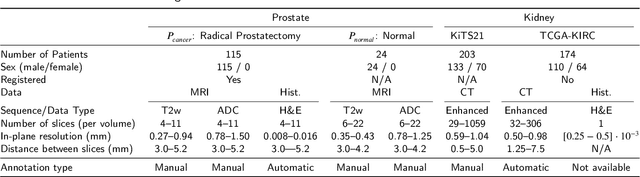
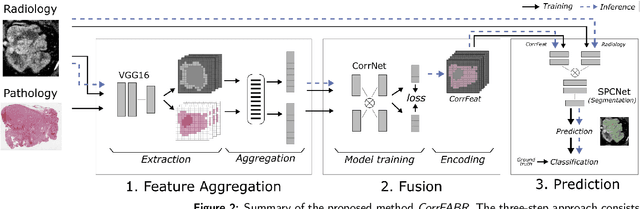

Abstract:Renal cell carcinoma (RCC) is a common cancer that varies in clinical behavior. Indolent RCC is often low-grade without necrosis and can be monitored without treatment. Aggressive RCC is often high-grade and can cause metastasis and death if not promptly detected and treated. While most kidney cancers are detected on CT scans, grading is based on histology from invasive biopsy or surgery. Determining aggressiveness on CT images is clinically important as it facilitates risk stratification and treatment planning. This study aims to use machine learning methods to identify radiology features that correlate with features on pathology to facilitate assessment of cancer aggressiveness on CT images instead of histology. This paper presents a novel automated method, Correlated Feature Aggregation By Region (CorrFABR), for classifying aggressiveness of clear cell RCC by leveraging correlations between radiology and corresponding unaligned pathology images. CorrFABR consists of three main steps: (1) Feature Aggregation where region-level features are extracted from radiology and pathology images, (2) Fusion where radiology features correlated with pathology features are learned on a region level, and (3) Prediction where the learned correlated features are used to distinguish aggressive from indolent clear cell RCC using CT alone as input. Thus, during training, CorrFABR learns from both radiology and pathology images, but during inference, CorrFABR will distinguish aggressive from indolent clear cell RCC using CT alone, in the absence of pathology images. CorrFABR improved classification performance over radiology features alone, with an increase in binary classification F1-score from 0.68 (0.04) to 0.73 (0.03). This demonstrates the potential of incorporating pathology disease characteristics for improved classification of aggressiveness of clear cell RCC on CT images.
Learning Representations with Contrastive Self-Supervised Learning for Histopathology Applications
Dec 10, 2021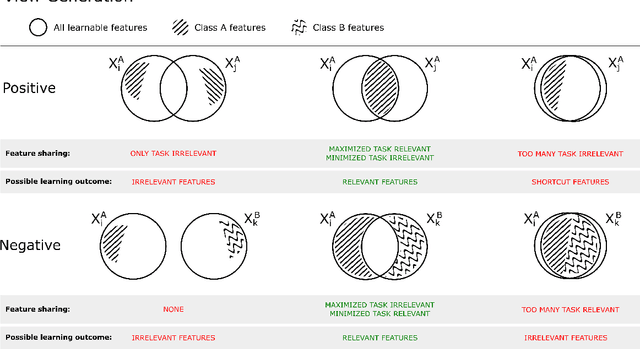
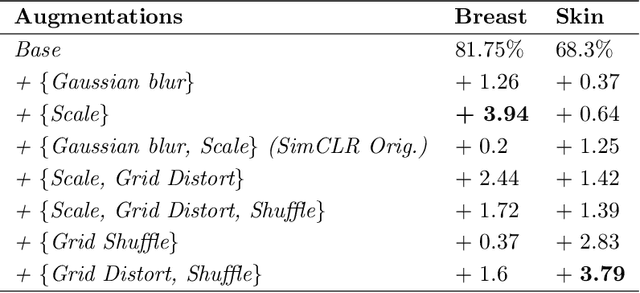
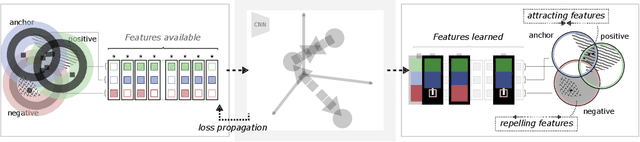
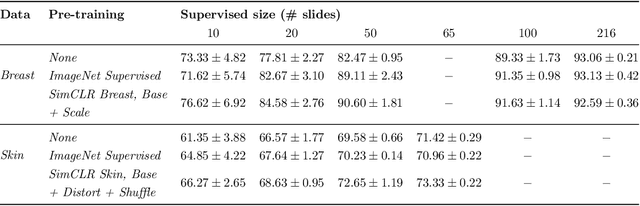
Abstract:Unsupervised learning has made substantial progress over the last few years, especially by means of contrastive self-supervised learning. The dominating dataset for benchmarking self-supervised learning has been ImageNet, for which recent methods are approaching the performance achieved by fully supervised training. The ImageNet dataset is however largely object-centric, and it is not clear yet what potential those methods have on widely different datasets and tasks that are not object-centric, such as in digital pathology. While self-supervised learning has started to be explored within this area with encouraging results, there is reason to look closer at how this setting differs from natural images and ImageNet. In this paper we make an in-depth analysis of contrastive learning for histopathology, pin-pointing how the contrastive objective will behave differently due to the characteristics of histopathology data. We bring forward a number of considerations, such as view generation for the contrastive objective and hyper-parameter tuning. In a large battery of experiments, we analyze how the downstream performance in tissue classification will be affected by these considerations. The results point to how contrastive learning can reduce the annotation effort within digital pathology, but that the specific dataset characteristics need to be considered. To take full advantage of the contrastive learning objective, different calibrations of view generation and hyper-parameters are required. Our results pave the way for realizing the full potential of self-supervised learning for histopathology applications.
Primary Tumor and Inter-Organ Augmentations for Supervised Lymph Node Colon Adenocarcinoma Metastasis Detection
Sep 17, 2021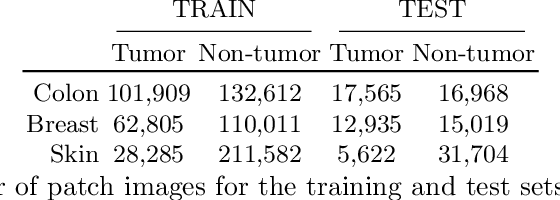
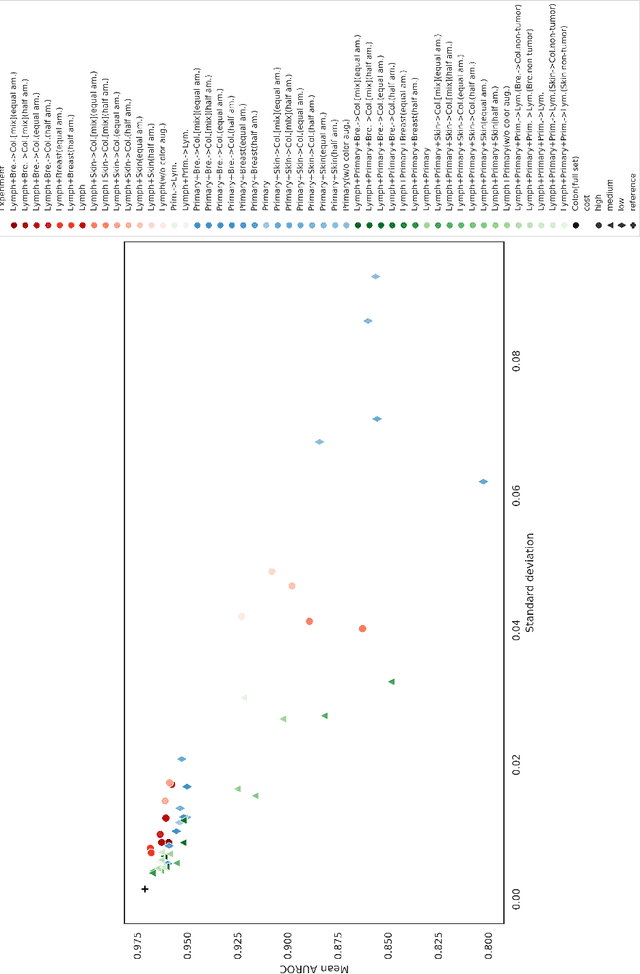
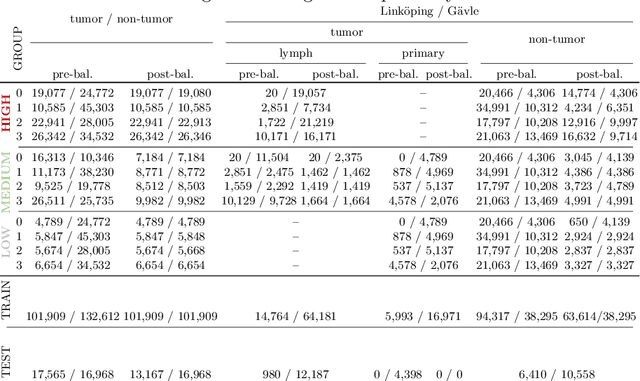
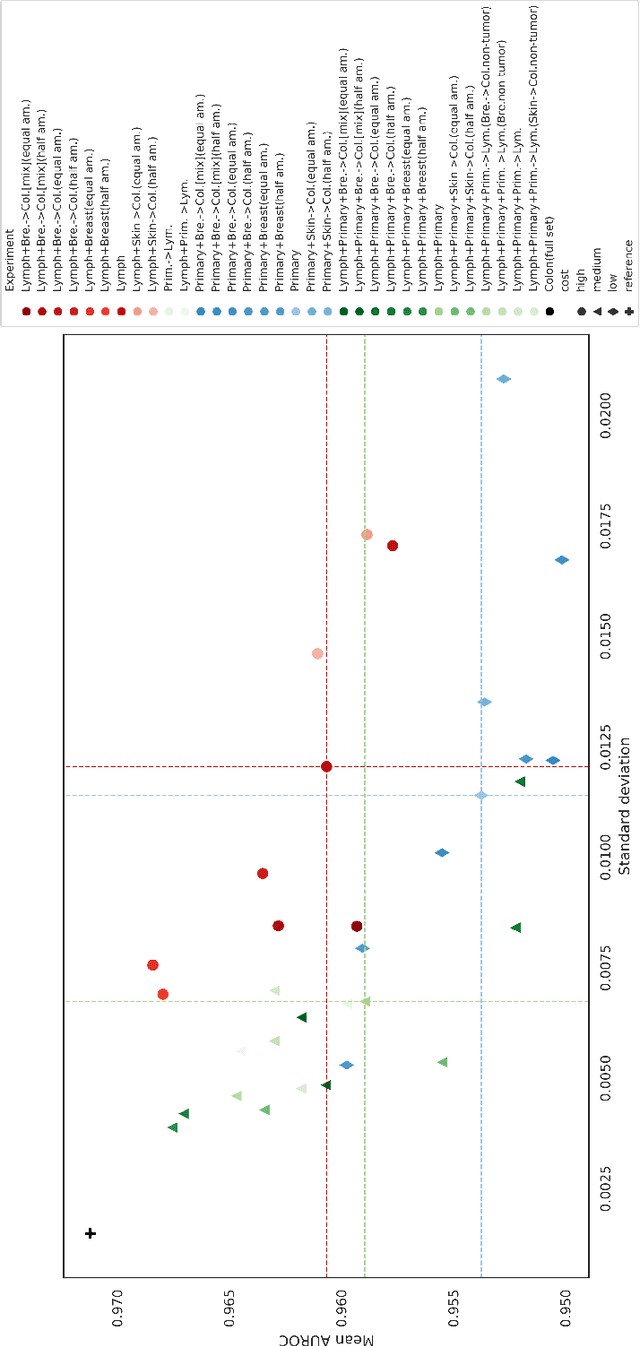
Abstract:The scarcity of labeled data is a major bottleneck for developing accurate and robust deep learning-based models for histopathology applications. The problem is notably prominent for the task of metastasis detection in lymph nodes, due to the tissue's low tumor-to-non-tumor ratio, resulting in labor- and time-intensive annotation processes for the pathologists. This work explores alternatives on how to augment the training data for colon carcinoma metastasis detection when there is limited or no representation of the target domain. Through an exhaustive study of cross-validated experiments with limited training data availability, we evaluate both an inter-organ approach utilizing already available data for other tissues, and an intra-organ approach, utilizing the primary tumor. Both these approaches result in little to no extra annotation effort. Our results show that these data augmentation strategies can be an efficient way of increasing accuracy on metastasis detection, but fore-most increase robustness.
A Study of Deep Learning Colon Cancer Detection in Limited Data Access Scenarios
May 22, 2020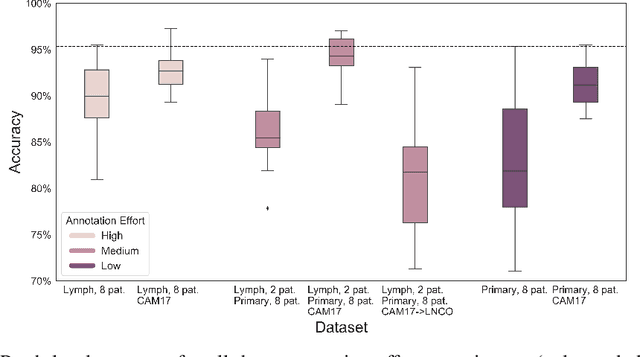
Abstract:Digitization of histopathology slides has led to several advances, from easy data sharing and collaborations to the development of digital diagnostic tools. Deep learning (DL) methods for classification and detection have shown great potential, but often require large amounts of training data that are hard to collect, and annotate. For many cancer types, the scarceness of data creates barriers for training DL models. One such scenario relates to detecting tumor metastasis in lymph node tissue, where the low ratio of tumor to non-tumor cells makes the diagnostic task hard and time-consuming. DL-based tools can allow faster diagnosis, with potentially increased quality. Unfortunately, due to the sparsity of tumor cells, annotating this type of data demands a high level of effort from pathologists. Using weak annotations from slide-level images have shown great potential, but demand access to a substantial amount of data as well. In this study, we investigate mitigation strategies for limited data access scenarios. Particularly, we address whether it is possible to exploit mutual structure between tissues to develop general techniques, wherein data from one type of cancer in a particular tissue could have diagnostic value for other cancers in other tissues. Our case is exemplified by a DL model for metastatic colon cancer detection in lymph nodes. Could such a model be trained with little or even no lymph node data? As alternative data sources, we investigate 1) tumor cells taken from the primary colon tumor tissue, and 2) cancer data from a different organ (breast), either as is or transformed to the target domain (colon) using Cycle-GANs. We show that the suggested approaches make it possible to detect cancer metastasis with no or very little lymph node data, opening up for the possibility that existing, annotated histopathology data could generalize to other domains.
A Closer Look at Domain Shift for Deep Learning in Histopathology
Sep 26, 2019

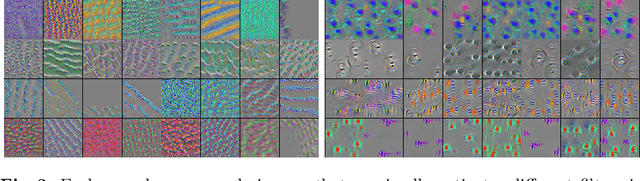
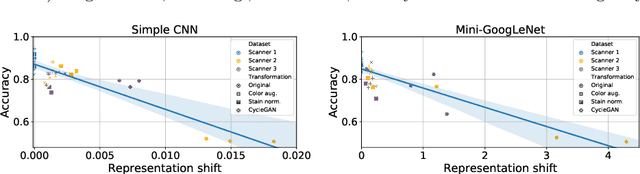
Abstract:Domain shift is a significant problem in histopathology. There can be large differences in data characteristics of whole-slide images between medical centers and scanners, making generalization of deep learning to unseen data difficult. To gain a better understanding of the problem, we present a study on convolutional neural networks trained for tumor classification of H&E stained whole-slide images. We analyze how augmentation and normalization strategies affect performance and learned representations, and what features a trained model respond to. Most centrally, we present a novel measure for evaluating the distance between domains in the context of the learned representation of a particular model. This measure can reveal how sensitive a model is to domain variations, and can be used to detect new data that a model will have problems generalizing to. The results show how learning is heavily influenced by the preparation of training data, and that the latent representation used to do classification is sensitive to changes in data distribution, especially when training without augmentation or normalization.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge