Kallil M. Zielinski
VORTEX: Challenging CNNs at Texture Recognition by using Vision Transformers with Orderless and Randomized Token Encodings
Mar 09, 2025



Abstract:Texture recognition has recently been dominated by ImageNet-pre-trained deep Convolutional Neural Networks (CNNs), with specialized modifications and feature engineering required to achieve state-of-the-art (SOTA) performance. However, although Vision Transformers (ViTs) were introduced a few years ago, little is known about their texture recognition ability. Therefore, in this work, we introduce VORTEX (ViTs with Orderless and Randomized Token Encodings for Texture Recognition), a novel method that enables the effective use of ViTs for texture analysis. VORTEX extracts multi-depth token embeddings from pre-trained ViT backbones and employs a lightweight module to aggregate hierarchical features and perform orderless encoding, obtaining a better image representation for texture recognition tasks. This approach allows seamless integration with any ViT with the common transformer architecture. Moreover, no fine-tuning of the backbone is performed, since they are used only as frozen feature extractors, and the features are fed to a linear SVM. We evaluate VORTEX on nine diverse texture datasets, demonstrating its ability to achieve or surpass SOTA performance in a variety of texture analysis scenarios. By bridging the gap between texture recognition with CNNs and transformer-based architectures, VORTEX paves the way for adopting emerging transformer foundation models. Furthermore, VORTEX demonstrates robust computational efficiency when coupled with ViT backbones compared to CNNs with similar costs. The method implementation and experimental scripts are publicly available in our online repository.
A Comparative Survey of Vision Transformers for Feature Extraction in Texture Analysis
Jun 10, 2024



Abstract:Texture, a significant visual attribute in images, has been extensively investigated across various image recognition applications. Convolutional Neural Networks (CNNs), which have been successful in many computer vision tasks, are currently among the best texture analysis approaches. On the other hand, Vision Transformers (ViTs) have been surpassing the performance of CNNs on tasks such as object recognition, causing a paradigm shift in the field. However, ViTs have so far not been scrutinized for texture recognition, hindering a proper appreciation of their potential in this specific setting. For this reason, this work explores various pre-trained ViT architectures when transferred to tasks that rely on textures. We review 21 different ViT variants and perform an extensive evaluation and comparison with CNNs and hand-engineered models on several tasks, such as assessing robustness to changes in texture rotation, scale, and illumination, and distinguishing color textures, material textures, and texture attributes. The goal is to understand the potential and differences among these models when directly applied to texture recognition, using pre-trained ViTs primarily for feature extraction and employing linear classifiers for evaluation. We also evaluate their efficiency, which is one of the main drawbacks in contrast to other methods. Our results show that ViTs generally outperform both CNNs and hand-engineered models, especially when using stronger pre-training and tasks involving in-the-wild textures (images from the internet). We highlight the following promising models: ViT-B with DINO pre-training, BeiTv2, and the Swin architecture, as well as the EfficientFormer as a low-cost alternative. In terms of efficiency, although having a higher number of GFLOPs and parameters, ViT-B and BeiT(v2) can achieve a lower feature extraction time on GPUs compared to ResNet50.
Advanced wood species identification based on multiple anatomical sections and using deep feature transfer and fusion
Apr 12, 2024Abstract:In recent years, we have seen many advancements in wood species identification. Methods like DNA analysis, Near Infrared (NIR) spectroscopy, and Direct Analysis in Real Time (DART) mass spectrometry complement the long-established wood anatomical assessment of cell and tissue morphology. However, most of these methods have some limitations such as high costs, the need for skilled experts for data interpretation, and the lack of good datasets for professional reference. Therefore, most of these methods, and certainly the wood anatomical assessment, may benefit from tools based on Artificial Intelligence. In this paper, we apply two transfer learning techniques with Convolutional Neural Networks (CNNs) to a multi-view Congolese wood species dataset including sections from different orientations and viewed at different microscopic magnifications. We explore two feature extraction methods in detail, namely Global Average Pooling (GAP) and Random Encoding of Aggregated Deep Activation Maps (RADAM), for efficient and accurate wood species identification. Our results indicate superior accuracy on diverse datasets and anatomical sections, surpassing the results of other methods. Our proposal represents a significant advancement in wood species identification, offering a robust tool to support the conservation of forest ecosystems and promote sustainable forestry practices.
RADAM: Texture Recognition through Randomized Aggregated Encoding of Deep Activation Maps
Mar 08, 2023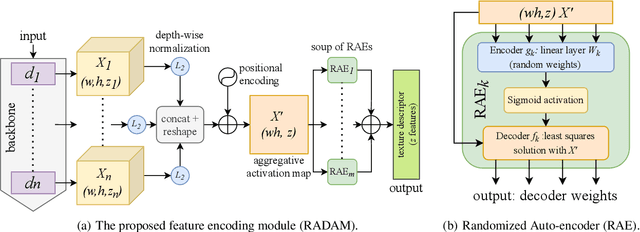
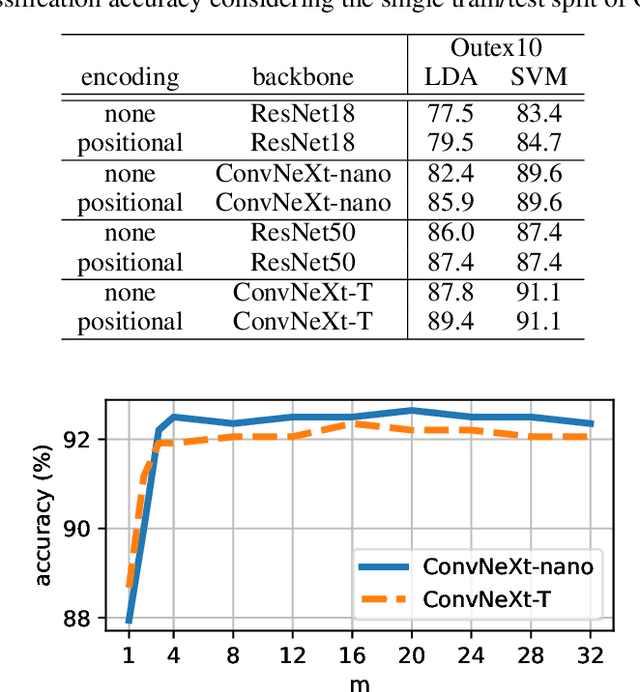
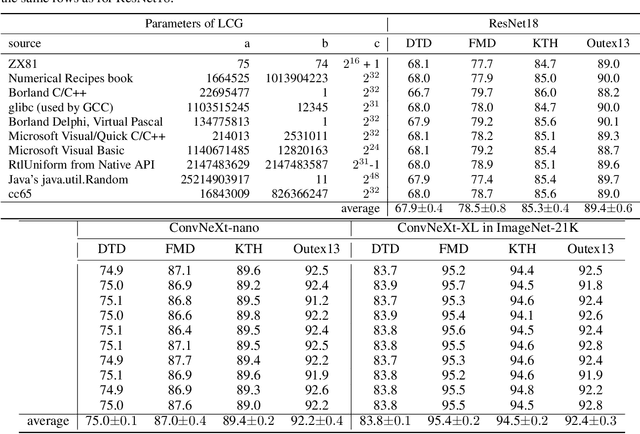
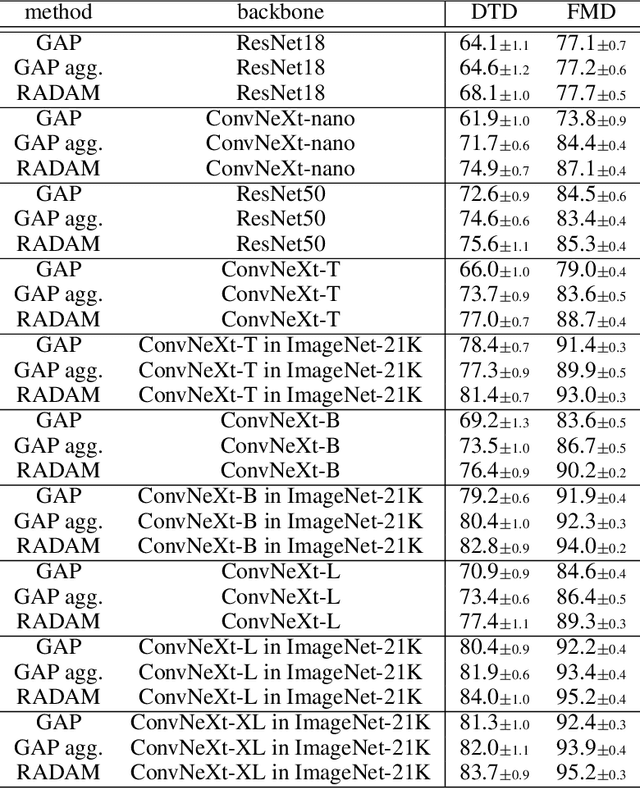
Abstract:Texture analysis is a classical yet challenging task in computer vision for which deep neural networks are actively being applied. Most approaches are based on building feature aggregation modules around a pre-trained backbone and then fine-tuning the new architecture on specific texture recognition tasks. Here we propose a new method named \textbf{R}andom encoding of \textbf{A}ggregated \textbf{D}eep \textbf{A}ctivation \textbf{M}aps (RADAM) which extracts rich texture representations without ever changing the backbone. The technique consists of encoding the output at different depths of a pre-trained deep convolutional network using a Randomized Autoencoder (RAE). The RAE is trained locally to each image using a closed-form solution, and its decoder weights are used to compose a 1-dimensional texture representation that is fed into a linear SVM. This means that no fine-tuning or backpropagation is needed. We explore RADAM on several texture benchmarks and achieve state-of-the-art results with different computational budgets. Our results suggest that pre-trained backbones may not require additional fine-tuning for texture recognition if their learned representations are better encoded.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge