Julien Bert
Machine Learning-Based Modeling of the Anode Heel Effect in X-ray Beam Monte Carlo Simulations
Apr 27, 2025Abstract:This study enhances Monte Carlo simulation accuracy in X-ray imaging by developing an AI-driven model for the anode heel effect, achieving improved beam intensity distribution and dosimetric precision. Through dynamic adjustments to beam weights on the anode and cathode sides of the X-ray tube, our machine learning model effectively replicates the asymmetry characteristic of clinical X-ray beams. Experimental results reveal dose rate increases of up to 9.6% on the cathode side and reductions of up to 12.5% on the anode side, for energy levels between 50 and 120 kVp. These experimentally optimized beam weights were integrated into the OpenGATE and GGEMS Monte Carlo toolkits, significantly advancing dosimetric simulation accuracy and the image quality which closely resembles the clinical imaging. Validation with fluence and dose actors demonstrated that the AI-based model closely mirrors clinical beam behavior, providing substantial improvements in dose consistency and accuracy over conventional X-ray models. This approach provides a robust framework for improving X-ray dosimetry, with potential applications in dose optimization, imaging quality enhancement, and radiation safety in both clinical and research settings.
Semi-Supervised Learning for Dose Prediction in Targeted Radionuclide: A Synthetic Data Study
Mar 07, 2025Abstract:Targeted Radionuclide Therapy (TRT) is a modern strategy in radiation oncology that aims to administer a potent radiation dose specifically to cancer cells using cancer-targeting radiopharmaceuticals. Accurate radiation dose estimation tailored to individual patients is crucial. Deep learning, particularly with pre-therapy imaging, holds promise for personalizing TRT doses. However, current methods require large time series of SPECT imaging, which is hardly achievable in routine clinical practice, and thus raises issues of data availability. Our objective is to develop a semi-supervised learning (SSL) solution to personalize dosimetry using pre-therapy images. The aim is to develop an approach that achieves accurate results when PET/CT images are available, but are associated with only a few post-therapy dosimetry data provided by SPECT images. In this work, we introduce an SSL method using a pseudo-label generation approach for regression tasks inspired by the FixMatch framework. The feasibility of the proposed solution was preliminarily evaluated through an in-silico study using synthetic data and Monte Carlo simulation. Experimental results for organ dose prediction yielded promising outcomes, showing that the use of pseudo-labeled data provides better accuracy compared to using only labeled data.
CT respiratory motion synthesis using joint supervised and adversarial learning
Mar 29, 2024Abstract:Objective: Four-dimensional computed tomography (4DCT) imaging consists in reconstructing a CT acquisition into multiple phases to track internal organ and tumor motion. It is commonly used in radiotherapy treatment planning to establish planning target volumes. However, 4DCT increases protocol complexity, may not align with patient breathing during treatment, and lead to higher radiation delivery. Approach: In this study, we propose a deep synthesis method to generate pseudo respiratory CT phases from static images for motion-aware treatment planning. The model produces patient-specific deformation vector fields (DVFs) by conditioning synthesis on external patient surface-based estimation, mimicking respiratory monitoring devices. A key methodological contribution is to encourage DVF realism through supervised DVF training while using an adversarial term jointly not only on the warped image but also on the magnitude of the DVF itself. This way, we avoid excessive smoothness typically obtained through deep unsupervised learning, and encourage correlations with the respiratory amplitude. Main results: Performance is evaluated using real 4DCT acquisitions with smaller tumor volumes than previously reported. Results demonstrate for the first time that the generated pseudo-respiratory CT phases can capture organ and tumor motion with similar accuracy to repeated 4DCT scans of the same patient. Mean inter-scans tumor center-of-mass distances and Dice similarity coefficients were $1.97$mm and $0.63$, respectively, for real 4DCT phases and $2.35$mm and $0.71$ for synthetic phases, and compares favorably to a state-of-the-art technique (RMSim).
Cross-modal tumor segmentation using generative blending augmentation and self training
Apr 04, 2023Abstract:Deep learning for medical imaging is limited by data scarcity and domain shift, which lead to biased training sets that do not accurately represent deployment conditions. A related practical problem is cross-modal segmentation where the objective is to segment unlabelled domains using previously labelled images from other modalites, which is the context of the MICCAI CrossMoDA 2022 challenge on vestibular schwannoma (VS) segmentation. In this context, we propose a VS segmentation method that leverages conventional image-to-image translation and segmentation using iterative self training combined to a dedicated data augmentation technique called Generative Blending Augmentation (GBA). GBA is based on a one-shot 2D SinGAN generative model that allows to realistically diversify target tumor appearances in a downstream segmentation model, improving its generalization power at test time. Our solution ranked first on the VS segmentation task during the validation and test phase of the CrossModa 2022 challenge.
Regularized directional representations for medical image registration
Nov 30, 2021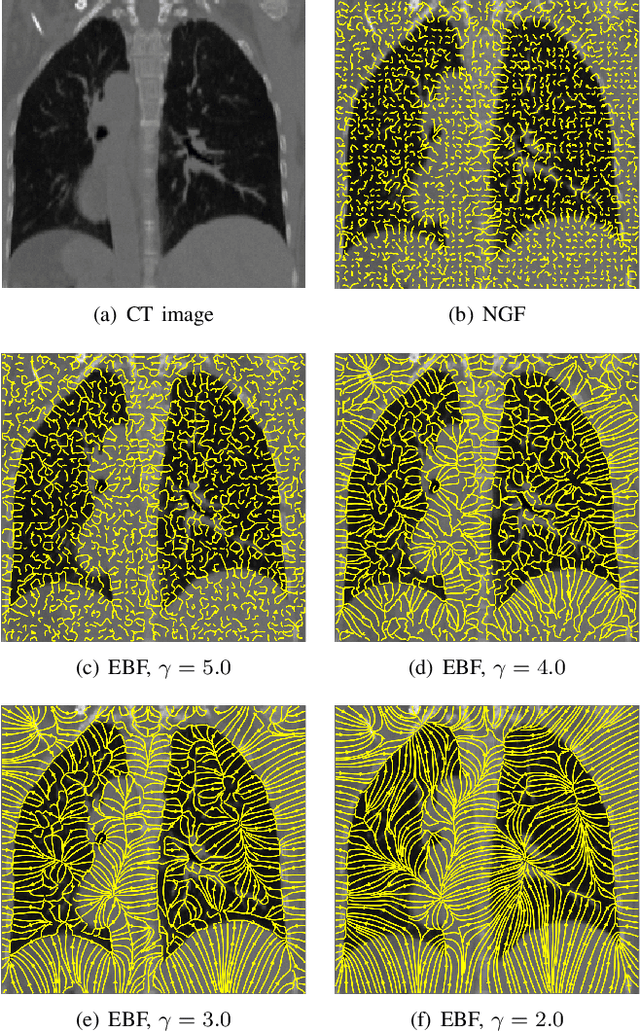
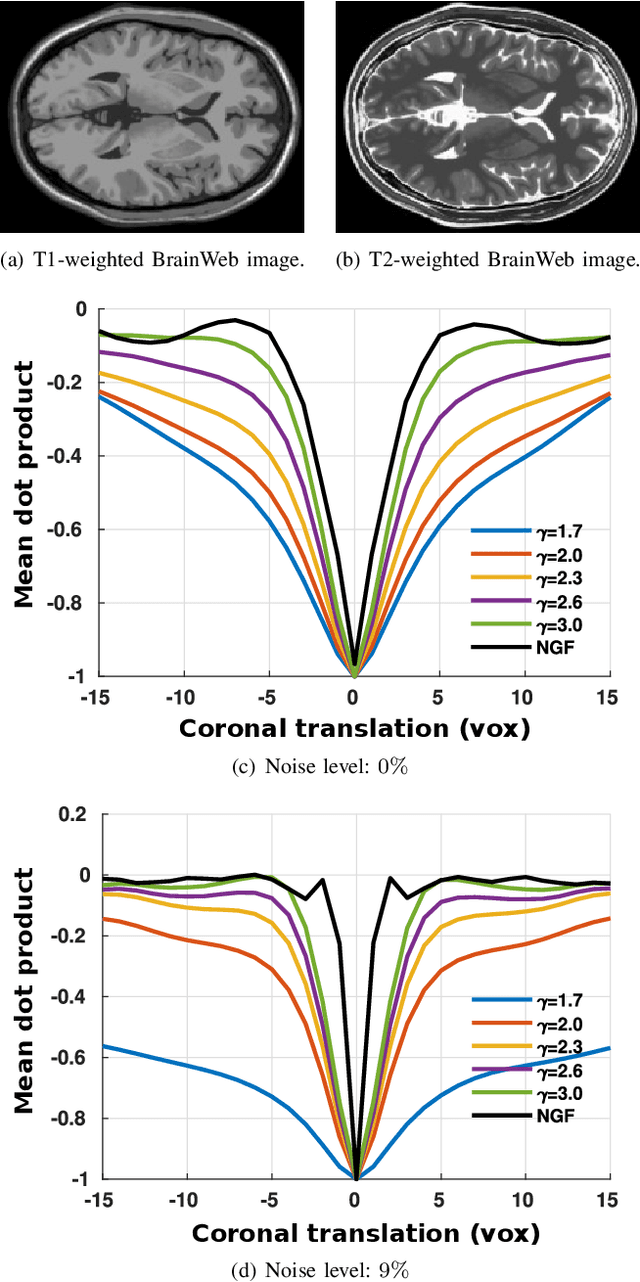
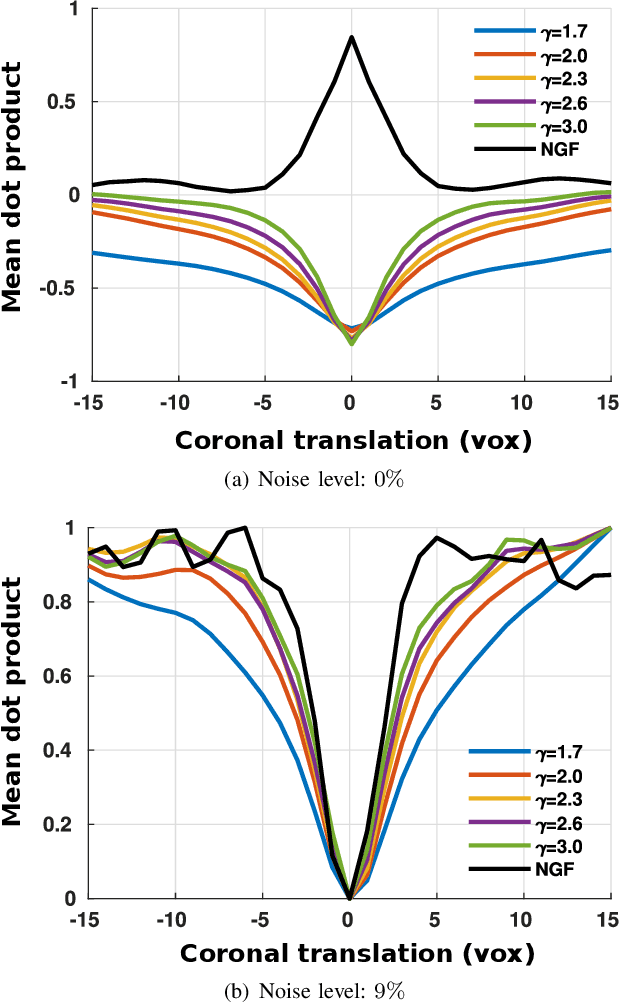
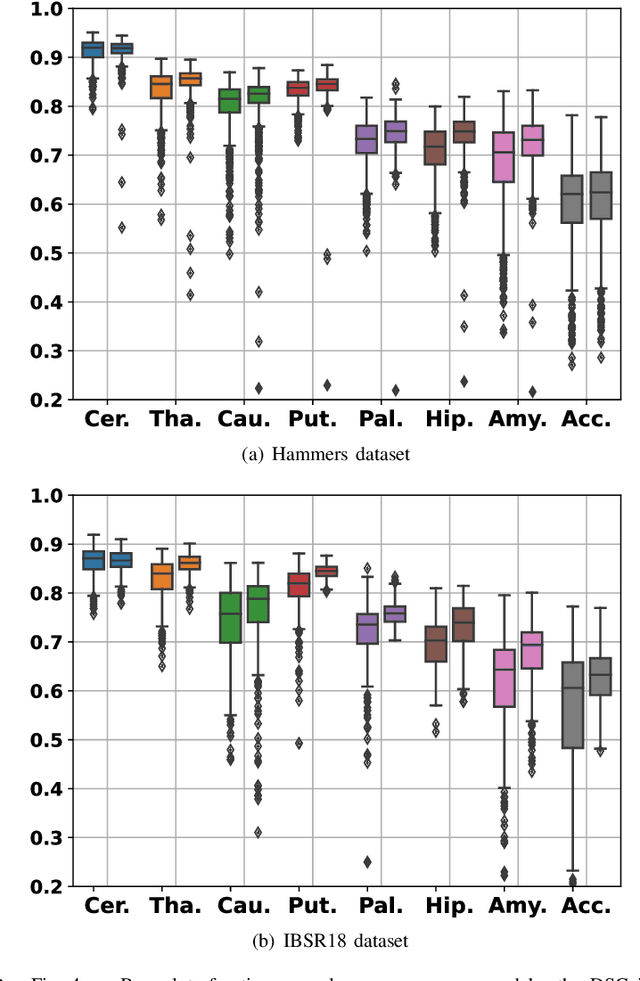
Abstract:In image registration, many efforts have been devoted to the development of alternatives to the popular normalized mutual information criterion. Concurrently to these efforts, an increasing number of works have demonstrated that substantial gains in registration accuracy can also be achieved by aligning structural representations of images rather than images themselves. Following this research path, we propose a new method for mono- and multimodal image registration based on the alignment of regularized vector fields derived from structural information such as gradient vector flow fields, a technique we call \textit{vector field similarity}. Our approach can be combined in a straightforward fashion with any existing registration framework by substituting vector field similarity to intensity-based registration. In our experiments, we show that the proposed approach compares favourably with conventional image alignment on several public image datasets using a diversity of imaging modalities and anatomical locations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge