Juan Gómez Luna
Hardware Acceleration for Knowledge Graph Processing: Challenges & Recent Developments
Aug 22, 2024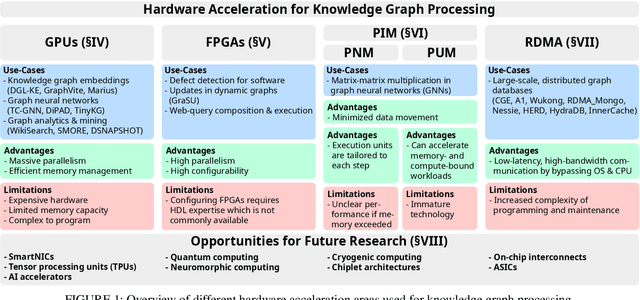
Abstract:Knowledge graphs (KGs) have achieved significant attention in recent years, particularly in the area of the Semantic Web as well as gaining popularity in other application domains such as data mining and search engines. Simultaneously, there has been enormous progress in the development of different types of heterogeneous hardware, impacting the way KGs are processed. The aim of this paper is to provide a systematic literature review of knowledge graph hardware acceleration. For this, we present a classification of the primary areas in knowledge graph technology that harnesses different hardware units for accelerating certain knowledge graph functionalities. We then extensively describe respective works, focusing on how KG related schemes harness modern hardware accelerators. Based on our review, we identify various research gaps and future exploratory directions that are anticipated to be of significant value both for academics and industry practitioners.
ApHMM: Accelerating Profile Hidden Markov Models for Fast and Energy-Efficient Genome Analysis
Jul 20, 2022
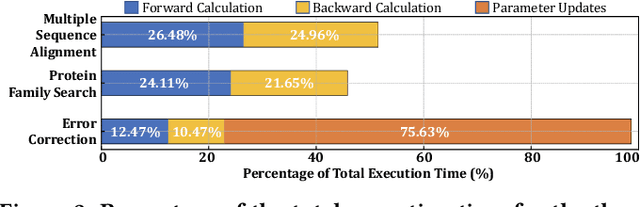
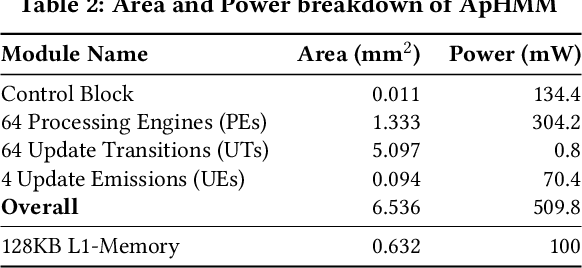
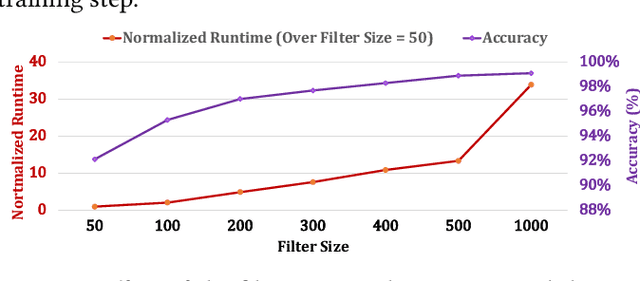
Abstract:Profile hidden Markov models (pHMMs) are widely used in many bioinformatics applications to accurately identify similarities between biological sequences (e.g., DNA or protein sequences). PHMMs use a commonly-adopted and highly-accurate method, called the Baum-Welch algorithm, to calculate these similarities. However, the Baum-Welch algorithm is computationally expensive, and existing works provide either software- or hardware-only solutions for a fixed pHMM design. When we analyze the state-of-the-art works, we find that there is a pressing need for a flexible, high-performant, and energy-efficient hardware-software co-design to efficiently and effectively solve all the major inefficiencies in the Baum-Welch algorithm for pHMMs. We propose ApHMM, the first flexible acceleration framework that can significantly reduce computational and energy overheads of the Baum-Welch algorithm for pHMMs. ApHMM leverages hardware-software co-design to solve the major inefficiencies in the Baum-Welch algorithm by 1) designing a flexible hardware to support different pHMMs designs, 2) exploiting the predictable data dependency pattern in an on-chip memory with memoization techniques, 3) quickly eliminating negligible computations with a hardware-based filter, and 4) minimizing the redundant computations. We implement our 1) hardware-software optimizations on a specialized hardware and 2) software optimizations for GPUs to provide the first flexible Baum-Welch accelerator for pHMMs. ApHMM provides significant speedups of 15.55x-260.03x, 1.83x-5.34x, and 27.97x compared to CPU, GPU, and FPGA implementations of the Baum-Welch algorithm, respectively. ApHMM outperforms the state-of-the-art CPU implementations of three important bioinformatics applications, 1) error correction, 2) protein family search, and 3) multiple sequence alignment, by 1.29x-59.94x, 1.03x-1.75x, and 1.03x-1.95x, respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge