Jeffrey Mendenhall
Deep Learning for Virtual Screening: Five Reasons to Use ROC Cost Functions
Jun 25, 2020Abstract:Computer-aided drug discovery is an essential component of modern drug development. Therein, deep learning has become an important tool for rapid screening of billions of molecules in silico for potential hits containing desired chemical features. Despite its importance, substantial challenges persist in training these models, such as severe class imbalance, high decision thresholds, and lack of ground truth labels in some datasets. In this work we argue in favor of directly optimizing the receiver operating characteristic (ROC) in such cases, due to its robustness to class imbalance, its ability to compromise over different decision thresholds, certain freedom to influence the relative weights in this compromise, fidelity to typical benchmarking measures, and equivalence to positive/unlabeled learning. We also propose new training schemes (coherent mini-batch arrangement, and usage of out-of-batch samples) for cost functions based on the ROC, as well as a cost function based on the logAUC metric that facilitates early enrichment (i.e. improves performance at high decision thresholds, as often desired when synthesizing predicted hit compounds). We demonstrate that these approaches outperform standard deep learning approaches on a series of PubChem high-throughput screening datasets that represent realistic and diverse drug discovery campaigns on major drug target families.
3D Deep Learning for Biological Function Prediction from Physical Fields
Apr 13, 2017
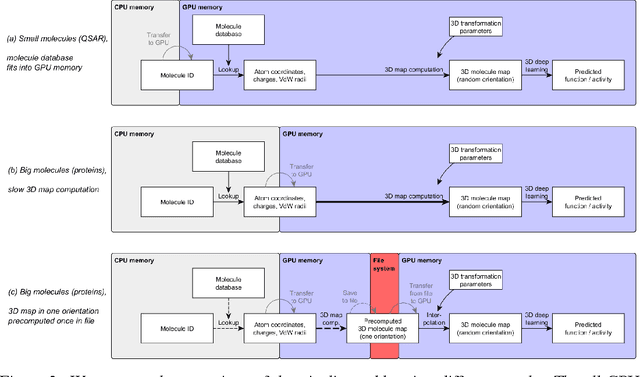
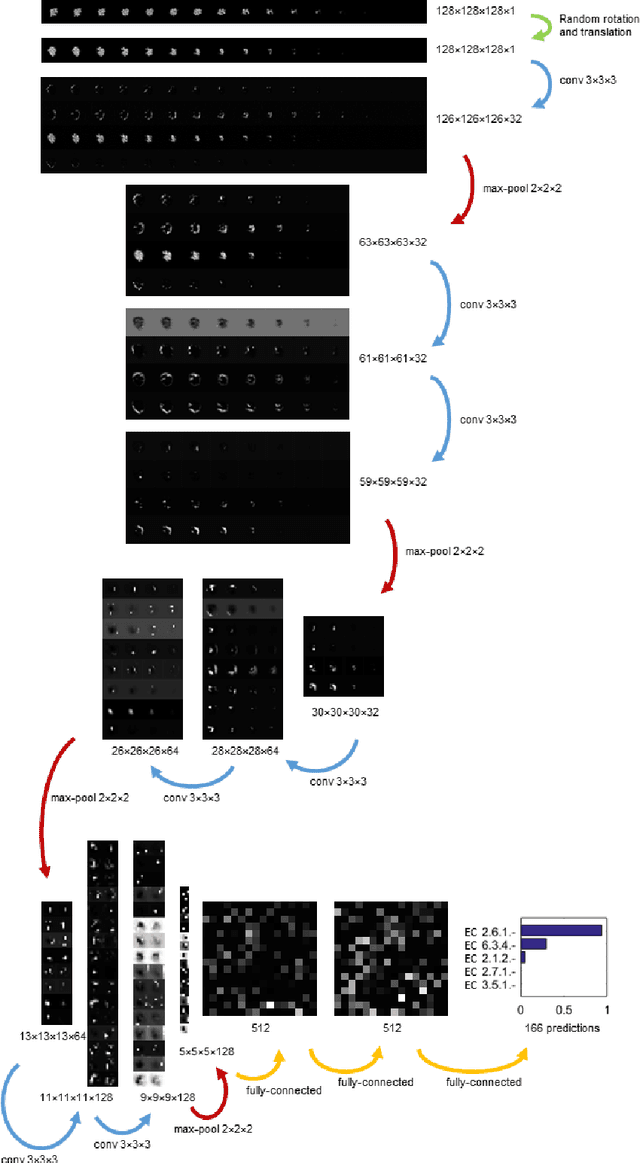
Abstract:Predicting the biological function of molecules, be it proteins or drug-like compounds, from their atomic structure is an important and long-standing problem. Function is dictated by structure, since it is by spatial interactions that molecules interact with each other, both in terms of steric complementarity, as well as intermolecular forces. Thus, the electron density field and electrostatic potential field of a molecule contain the "raw fingerprint" of how this molecule can fit to binding partners. In this paper, we show that deep learning can predict biological function of molecules directly from their raw 3D approximated electron density and electrostatic potential fields. Protein function based on EC numbers is predicted from the approximated electron density field. In another experiment, the activity of small molecules is predicted with quality comparable to state-of-the-art descriptor-based methods. We propose several alternative computational models for the GPU with different memory and runtime requirements for different sizes of molecules and of databases. We also propose application-specific multi-channel data representations. With future improvements of training datasets and neural network settings in combination with complementary information sources (sequence, genomic context, expression level), deep learning can be expected to show its generalization power and revolutionize the field of molecular function prediction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge