Javier L. Prieto
Insulin Resistance Prediction From Wearables and Routine Blood Biomarkers
Apr 30, 2025Abstract:Insulin resistance, a precursor to type 2 diabetes, is characterized by impaired insulin action in tissues. Current methods for measuring insulin resistance, while effective, are expensive, inaccessible, not widely available and hinder opportunities for early intervention. In this study, we remotely recruited the largest dataset to date across the US to study insulin resistance (N=1,165 participants, with median BMI=28 kg/m2, age=45 years, HbA1c=5.4%), incorporating wearable device time series data and blood biomarkers, including the ground-truth measure of insulin resistance, homeostatic model assessment for insulin resistance (HOMA-IR). We developed deep neural network models to predict insulin resistance based on readily available digital and blood biomarkers. Our results show that our models can predict insulin resistance by combining both wearable data and readily available blood biomarkers better than either of the two data sources separately (R2=0.5, auROC=0.80, Sensitivity=76%, and specificity 84%). The model showed 93% sensitivity and 95% adjusted specificity in obese and sedentary participants, a subpopulation most vulnerable to developing type 2 diabetes and who could benefit most from early intervention. Rigorous evaluation of model performance, including interpretability, and robustness, facilitates generalizability across larger cohorts, which is demonstrated by reproducing the prediction performance on an independent validation cohort (N=72 participants). Additionally, we demonstrated how the predicted insulin resistance can be integrated into a large language model agent to help understand and contextualize HOMA-IR values, facilitating interpretation and safe personalized recommendations. This work offers the potential for early detection of people at risk of type 2 diabetes and thereby facilitate earlier implementation of preventative strategies.
A Scalable Framework for Evaluating Health Language Models
Apr 01, 2025Abstract:Large language models (LLMs) have emerged as powerful tools for analyzing complex datasets. Recent studies demonstrate their potential to generate useful, personalized responses when provided with patient-specific health information that encompasses lifestyle, biomarkers, and context. As LLM-driven health applications are increasingly adopted, rigorous and efficient one-sided evaluation methodologies are crucial to ensure response quality across multiple dimensions, including accuracy, personalization and safety. Current evaluation practices for open-ended text responses heavily rely on human experts. This approach introduces human factors and is often cost-prohibitive, labor-intensive, and hinders scalability, especially in complex domains like healthcare where response assessment necessitates domain expertise and considers multifaceted patient data. In this work, we introduce Adaptive Precise Boolean rubrics: an evaluation framework that streamlines human and automated evaluation of open-ended questions by identifying gaps in model responses using a minimal set of targeted rubrics questions. Our approach is based on recent work in more general evaluation settings that contrasts a smaller set of complex evaluation targets with a larger set of more precise, granular targets answerable with simple boolean responses. We validate this approach in metabolic health, a domain encompassing diabetes, cardiovascular disease, and obesity. Our results demonstrate that Adaptive Precise Boolean rubrics yield higher inter-rater agreement among expert and non-expert human evaluators, and in automated assessments, compared to traditional Likert scales, while requiring approximately half the evaluation time of Likert-based methods. This enhanced efficiency, particularly in automated evaluation and non-expert contributions, paves the way for more extensive and cost-effective evaluation of LLMs in health.
Lifestyle-Informed Personalized Blood Biomarker Prediction via Novel Representation Learning
Jul 09, 2024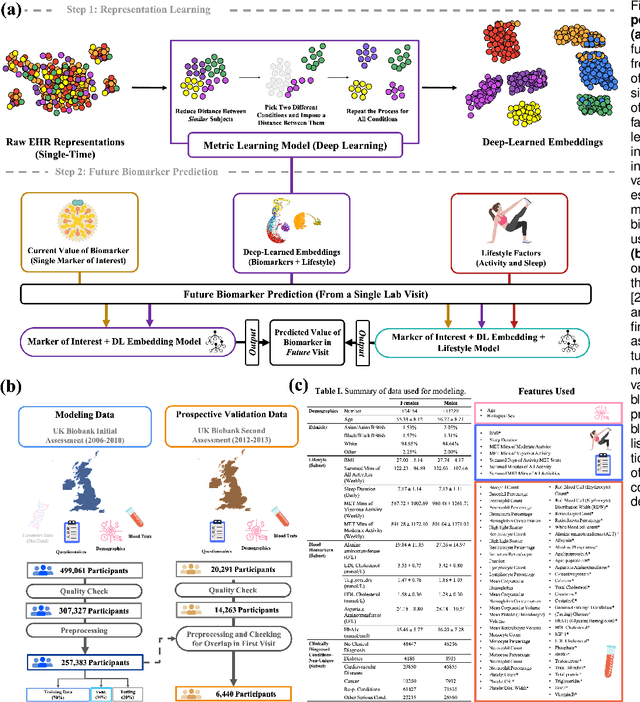
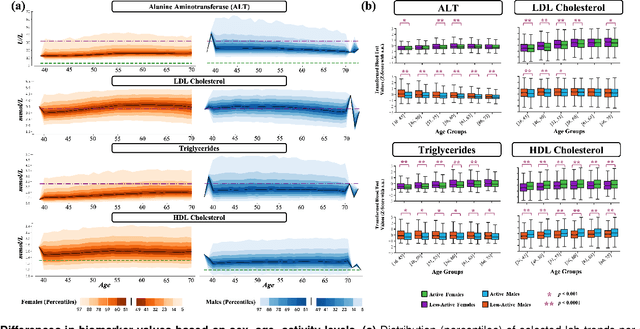
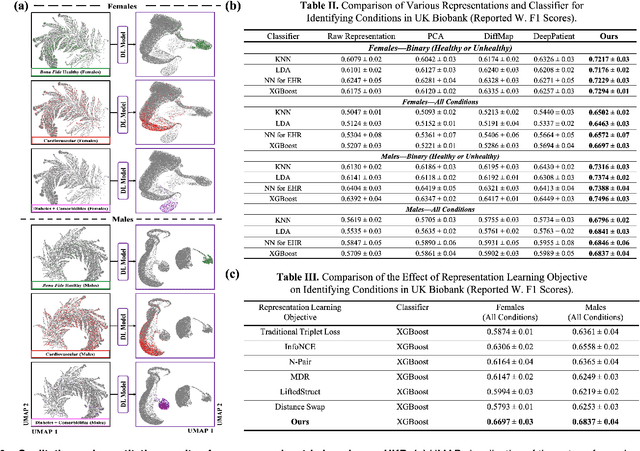
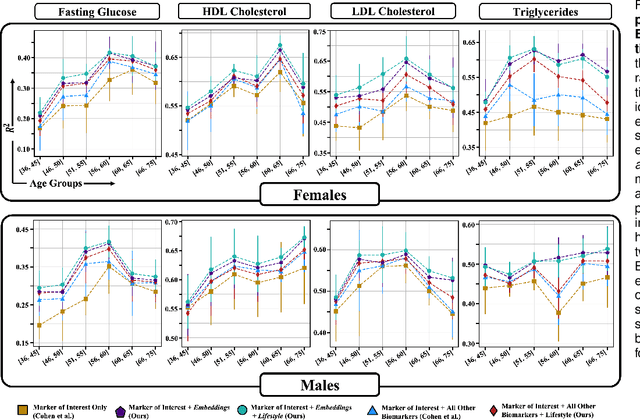
Abstract:Blood biomarkers are an essential tool for healthcare providers to diagnose, monitor, and treat a wide range of medical conditions. Current reference values and recommended ranges often rely on population-level statistics, which may not adequately account for the influence of inter-individual variability driven by factors such as lifestyle and genetics. In this work, we introduce a novel framework for predicting future blood biomarker values and define personalized references through learned representations from lifestyle data (physical activity and sleep) and blood biomarkers. Our proposed method learns a similarity-based embedding space that captures the complex relationship between biomarkers and lifestyle factors. Using the UK Biobank (257K participants), our results show that our deep-learned embeddings outperform traditional and current state-of-the-art representation learning techniques in predicting clinical diagnosis. Using a subset of UK Biobank of 6440 participants who have follow-up visits, we validate that the inclusion of these embeddings and lifestyle factors directly in blood biomarker models improves the prediction of future lab values from a single lab visit. This personalized modeling approach provides a foundation for developing more accurate risk stratification tools and tailoring preventative care strategies. In clinical settings, this translates to the potential for earlier disease detection, more timely interventions, and ultimately, a shift towards personalized healthcare.
No Pairs Left Behind: Improving Metric Learning with Regularized Triplet Objective
Oct 18, 2022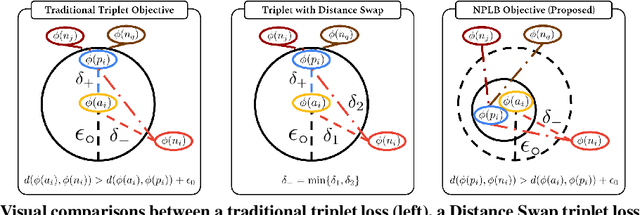

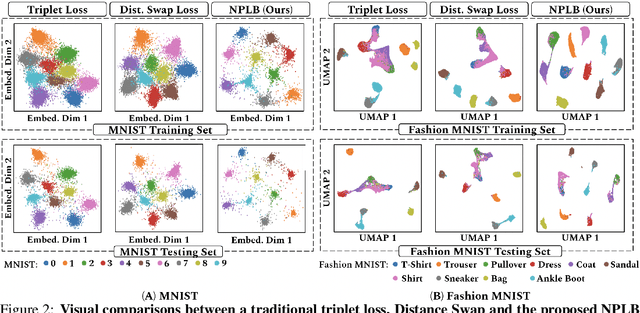
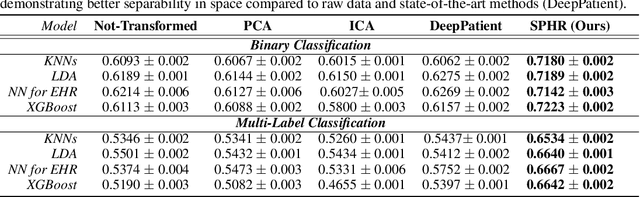
Abstract:We propose a novel formulation of the triplet objective function that improves metric learning without additional sample mining or overhead costs. Our approach aims to explicitly regularize the distance between the positive and negative samples in a triplet with respect to the anchor-negative distance. As an initial validation, we show that our method (called No Pairs Left Behind [NPLB]) improves upon the traditional and current state-of-the-art triplet objective formulations on standard benchmark datasets. To show the effectiveness and potentials of NPLB on real-world complex data, we evaluate our approach on a large-scale healthcare dataset (UK Biobank), demonstrating that the embeddings learned by our model significantly outperform all other current representations on tested downstream tasks. Additionally, we provide a new model-agnostic single-time health risk definition that, when used in tandem with the learned representations, achieves the most accurate prediction of subjects' future health complications. Our results indicate that NPLB is a simple, yet effective framework for improving existing deep metric learning models, showcasing the potential implications of metric learning in more complex applications, especially in the biological and healthcare domains.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge