Jacobus Preller
Recent Methodological Advances in Federated Learning for Healthcare
Oct 04, 2023
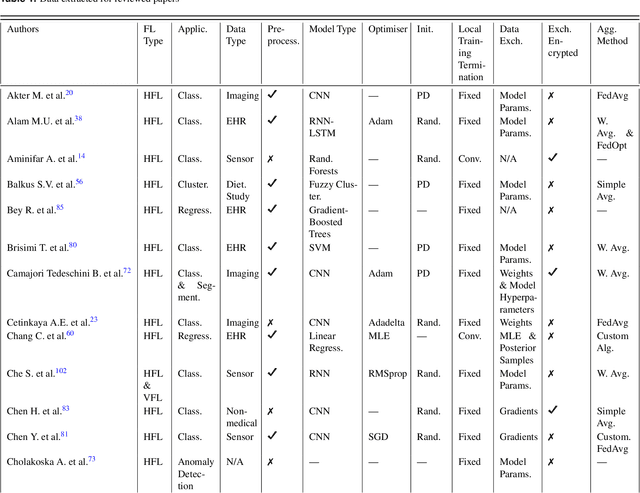
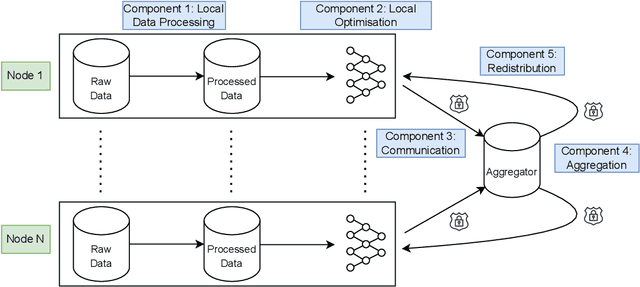
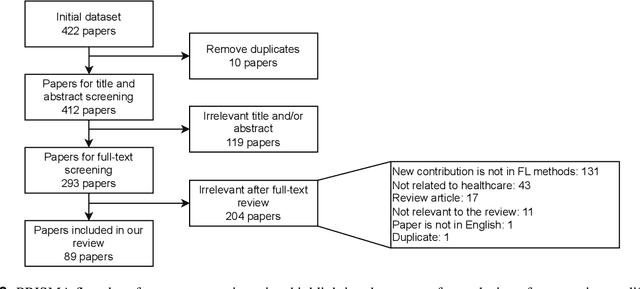
Abstract:For healthcare datasets, it is often not possible to combine data samples from multiple sites due to ethical, privacy or logistical concerns. Federated learning allows for the utilisation of powerful machine learning algorithms without requiring the pooling of data. Healthcare data has many simultaneous challenges which require new methodologies to address, such as highly-siloed data, class imbalance, missing data, distribution shifts and non-standardised variables. Federated learning adds significant methodological complexity to conventional centralised machine learning, requiring distributed optimisation, communication between nodes, aggregation of models and redistribution of models. In this systematic review, we consider all papers on Scopus that were published between January 2015 and February 2023 and which describe new federated learning methodologies for addressing challenges with healthcare data. We performed a detailed review of the 89 papers which fulfilled these criteria. Significant systemic issues were identified throughout the literature which compromise the methodologies in many of the papers reviewed. We give detailed recommendations to help improve the quality of the methodology development for federated learning in healthcare.
Reinterpreting survival analysis in the universal approximator age
Jul 25, 2023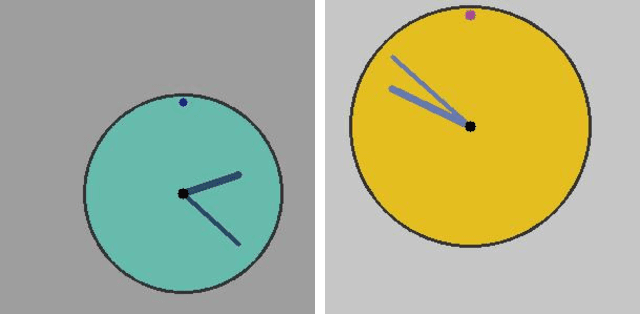
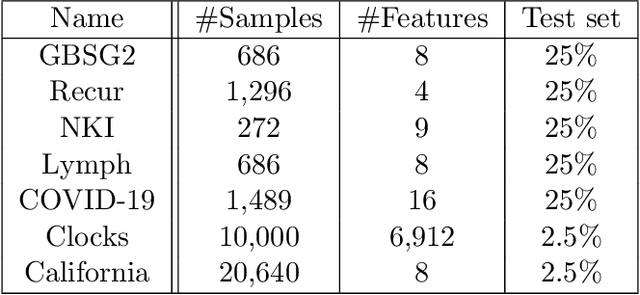
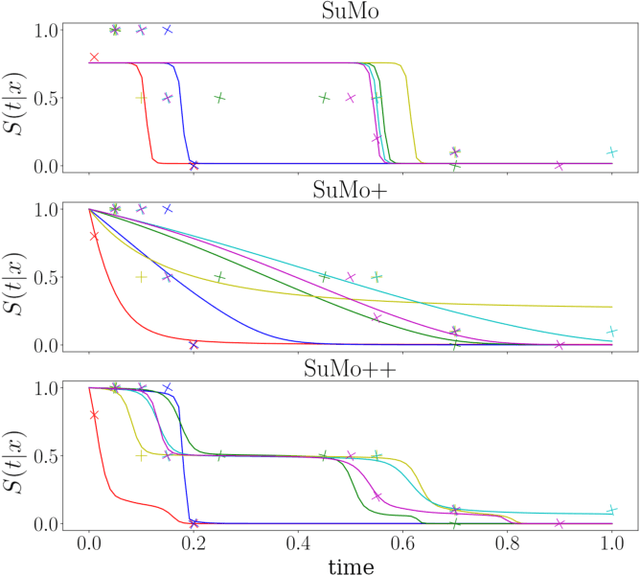
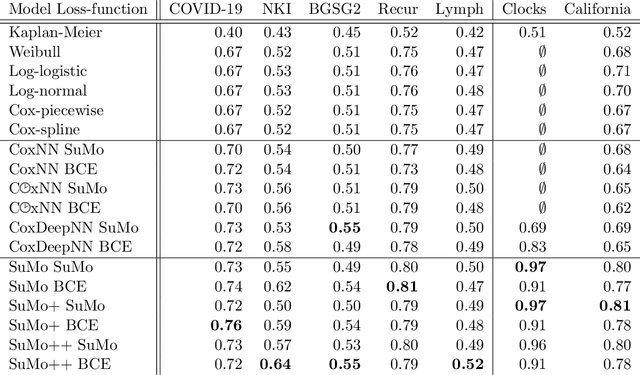
Abstract:Survival analysis is an integral part of the statistical toolbox. However, while most domains of classical statistics have embraced deep learning, survival analysis only recently gained some minor attention from the deep learning community. This recent development is likely in part motivated by the COVID-19 pandemic. We aim to provide the tools needed to fully harness the potential of survival analysis in deep learning. On the one hand, we discuss how survival analysis connects to classification and regression. On the other hand, we provide technical tools. We provide a new loss function, evaluation metrics, and the first universal approximating network that provably produces survival curves without numeric integration. We show that the loss function and model outperform other approaches using a large numerical study.
Dis-AE: Multi-domain & Multi-task Generalisation on Real-World Clinical Data
Jun 15, 2023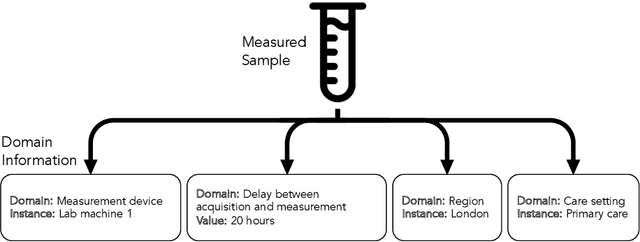
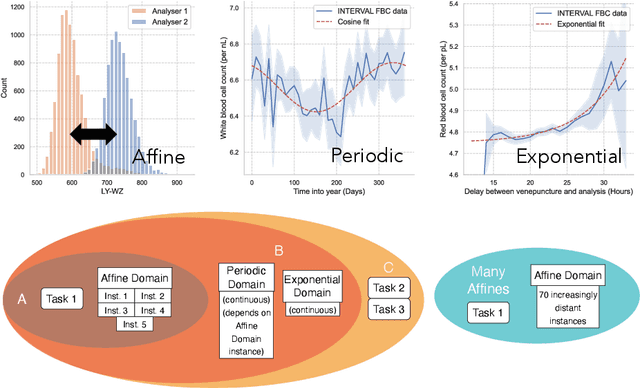
Abstract:Clinical data is often affected by clinically irrelevant factors such as discrepancies between measurement devices or differing processing methods between sites. In the field of machine learning (ML), these factors are known as domains and the distribution differences they cause in the data are known as domain shifts. ML models trained using data from one domain often perform poorly when applied to data from another domain, potentially leading to wrong predictions. As such, developing machine learning models that can generalise well across multiple domains is a challenging yet essential task in the successful application of ML in clinical practice. In this paper, we propose a novel disentangled autoencoder (Dis-AE) neural network architecture that can learn domain-invariant data representations for multi-label classification of medical measurements even when the data is influenced by multiple interacting domain shifts at once. The model utilises adversarial training to produce data representations from which the domain can no longer be determined. We evaluate the model's domain generalisation capabilities on synthetic datasets and full blood count (FBC) data from blood donors as well as primary and secondary care patients, showing that Dis-AE improves model generalisation on multiple domains simultaneously while preserving clinically relevant information.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge