Harvir Sahota
Lizard: A Large-Scale Dataset for Colonic Nuclear Instance Segmentation and Classification
Aug 25, 2021
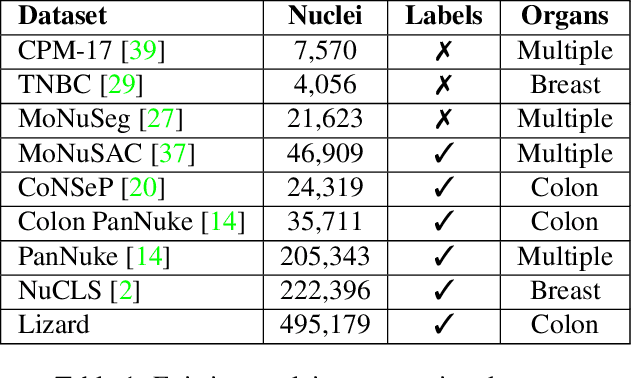
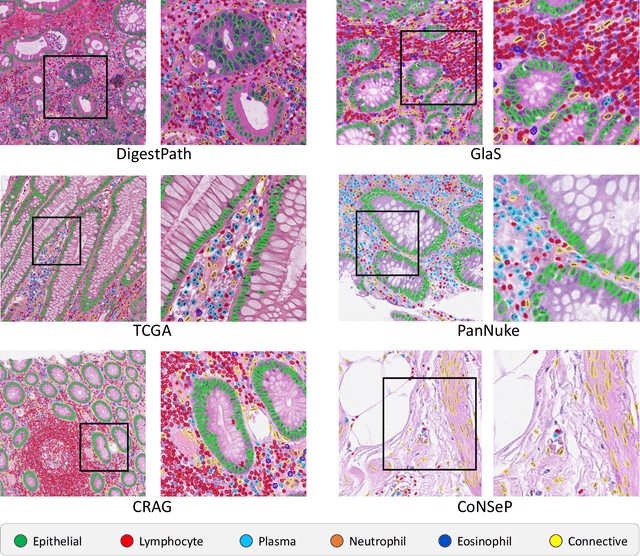
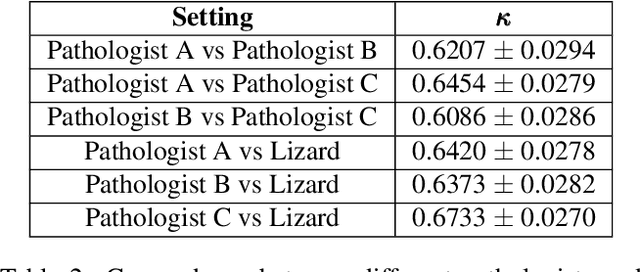
Abstract:The development of deep segmentation models for computational pathology (CPath) can help foster the investigation of interpretable morphological biomarkers. Yet, there is a major bottleneck in the success of such approaches because supervised deep learning models require an abundance of accurately labelled data. This issue is exacerbated in the field of CPath because the generation of detailed annotations usually demands the input of a pathologist to be able to distinguish between different tissue constructs and nuclei. Manually labelling nuclei may not be a feasible approach for collecting large-scale annotated datasets, especially when a single image region can contain thousands of different cells. However, solely relying on automatic generation of annotations will limit the accuracy and reliability of ground truth. Therefore, to help overcome the above challenges, we propose a multi-stage annotation pipeline to enable the collection of large-scale datasets for histology image analysis, with pathologist-in-the-loop refinement steps. Using this pipeline, we generate the largest known nuclear instance segmentation and classification dataset, containing nearly half a million labelled nuclei in H&E stained colon tissue. We have released the dataset and encourage the research community to utilise it to drive forward the development of downstream cell-based models in CPath.
Semantic annotation for computational pathology: Multidisciplinary experience and best practice recommendations
Jun 25, 2021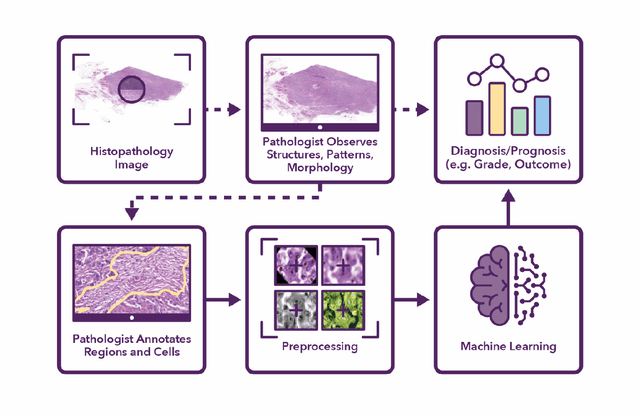
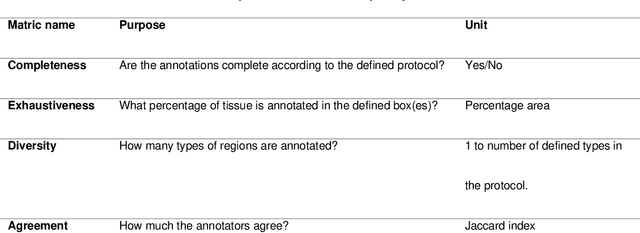

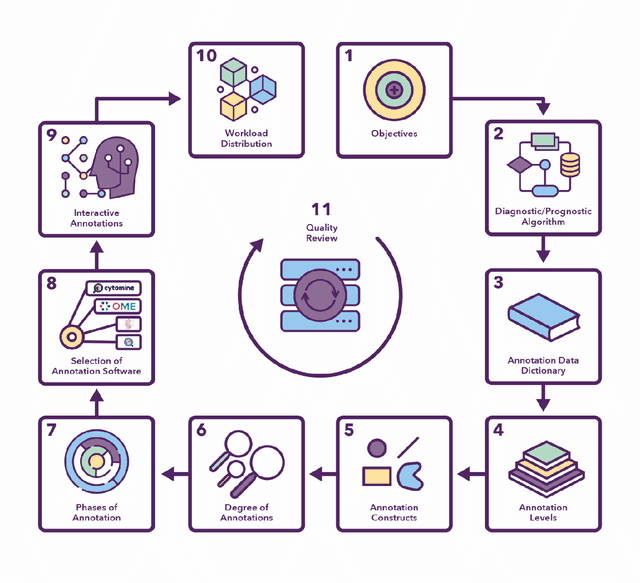
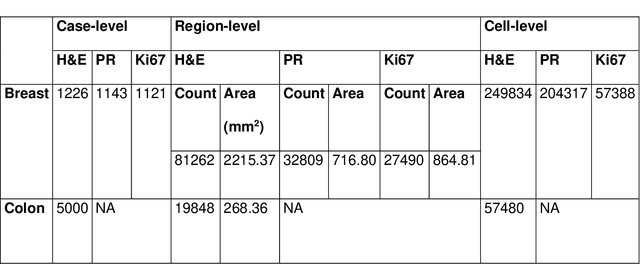
Abstract:Recent advances in whole slide imaging (WSI) technology have led to the development of a myriad of computer vision and artificial intelligence (AI) based diagnostic, prognostic, and predictive algorithms. Computational Pathology (CPath) offers an integrated solution to utilize information embedded in pathology WSIs beyond what we obtain through visual assessment. For automated analysis of WSIs and validation of machine learning (ML) models, annotations at the slide, tissue and cellular levels are required. The annotation of important visual constructs in pathology images is an important component of CPath projects. Improper annotations can result in algorithms which are hard to interpret and can potentially produce inaccurate and inconsistent results. Despite the crucial role of annotations in CPath projects, there are no well-defined guidelines or best practices on how annotations should be carried out. In this paper, we address this shortcoming by presenting the experience and best practices acquired during the execution of a large-scale annotation exercise involving a multidisciplinary team of pathologists, ML experts and researchers as part of the Pathology image data Lake for Analytics, Knowledge and Education (PathLAKE) consortium. We present a real-world case study along with examples of different types of annotations, diagnostic algorithm, annotation data dictionary and annotation constructs. The analyses reported in this work highlight best practice recommendations that can be used as annotation guidelines over the lifecycle of a CPath project.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge