Hans Pinckaers
Improved Multimodal Fusion for Small Datasets with Auxiliary Supervision
Apr 01, 2023Abstract:Prostate cancer is one of the leading causes of cancer-related death in men worldwide. Like many cancers, diagnosis involves expert integration of heterogeneous patient information such as imaging, clinical risk factors, and more. For this reason, there have been many recent efforts toward deep multimodal fusion of image and non-image data for clinical decision tasks. Many of these studies propose methods to fuse learned features from each patient modality, providing significant downstream improvements with techniques like cross-modal attention gating, Kronecker product fusion, orthogonality regularization, and more. While these enhanced fusion operations can improve upon feature concatenation, they often come with an extremely high learning capacity, meaning they are likely to overfit when applied even to small or low-dimensional datasets. Rather than designing a highly expressive fusion operation, we propose three simple methods for improved multimodal fusion with small datasets that aid optimization by generating auxiliary sources of supervision during training: extra supervision, clinical prediction, and dense fusion. We validate the proposed approaches on prostate cancer diagnosis from paired histopathology imaging and tabular clinical features. The proposed methods are straightforward to implement and can be applied to any classification task with paired image and non-image data.
Detection of prostate cancer in whole-slide images through end-to-end training with image-level labels
Jun 05, 2020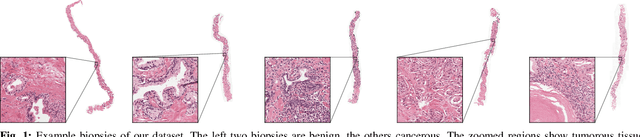
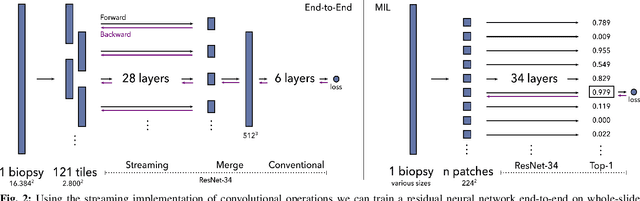
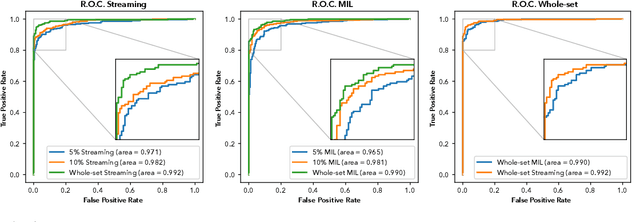
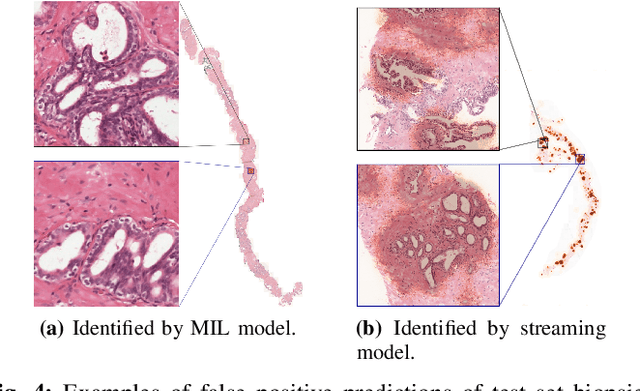
Abstract:Prostate cancer is the most prevalent cancer among men in Western countries, with 1.1 million new diagnoses every year. The gold standard for the diagnosis of prostate cancer is a pathologists' evaluation of prostate tissue. To potentially assist pathologists deep-learning-based cancer detection systems have been developed. Many of the state-of-the-art models are patch-based convolutional neural networks, as the use of entire scanned slides is hampered by memory limitations on accelerator cards. Patch-based systems typically require detailed, pixel-level annotations for effective training. However, such annotations are seldom readily available, in contrast to the clinical reports of pathologists, which contain slide-level labels. As such, developing algorithms which do not require manual pixel-wise annotations, but can learn using only the clinical report would be a significant advancement for the field. In this paper, we propose to use a streaming implementation of convolutional layers, to train a modern CNN (ResNet-34) with 21 million parameters end-to-end on 4712 prostate biopsies. The method enables the use of entire biopsy images at high-resolution directly by reducing the GPU memory requirements by 2.4 TB. We show that modern CNNs, trained using our streaming approach, can extract meaningful features from high-resolution images without additional heuristics, reaching similar performance as state-of-the-art patch-based and multiple-instance learning methods. By circumventing the need for manual annotations, this approach can function as a blueprint for other tasks in histopathological diagnosis. The source code to reproduce the streaming models is available at https://github.com/DIAGNijmegen/pathology-streaming-pipeline .
Streaming convolutional neural networks for end-to-end learning with multi-megapixel images
Nov 11, 2019
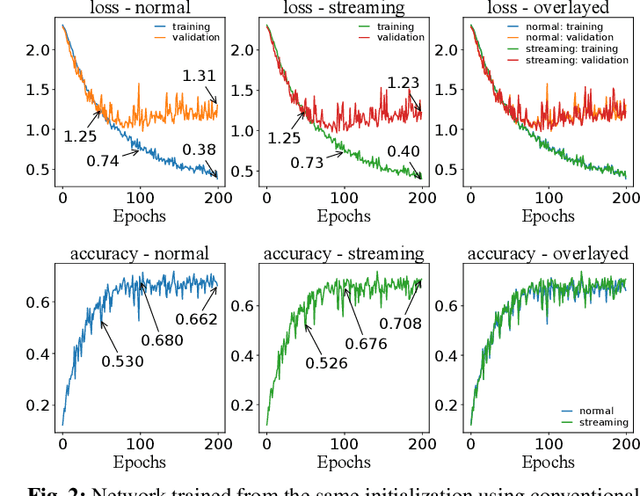

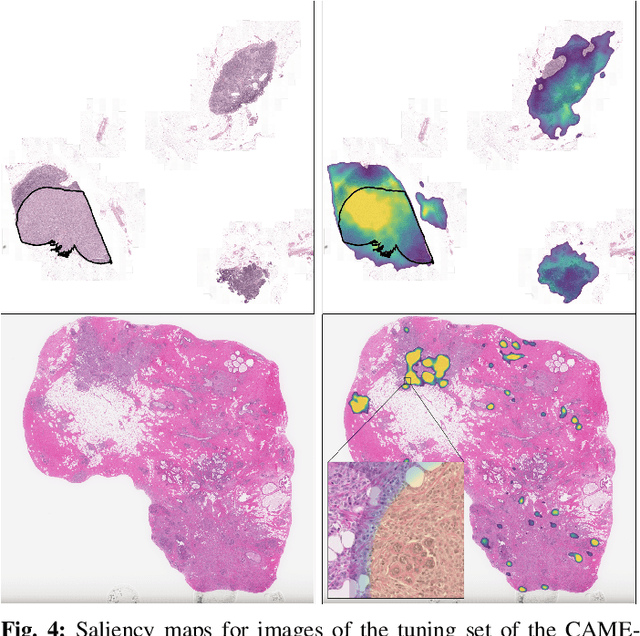
Abstract:Due to memory constraints on current hardware, most convolution neural networks (CNN) are trained on sub-megapixel images. For example, most popular datasets in computer vision contain images much less than a megapixel in size (0.09MP for ImageNet and 0.001MP for CIFAR-10). In some domains such as medical imaging, multi-megapixel images are needed to identify the presence of disease accurately. We propose a novel method to directly train convolutional neural networks using any input image size end-to-end. This method exploits the locality of most operations in modern convolutional neural networks by performing the forward and backward pass on smaller tiles of the image. In this work, we show a proof of concept using images of up to 66-megapixels (8192x8192), saving approximately 50GB of memory per image. Using two public challenge datasets, we demonstrate that CNNs can learn to extract relevant information from these large images and benefit from increasing resolution. We improved the area under the receiver-operating characteristic curve from 0.580 (4MP) to 0.706 (66MP) for metastasis detection in breast cancer (CAMELYON17). We also obtained a Spearman correlation metric approaching state-of-the-art performance on the TUPAC16 dataset, from 0.485 (1MP) to 0.570 (16MP). Code to reproduce a subset of the experiments is available at https://github.com/DIAGNijmegen/StreamingCNN.
Neural Ordinary Differential Equations for Semantic Segmentation of Individual Colon Glands
Oct 23, 2019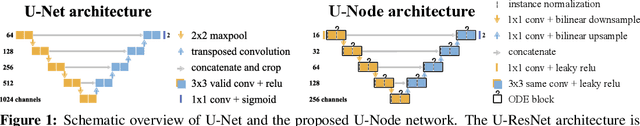
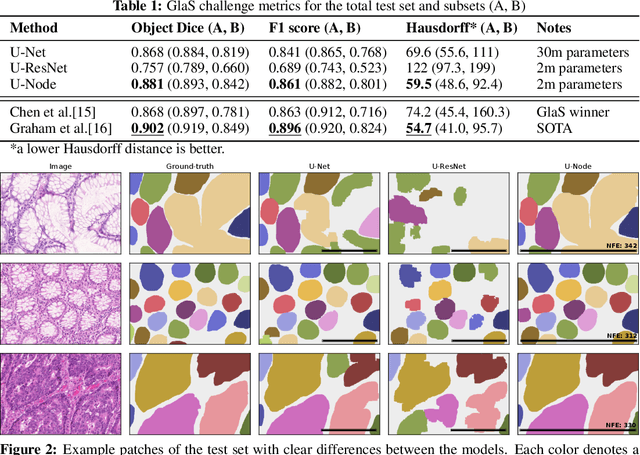
Abstract:Automated medical image segmentation plays a key role in quantitative research and diagnostics. Convolutional neural networks based on the U-Net architecture are the state-of-the-art. A key disadvantage is the hard-coding of the receptive field size, which requires architecture optimization for each segmentation task. Furthermore, increasing the receptive field results in an increasing number of weights. Recently, Neural Ordinary Differential Equations (NODE) have been proposed, a new type of continuous depth deep neural network. This framework allows for a dynamic receptive field at a fixed memory cost and a smaller amount of parameters. We show on a colon gland segmentation dataset (GlaS) that these NODEs can be used within the U-Net framework to improve segmentation results while reducing memory load and parameter counts.
Automated Gleason Grading of Prostate Biopsies using Deep Learning
Jul 18, 2019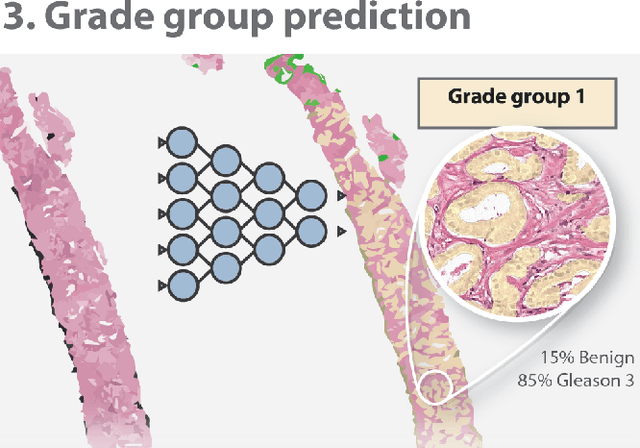

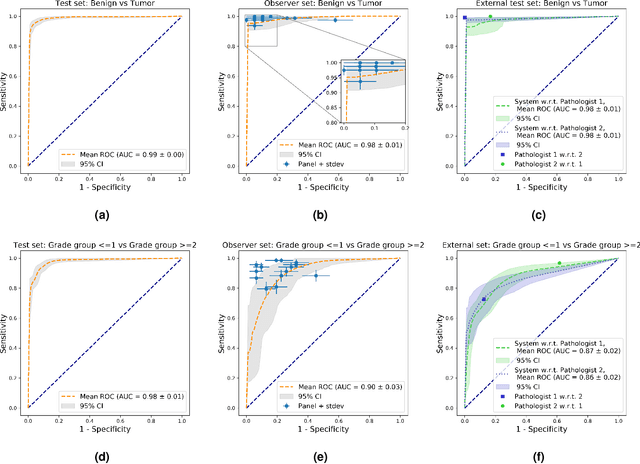
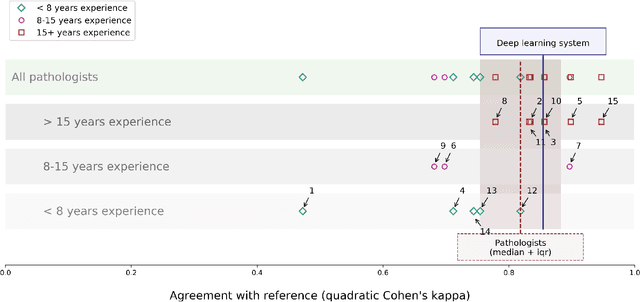
Abstract:The Gleason score is the most important prognostic marker for prostate cancer patients but suffers from significant inter-observer variability. We developed a fully automated deep learning system to grade prostate biopsies. The system was developed using 5834 biopsies from 1243 patients. A semi-automatic labeling technique was used to circumvent the need for full manual annotation by pathologists. The developed system achieved a high agreement with the reference standard. In a separate observer experiment, the deep learning system outperformed 10 out of 15 pathologists. The system has the potential to improve prostate cancer prognostics by acting as a first or second reader.
Training convolutional neural networks with megapixel images
Apr 16, 2018
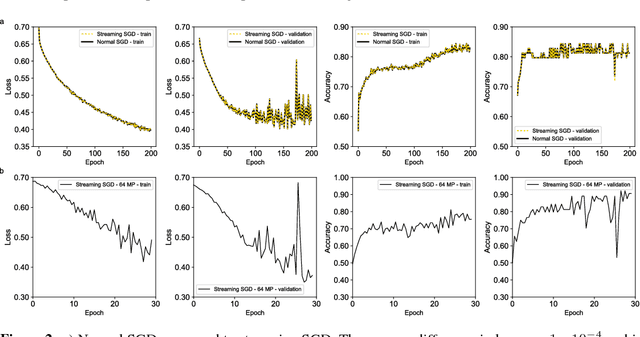
Abstract:To train deep convolutional neural networks, the input data and the intermediate activations need to be kept in memory to calculate the gradient descent step. Given the limited memory available in the current generation accelerator cards, this limits the maximum dimensions of the input data. We demonstrate a method to train convolutional neural networks holding only parts of the image in memory while giving equivalent results. We quantitatively compare this new way of training convolutional neural networks with conventional training. In addition, as a proof of concept, we train a convolutional neural network with 64 megapixel images, which requires 97% less memory than the conventional approach.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge