Hanns-Christian Breit
TotalSegmentator: robust segmentation of 104 anatomical structures in CT images
Aug 11, 2022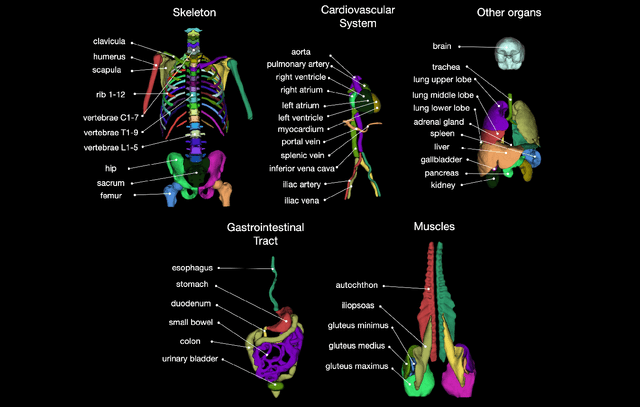
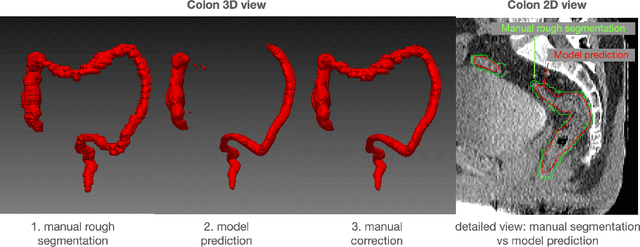
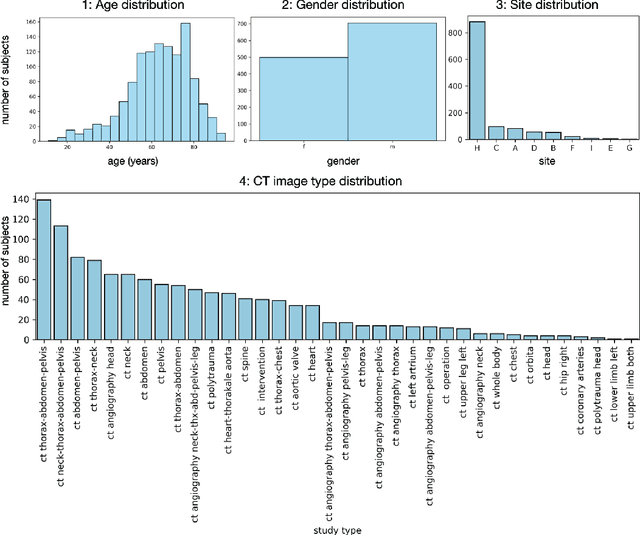

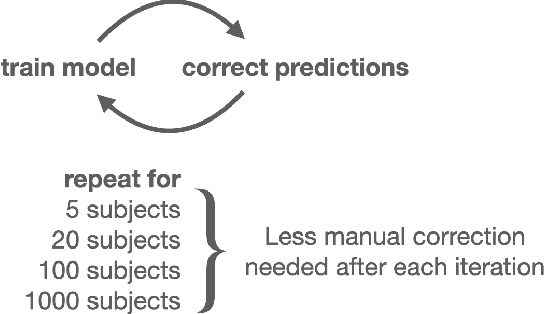
Abstract:In this work we focus on automatic segmentation of multiple anatomical structures in (whole body) CT images. Many segmentation algorithms exist for this task. However, in most cases they suffer from 3 problems: 1. They are difficult to use (the code and data is not publicly available or difficult to use). 2. They do not generalize (often the training dataset was curated to only contain very clean images which do not reflect the image distribution found during clinical routine), 3. The algorithm can only segment one anatomical structure. For more structures several algorithms have to be used which increases the effort required to set up the system. In this work we publish a new dataset and segmentation toolkit which solves all three of these problems: In 1204 CT images we segmented 104 anatomical structures (27 organs, 59 bones, 10 muscles, 8 vessels) covering a majority of relevant classes for most use cases. We show an improved workflow for the creation of ground truth segmentations which speeds up the process by over 10x. The CT images were randomly sampled from clinical routine, thus representing a real world dataset which generalizes to clinical application. The dataset contains a wide range of different pathologies, scanners, sequences and sites. Finally, we train a segmentation algorithm on this new dataset. We call this algorithm TotalSegmentator and make it easily available as a pretrained python pip package (pip install totalsegmentator). Usage is as simple as TotalSegmentator -i ct.nii.gz -o seg and it works well for most CT images. The code is available at https://github.com/wasserth/TotalSegmentator and the dataset at https://doi.org/10.5281/zenodo.6802613.
Improved Segmentation and Detection Sensitivity of Diffusion-Weighted Brain Infarct Lesions with Synthetically Enhanced Deep Learning
Dec 29, 2020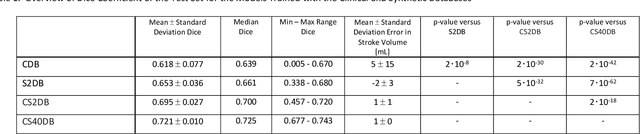
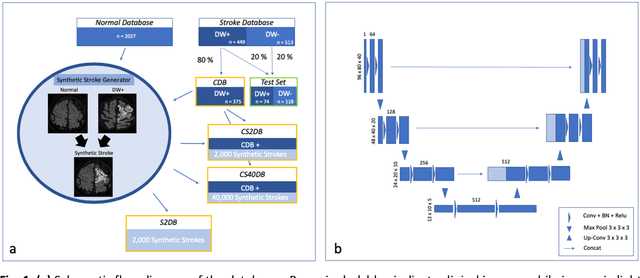
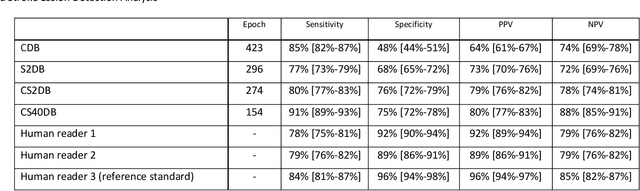
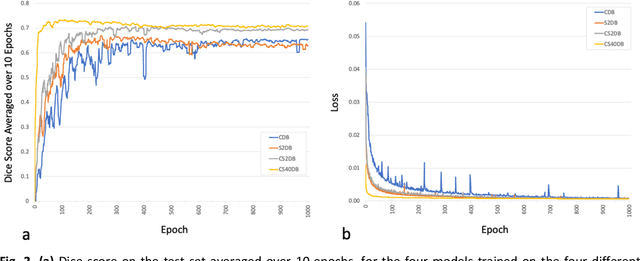
Abstract:Purpose: To compare the segmentation and detection performance of a deep learning model trained on a database of human-labelled clinical diffusion-weighted (DW) stroke lesions to a model trained on the same database enhanced with synthetic DW stroke lesions. Methods: In this institutional review board approved study, a stroke database of 962 cases (mean age 65+/-17 years, 255 males, 449 scans with DW positive stroke lesions) and a normal database of 2,027 patients (mean age 38+/-24 years,1088 females) were obtained. Brain volumes with synthetic DW stroke lesions were produced by warping the relative signal increase of real strokes to normal brain volumes. A generic 3D U-Net was trained on four different databases to generate four different models: (a) 375 neuroradiologist-labeled clinical DW positive stroke cases(CDB);(b) 2,000 synthetic cases(S2DB);(c) CDB+2,000 synthetic cases(CS2DB); or (d) CDB+40,000 synthetic cases(CS40DB). The models were tested on 20%(n=192) of the cases of the stroke database, which were excluded from the training set. Segmentation accuracy was characterized using Dice score and lesion volume of the stroke segmentation, and statistical significance was tested using a paired, two-tailed, Student's t-test. Detection sensitivity and specificity was compared to three neuroradiologists. Results: The performance of the 3D U-Net model trained on the CS40DB(mean Dice 0.72) was better than models trained on the CS2DB (0.70,P <0.001) or the CDB(0.65,P<0.001). The deep learning model was also more sensitive (91%[89%-93%]) than each of the three human readers(84%[81%-87%],78%[75%-81%],and 79%[76%-82%]), but less specific(75%[72%-78%] vs for the three human readers (96%[94%-97%],92%[90%-94%] and 89%[86%-91%]). Conclusion: Deep learning training for segmentation and detection of DW stroke lesions was significantly improved by enhancing the training set with synthetic lesions.
* This manuscript has been accepted for publication in Radiology: Artificial Intelligence
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge