Haiming Li
OMGs: A multi-agent system supporting MDT decision-making across the ovarian tumour care continuum
Feb 14, 2026Abstract:Ovarian tumour management has increasingly relied on multidisciplinary tumour board (MDT) deliberation to address treatment complexity and disease heterogeneity. However, most patients worldwide lack access to timely expert consensus, particularly in resource-constrained centres where MDT resources are scarce or unavailable. Here we present OMGs (Ovarian tumour Multidisciplinary intelligent aGent System), a multi-agent AI framework where domain-specific agents deliberate collaboratively to integrate multidisciplinary evidence and generate MDT-style recommendations with transparent rationales. To systematically evaluate MDT recommendation quality, we developed SPEAR (Safety, Personalization, Evidence, Actionability, Robustness) and validated OMGs across diverse clinical scenarios spanning the care continuum. In multicentre re-evaluation, OMGs achieved performance comparable to expert MDT consensus ($4.45 \pm 0.30$ versus $4.53 \pm 0.23$), with higher Evidence scores (4.57 versus 3.92). In prospective multicentre evaluation (59 patients), OMGs demonstrated high concordance with routine MDT decisions. Critically, in paired human-AI studies, OMGs most substantially enhanced clinicians' recommendations in Evidence and Robustness, the dimensions most compromised when multidisciplinary expertise is unavailable. These findings suggest that multi-agent deliberative systems can achieve performance comparable to expert MDT consensus, with potential to expand access to specialized oncology expertise in resource-limited settings.
WSSS4LUAD: Grand Challenge on Weakly-supervised Tissue Semantic Segmentation for Lung Adenocarcinoma
Apr 14, 2022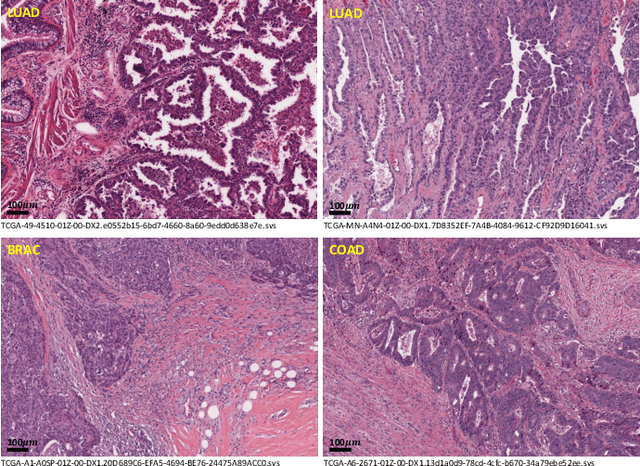
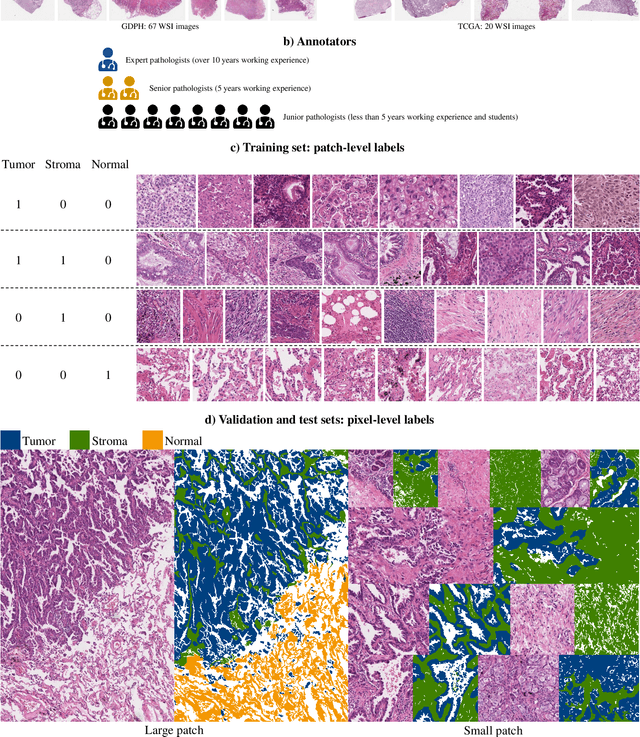
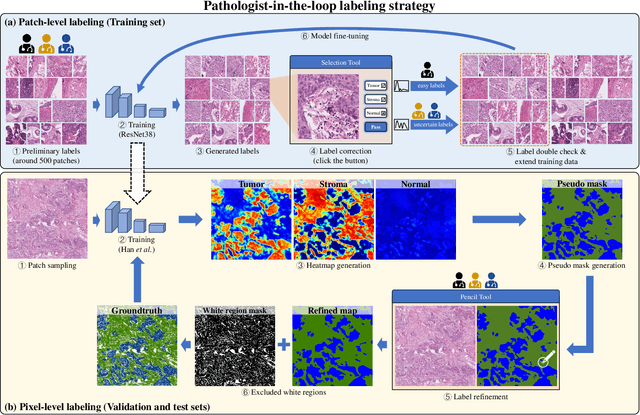
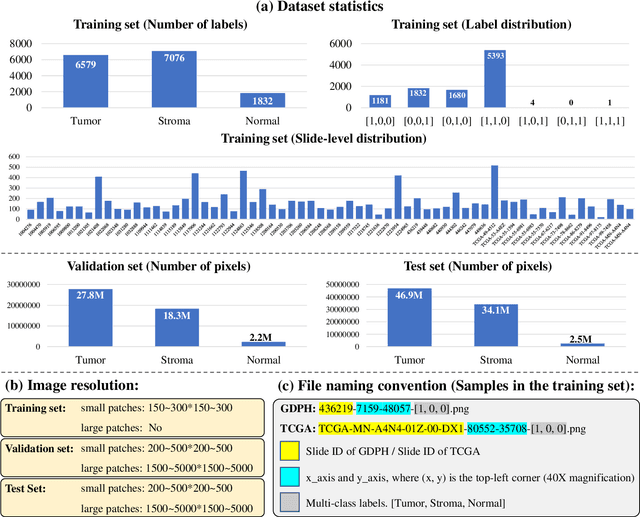
Abstract:Lung cancer is the leading cause of cancer death worldwide, and adenocarcinoma (LUAD) is the most common subtype. Exploiting the potential value of the histopathology images can promote precision medicine in oncology. Tissue segmentation is the basic upstream task of histopathology image analysis. Existing deep learning models have achieved superior segmentation performance but require sufficient pixel-level annotations, which is time-consuming and expensive. To enrich the label resources of LUAD and to alleviate the annotation efforts, we organize this challenge WSSS4LUAD to call for the outstanding weakly-supervised semantic segmentation (WSSS) techniques for histopathology images of LUAD. Participants have to design the algorithm to segment tumor epithelial, tumor-associated stroma and normal tissue with only patch-level labels. This challenge includes 10,091 patch-level annotations (the training set) and over 130 million labeled pixels (the validation and test sets), from 87 WSIs (67 from GDPH, 20 from TCGA). All the labels were generated by a pathologist-in-the-loop pipeline with the help of AI models and checked by the label review board. Among 532 registrations, 28 teams submitted the results in the test phase with over 1,000 submissions. Finally, the first place team achieved mIoU of 0.8413 (tumor: 0.8389, stroma: 0.7931, normal: 0.8919). According to the technical reports of the top-tier teams, CAM is still the most popular approach in WSSS. Cutmix data augmentation has been widely adopted to generate more reliable samples. With the success of this challenge, we believe that WSSS approaches with patch-level annotations can be a complement to the traditional pixel annotations while reducing the annotation efforts. The entire dataset has been released to encourage more researches on computational pathology in LUAD and more novel WSSS techniques.
PDBL: Improving Histopathological Tissue Classification with Plug-and-Play Pyramidal Deep-Broad Learning
Nov 04, 2021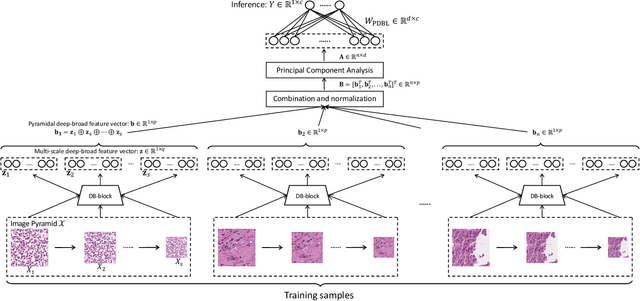
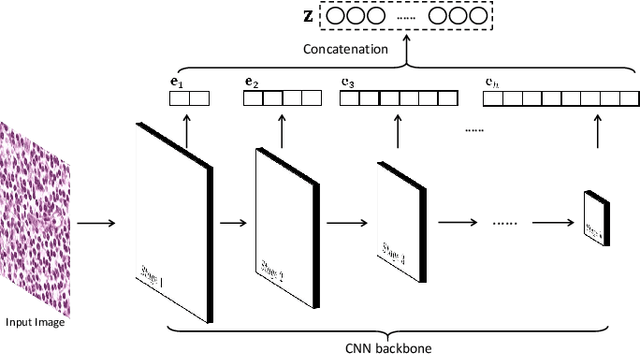
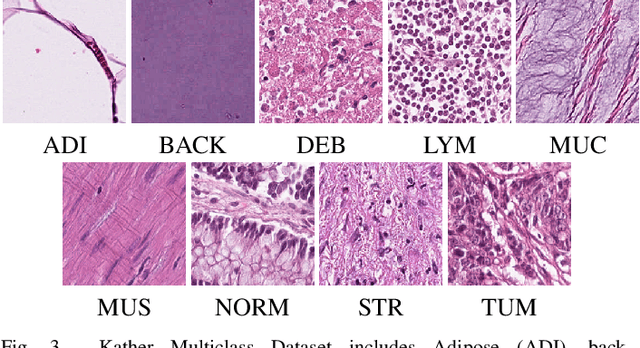
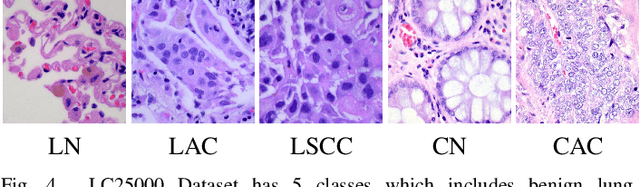
Abstract:Histopathological tissue classification is a fundamental task in pathomics cancer research. Precisely differentiating different tissue types is a benefit for the downstream researches, like cancer diagnosis, prognosis and etc. Existing works mostly leverage the popular classification backbones in computer vision to achieve histopathological tissue classification. In this paper, we proposed a super lightweight plug-and-play module, named Pyramidal Deep-Broad Learning (PDBL), for any well-trained classification backbone to further improve the classification performance without a re-training burden. We mimic how pathologists observe pathology slides in different magnifications and construct an image pyramid for the input image in order to obtain the pyramidal contextual information. For each level in the pyramid, we extract the multi-scale deep-broad features by our proposed Deep-Broad block (DB-block). We equipped PDBL in three popular classification backbones, ShuffLeNetV2, EfficientNetb0, and ResNet50 to evaluate the effectiveness and efficiency of our proposed module on two datasets (Kather Multiclass Dataset and the LC25000 Dataset). Experimental results demonstrate the proposed PDBL can steadily improve the tissue-level classification performance for any CNN backbones, especially for the lightweight models when given a small among of training samples (less than 10%), which greatly saves the computational time and annotation efforts.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge