Gerson Zaverucha
Neural-Symbolic Learning and Reasoning: A Survey and Interpretation
Nov 10, 2017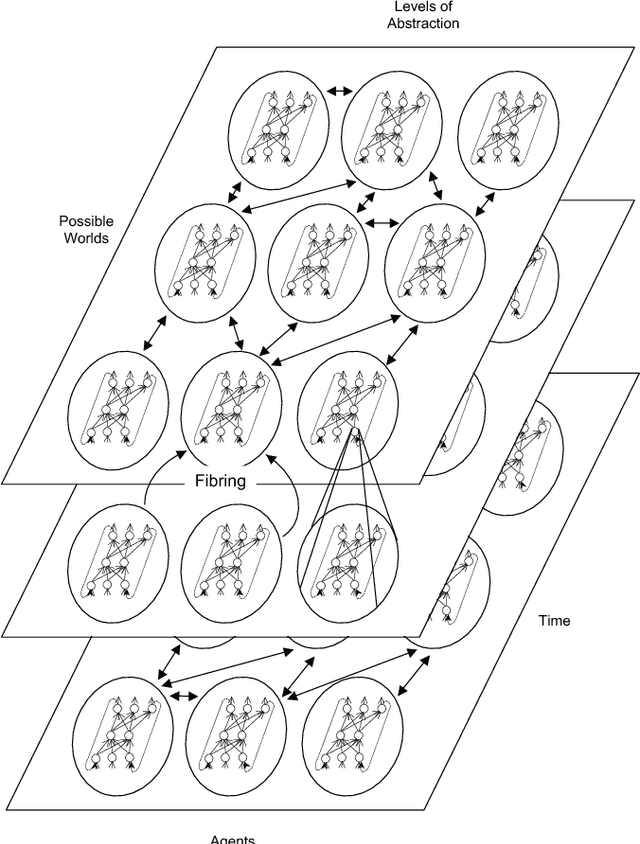

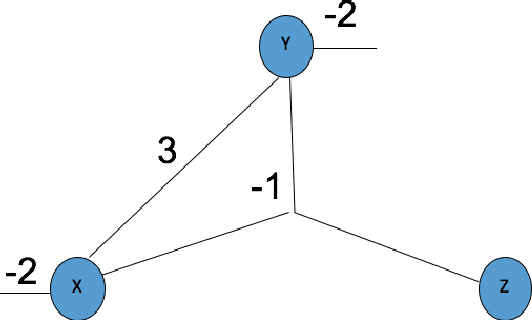
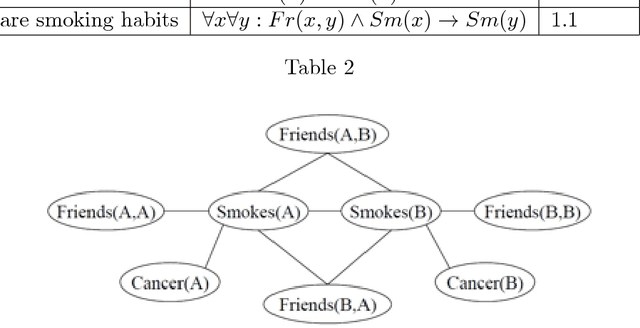
Abstract:The study and understanding of human behaviour is relevant to computer science, artificial intelligence, neural computation, cognitive science, philosophy, psychology, and several other areas. Presupposing cognition as basis of behaviour, among the most prominent tools in the modelling of behaviour are computational-logic systems, connectionist models of cognition, and models of uncertainty. Recent studies in cognitive science, artificial intelligence, and psychology have produced a number of cognitive models of reasoning, learning, and language that are underpinned by computation. In addition, efforts in computer science research have led to the development of cognitive computational systems integrating machine learning and automated reasoning. Such systems have shown promise in a range of applications, including computational biology, fault diagnosis, training and assessment in simulators, and software verification. This joint survey reviews the personal ideas and views of several researchers on neural-symbolic learning and reasoning. The article is organised in three parts: Firstly, we frame the scope and goals of neural-symbolic computation and have a look at the theoretical foundations. We then proceed to describe the realisations of neural-symbolic computation, systems, and applications. Finally we present the challenges facing the area and avenues for further research.
A study of structural properties on profiles HMMs
Dec 11, 2008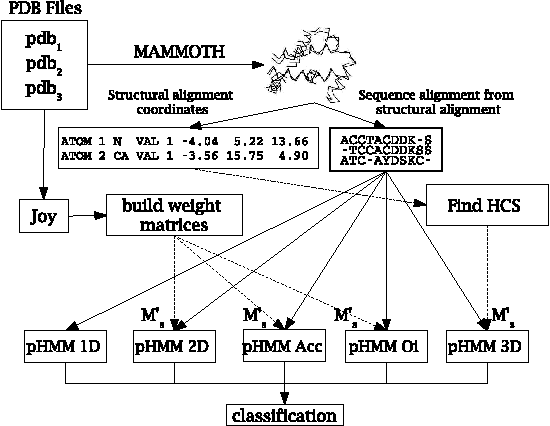
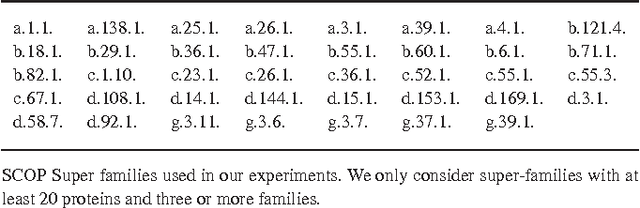
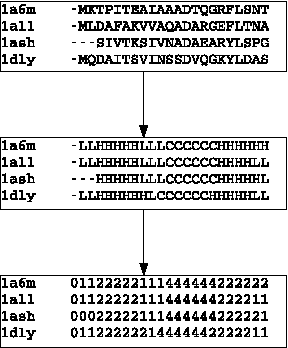
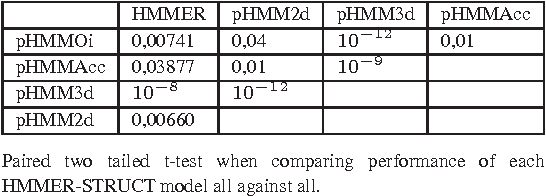
Abstract:Motivation: Profile hidden Markov Models (pHMMs) are a popular and very useful tool in the detection of the remote homologue protein families. Unfortunately, their performance is not always satisfactory when proteins are in the 'twilight zone'. We present HMMER-STRUCT, a model construction algorithm and tool that tries to improve pHMM performance by using structural information while training pHMMs. As a first step, HMMER-STRUCT constructs a set of pHMMs. Each pHMM is constructed by weighting each residue in an aligned protein according to a specific structural property of the residue. Properties used were primary, secondary and tertiary structures, accessibility and packing. HMMER-STRUCT then prioritizes the results by voting. Results: We used the SCOP database to perform our experiments. Throughout, we apply leave-one-family-out cross-validation over protein superfamilies. First, we used the MAMMOTH-mult structural aligner to align the training set proteins. Then, we performed two sets of experiments. In a first experiment, we compared structure weighted models against standard pHMMs and against each other. In a second experiment, we compared the voting model against individual pHMMs. We compare method performance through ROC curves and through Precision/Recall curves, and assess significance through the paired two tailed t-test. Our results show significant performance improvements of all structurally weighted models over default HMMER, and a significant improvement in sensitivity of the combined models over both the original model and the structurally weighted models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge