Dzhoshkun Ismail Shakir
Zero-shot super-resolution with a physically-motivated downsampling kernel for endomicroscopy
Mar 25, 2021
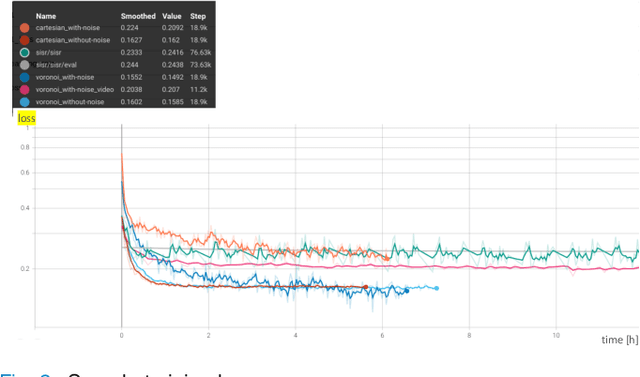
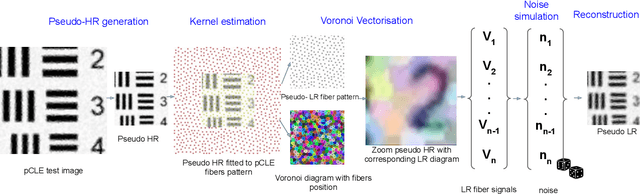
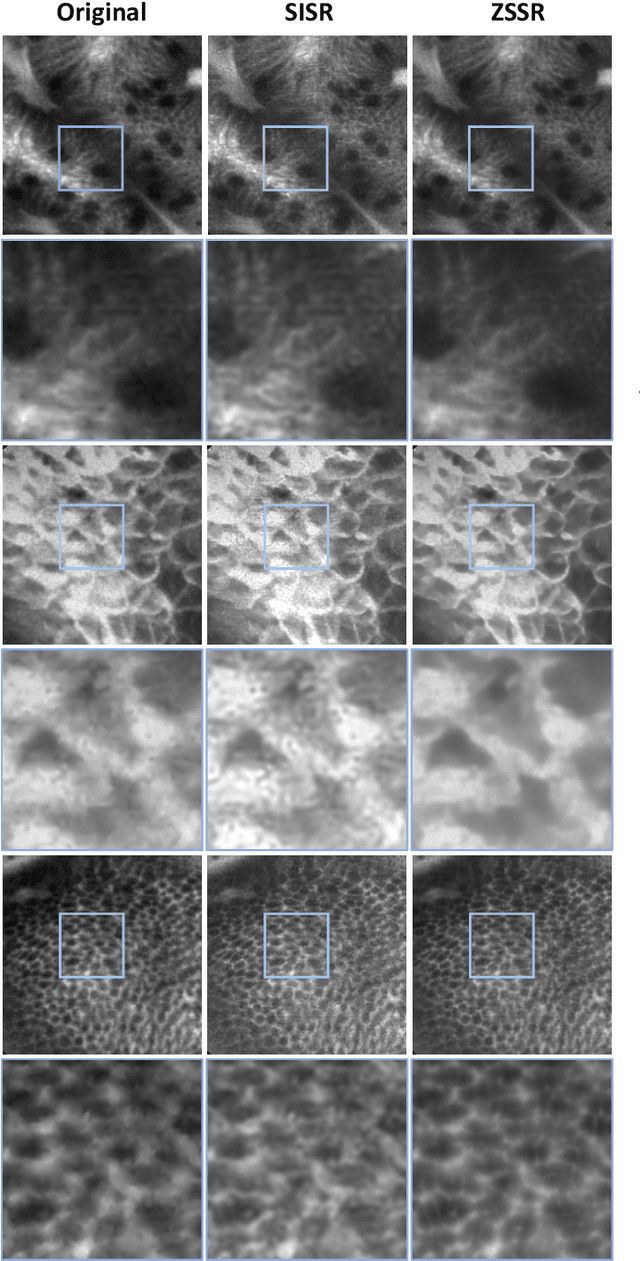
Abstract:Super-resolution (SR) methods have seen significant advances thanks to the development of convolutional neural networks (CNNs). CNNs have been successfully employed to improve the quality of endomicroscopy imaging. Yet, the inherent limitation of research on SR in endomicroscopy remains the lack of ground truth high-resolution (HR) images, commonly used for both supervised training and reference-based image quality assessment (IQA). Therefore, alternative methods, such as unsupervised SR are being explored. To address the need for non-reference image quality improvement, we designed a novel zero-shot super-resolution (ZSSR) approach that relies only on the endomicroscopy data to be processed in a self-supervised manner without the need for ground-truth HR images. We tailored the proposed pipeline to the idiosyncrasies of endomicroscopy by introducing both: a physically-motivated Voronoi downscaling kernel accounting for the endomicroscope's irregular fibre-based sampling pattern, and realistic noise patterns. We also took advantage of video sequences to exploit a sequence of images for self-supervised zero-shot image quality improvement. We run ablation studies to assess our contribution in regards to the downscaling kernel and noise simulation. We validate our methodology on both synthetic and original data. Synthetic experiments were assessed with reference-based IQA, while our results for original images were evaluated in a user study conducted with both expert and non-expert observers. The results demonstrated superior performance in image quality of ZSSR reconstructions in comparison to the baseline method. The ZSSR is also competitive when compared to supervised single-image SR, especially being the preferred reconstruction technique by experts.
Active Annotation of Informative Overlapping Frames in Video Mosaicking Applications
Dec 30, 2020
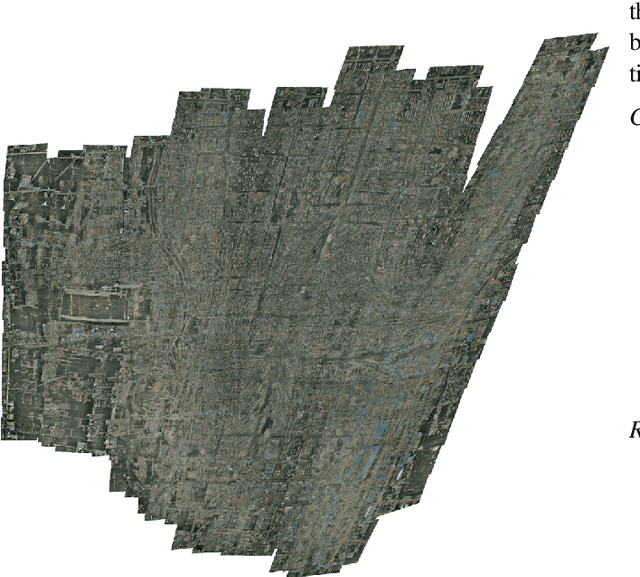
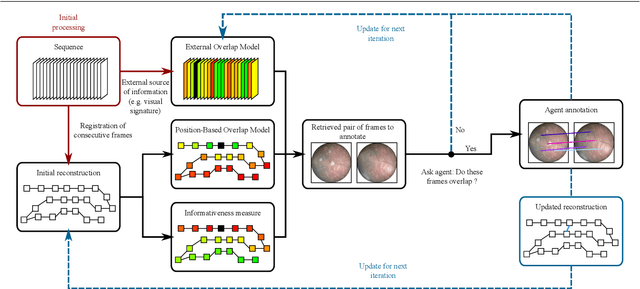

Abstract:Video mosaicking requires the registration of overlapping frames located at distant timepoints in the sequence to ensure global consistency of the reconstructed scene. However, fully automated registration of such long-range pairs is (i) challenging when the registration of images itself is difficult; and (ii) computationally expensive for long sequences due to the large number of candidate pairs for registration. In this paper, we introduce an efficient framework for the active annotation of long-range pairwise correspondences in a sequence. Our framework suggests pairs of images that are sought to be informative to an oracle agent (e.g., a human user, or a reliable matching algorithm) who provides visual correspondences on each suggested pair. Informative pairs are retrieved according to an iterative strategy based on a principled annotation reward coupled with two complementary and online adaptable models of frame overlap. In addition to the efficient construction of a mosaic, our framework provides, as a by-product, ground truth landmark correspondences which can be used for evaluation or learning purposes. We evaluate our approach in both automated and interactive scenarios via experiments on synthetic sequences, on a publicly available dataset for aerial imaging and on a clinical dataset for placenta mosaicking during fetal surgery.
Learning from Irregularly Sampled Data for Endomicroscopy Super-resolution: A Comparative Study of Sparse and Dense Approaches
Nov 29, 2019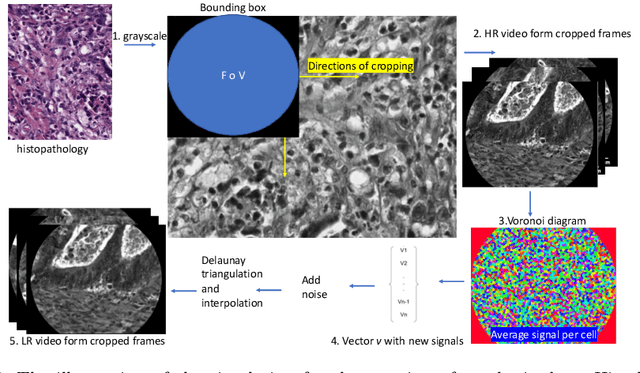
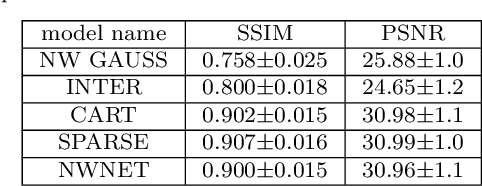
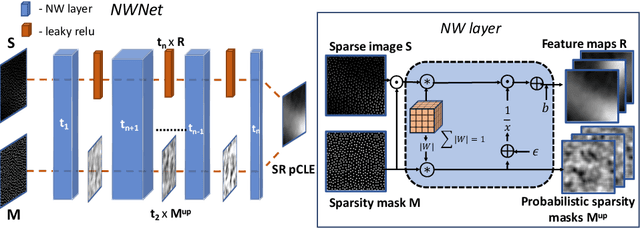
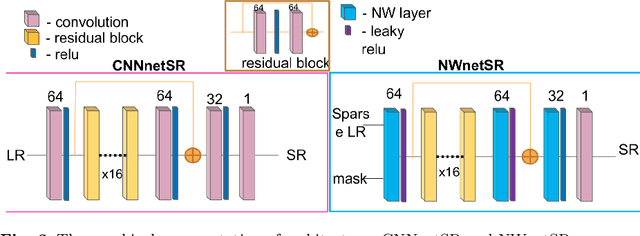
Abstract:Purpose: Probe-based Confocal Laser Endomicroscopy (pCLE) enables performing an optical biopsy, providing real-time microscopic images, via a probe. pCLE probes consist of multiple optical fibres arranged in a bundle, which taken together generate signals in an irregularly sampled pattern. Current pCLE reconstruction is based on interpolating irregular signals onto an over-sampled Cartesian grid, using a naive linear interpolation. It was shown that Convolutional Neural Networks (CNNs) could improve pCLE image quality. Although classical CNNs were applied to pCLE, input data were limited to reconstructed images in contrast to irregular data produced by pCLE. Methods: We compare pCLE reconstruction and super-resolution (SR) methods taking irregularly sampled or reconstructed pCLE images as input. We also propose to embed a Nadaraya-Watson (NW) kernel regression into the CNN framework as a novel trainable CNN layer. Using the NW layer and exemplar-based super-resolution, we design an NWNetSR architecture that allows for reconstructing high-quality pCLE images directly from the irregularly sampled input data. We created synthetic sparse pCLE images to evaluate our methodology. Results: The results were validated through an image quality assessment based on a combination of the following metrics: Peak signal-to-noise ratio, the Structural Similarity Index. Conclusion: Both dense and sparse CNNs outperform the reconstruction method currently used in the clinic. The main contributions of our study are a comparison of sparse and dense approach in pCLE image reconstruction, implementing trainable generalised NW kernel regression, and adaptation of synthetic data for training pCLE SR.
Effective deep learning training for single-image super-resolution in endomicroscopy exploiting video-registration-based reconstruction
Mar 23, 2018



Abstract:Purpose: Probe-based Confocal Laser Endomicroscopy (pCLE) is a recent imaging modality that allows performing in vivo optical biopsies. The design of pCLE hardware, and its reliance on an optical fibre bundle, fundamentally limits the image quality with a few tens of thousands fibres, each acting as the equivalent of a single-pixel detector, assembled into a single fibre bundle. Video-registration techniques can be used to estimate high-resolution (HR) images by exploiting the temporal information contained in a sequence of low-resolution (LR) images. However, the alignment of LR frames, required for the fusion, is computationally demanding and prone to artefacts. Methods: In this work, we propose a novel synthetic data generation approach to train exemplar-based Deep Neural Networks (DNNs). HR pCLE images with enhanced quality are recovered by the models trained on pairs of estimated HR images (generated by the video-registration algorithm) and realistic synthetic LR images. Performance of three different state-of-the-art DNNs techniques were analysed on a Smart Atlas database of 8806 images from 238 pCLE video sequences. The results were validated through an extensive Image Quality Assessment (IQA) that takes into account different quality scores, including a Mean Opinion Score (MOS). Results: Results indicate that the proposed solution produces an effective improvement in the quality of the obtained reconstructed image. Conclusion: The proposed training strategy and associated DNNs allows us to perform convincing super-resolution of pCLE images.
Retrieval and Registration of Long-Range Overlapping Frames for Scalable Mosaicking of In Vivo Fetoscopy
Feb 28, 2018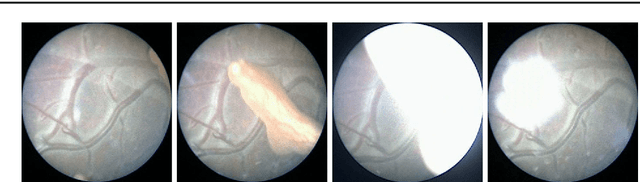
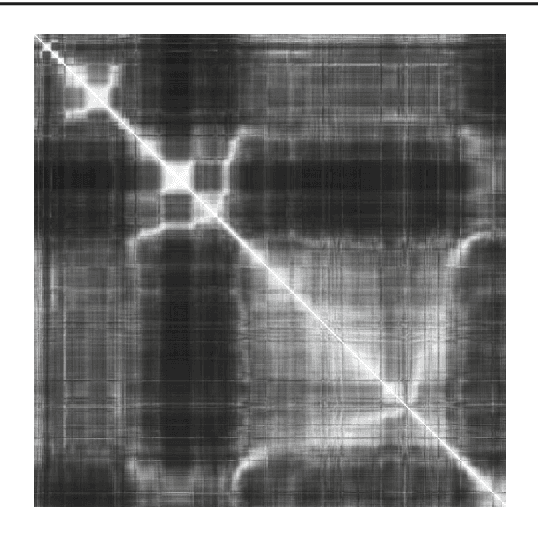


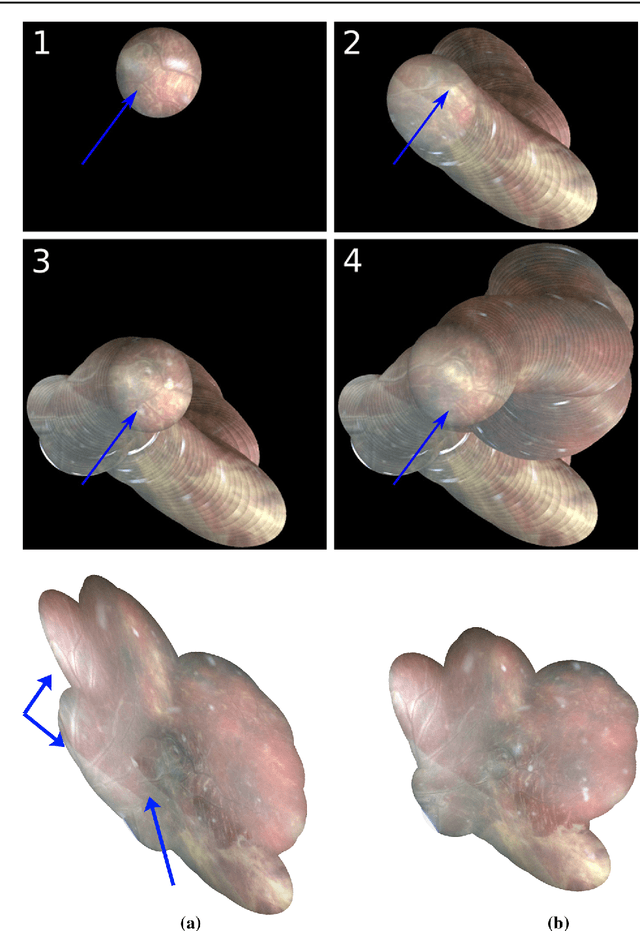
Abstract:Purpose: The standard clinical treatment of Twin-to-Twin Transfusion Syndrome consists in the photo-coagulation of undesired anastomoses located on the placenta which are responsible to a blood transfer between the two twins. While being the standard of care procedure, fetoscopy suffers from a limited field-of-view of the placenta resulting in missed anastomoses. To facilitate the task of the clinician, building a global map of the placenta providing a larger overview of the vascular network is highly desired. Methods: To overcome the challenging visual conditions inherent to in vivo sequences (low contrast, obstructions or presence of artifacts, among others), we propose the following contributions: (i) robust pairwise registration is achieved by aligning the orientation of the image gradients, and (ii) difficulties regarding long-range consistency (e.g. due to the presence of outliers) is tackled via a bag-of-word strategy, which identifies overlapping frames of the sequence to be registered regardless of their respective location in time. Results: In addition to visual difficulties, in vivo sequences are characterised by the intrinsic absence of gold standard. We present mosaics motivating qualitatively our methodological choices and demonstrating their promising aspect. We also demonstrate semi-quantitatively, via visual inspection of registration results, the efficacy of our registration approach in comparison to two standard baselines. Conclusion: This paper proposes the first approach for the construction of mosaics of placenta in in vivo fetoscopy sequences. Robustness to visual challenges during registration and long-range temporal consistency are proposed, offering first positive results on in vivo data for which standard mosaicking techniques are not applicable.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge