Djordje Miladinovic
In-silico biological discovery with large perturbation models
Mar 30, 2025
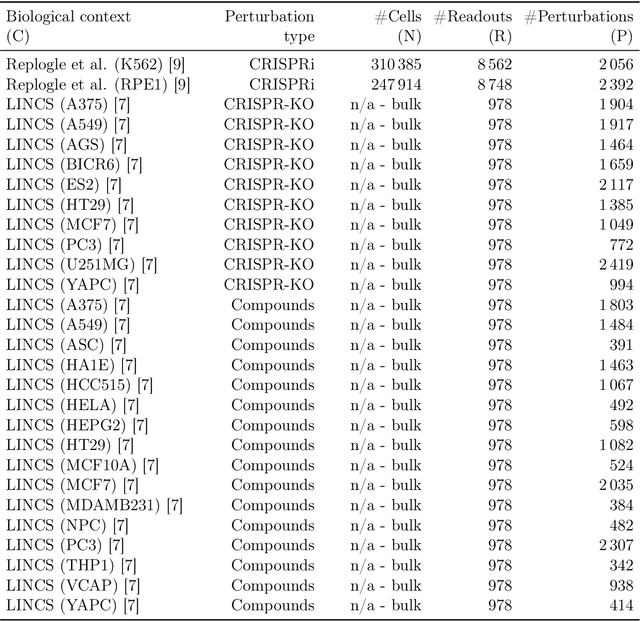
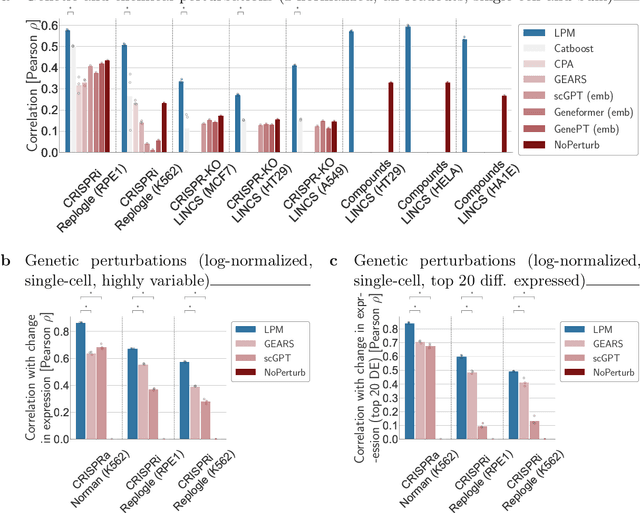
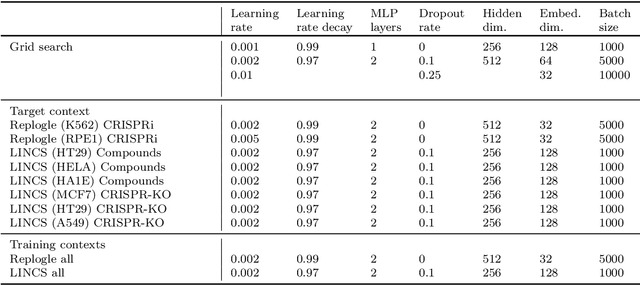
Abstract:Data generated in perturbation experiments link perturbations to the changes they elicit and therefore contain information relevant to numerous biological discovery tasks -- from understanding the relationships between biological entities to developing therapeutics. However, these data encompass diverse perturbations and readouts, and the complex dependence of experimental outcomes on their biological context makes it challenging to integrate insights across experiments. Here, we present the Large Perturbation Model (LPM), a deep-learning model that integrates multiple, heterogeneous perturbation experiments by representing perturbation, readout, and context as disentangled dimensions. LPM outperforms existing methods across multiple biological discovery tasks, including in predicting post-perturbation transcriptomes of unseen experiments, identifying shared molecular mechanisms of action between chemical and genetic perturbations, and facilitating the inference of gene-gene interaction networks.
Granger-causal Attentive Mixtures of Experts: Learning Important Features with Neural Networks
Oct 01, 2018
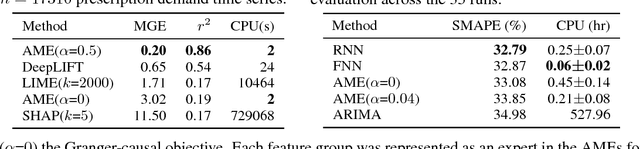
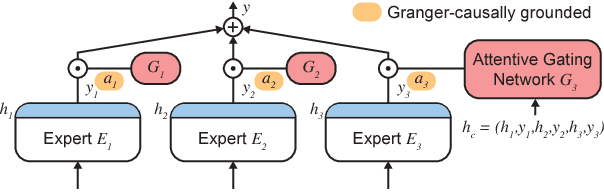
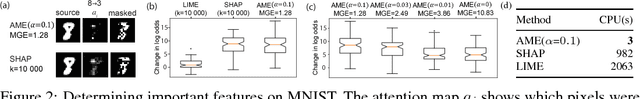

Abstract:Knowledge of the importance of input features towards decisions made by machine-learning models is essential to increase our understanding of both the models and the underlying data. Here, we present a new approach to estimating feature importance with neural networks based on the idea of distributing the features of interest among experts in an attentive mixture of experts (AME). AMEs use attentive gating networks trained with a Granger-causal objective to learn to jointly produce accurate predictions as well as estimates of feature importance in a single model. Our experiments on an established benchmark and two real-world datasets show (i) that the feature importance estimates provided by AMEs compare favourably to those provided by state-of-the-art methods, (ii) that AMEs are significantly faster than existing methods, and (iii) that the associations discovered by AMEs are consistent with those reported by domain experts.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge