Dietmar Frey
CLAIM - Charite Lab for AI in Medicine, Charite Universitatsmedizin Berlin, corporate member of Freie Universitat Berlin and Humboldt Universitat zu Berlin, Berlin, Germany, Department of Neurosurgery, Charite Universitatsmedizin Berlin, corporate member of Freie Universitat Berlin and Humboldt Universitat zu Berlin, Berlin, Germany
RELICT: A Replica Detection Framework for Medical Image Generation
Feb 24, 2025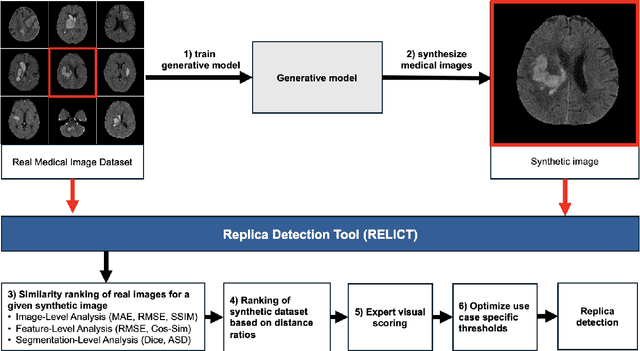
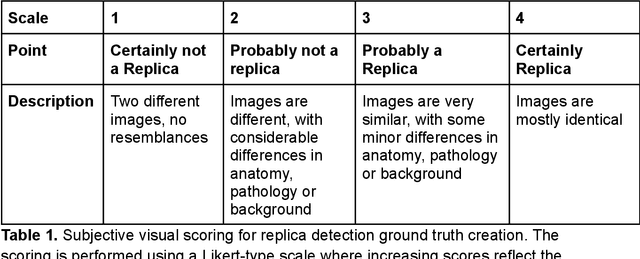
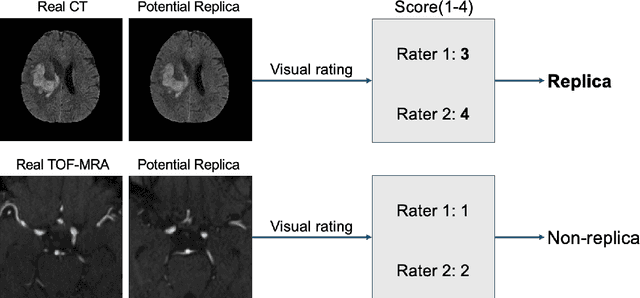
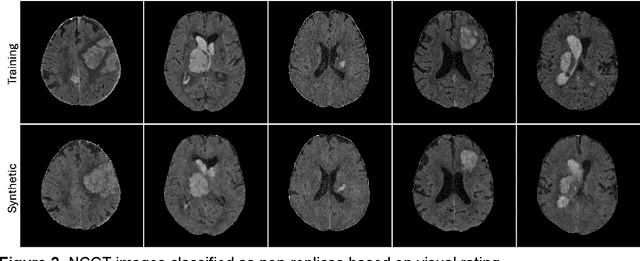
Abstract:Despite the potential of synthetic medical data for augmenting and improving the generalizability of deep learning models, memorization in generative models can lead to unintended leakage of sensitive patient information and limit model utility. Thus, the use of memorizing generative models in the medical domain can jeopardize patient privacy. We propose a framework for identifying replicas, i.e. nearly identical copies of the training data, in synthetic medical image datasets. Our REpLIca deteCTion (RELICT) framework for medical image generative models evaluates image similarity using three complementary approaches: (1) voxel-level analysis, (2) feature-level analysis by a pretrained medical foundation model, and (3) segmentation-level analysis. Two clinically relevant 3D generative modelling use cases were investigated: non-contrast head CT with intracerebral hemorrhage (N=774) and time-of-flight MR angiography of the Circle of Willis (N=1,782). Expert visual scoring was used as the reference standard to assess the presence of replicas. We report the balanced accuracy at the optimal threshold to assess replica classification performance. The reference visual rating identified 45 of 50 and 5 of 50 generated images as replicas for the NCCT and TOF-MRA use cases, respectively. Image-level and feature-level measures perfectly classified replicas with a balanced accuracy of 1 when an optimal threshold was selected for the NCCT use case. A perfect classification of replicas for the TOF-MRA case was not possible at any threshold, with the segmentation-level analysis achieving a balanced accuracy of 0.79. Replica detection is a crucial but neglected validation step for the development of generative models in medical imaging. The proposed RELICT framework provides a standardized, easy-to-use tool for replica detection and aims to facilitate responsible and ethical medical image synthesis.
Cross-modality image synthesis from TOF-MRA to CTA using diffusion-based models
Sep 16, 2024



Abstract:Cerebrovascular disease often requires multiple imaging modalities for accurate diagnosis, treatment, and monitoring. Computed Tomography Angiography (CTA) and Time-of-Flight Magnetic Resonance Angiography (TOF-MRA) are two common non-invasive angiography techniques, each with distinct strengths in accessibility, safety, and diagnostic accuracy. While CTA is more widely used in acute stroke due to its faster acquisition times and higher diagnostic accuracy, TOF-MRA is preferred for its safety, as it avoids radiation exposure and contrast agent-related health risks. Despite the predominant role of CTA in clinical workflows, there is a scarcity of open-source CTA data, limiting the research and development of AI models for tasks such as large vessel occlusion detection and aneurysm segmentation. This study explores diffusion-based image-to-image translation models to generate synthetic CTA images from TOF-MRA input. We demonstrate the modality conversion from TOF-MRA to CTA and show that diffusion models outperform a traditional U-Net-based approach. Our work compares different state-of-the-art diffusion architectures and samplers, offering recommendations for optimal model performance in this cross-modality translation task.
From Single-Hospital to Multi-Centre Applications: Enhancing the Generalisability of Deep Learning Models for Adverse Event Prediction in the ICU
Apr 07, 2023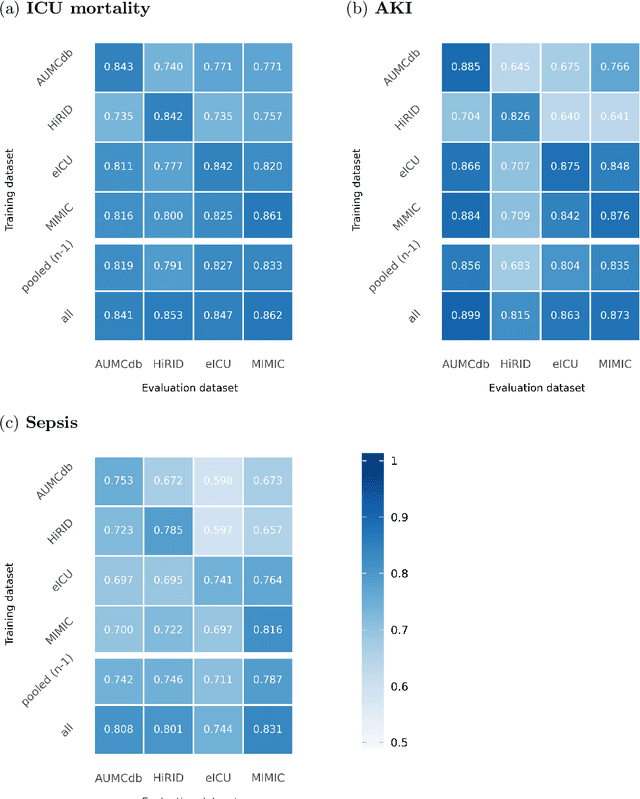

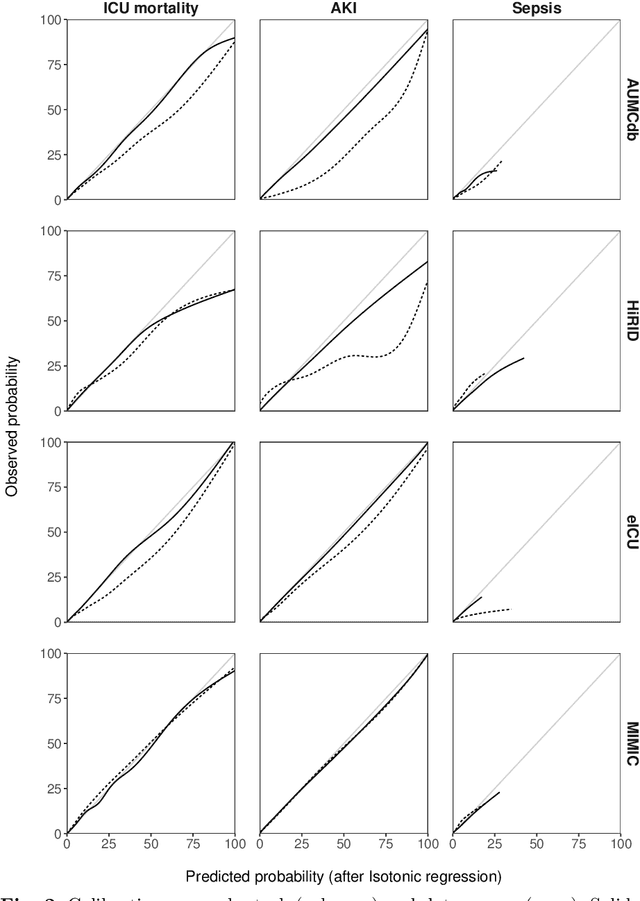
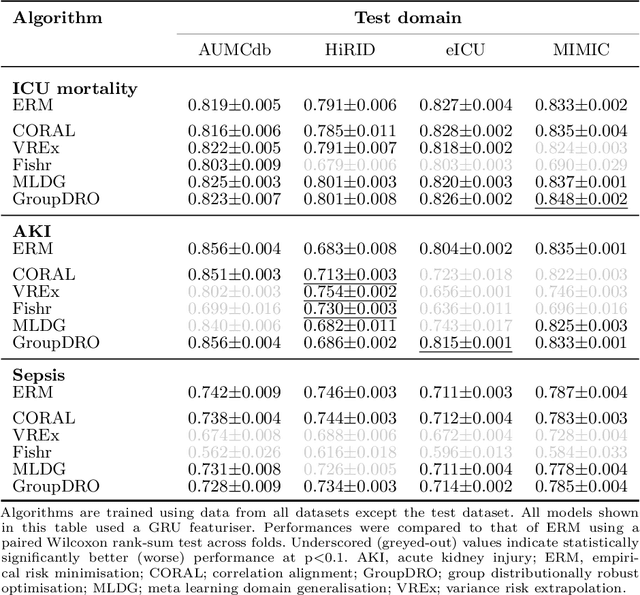
Abstract:Deep learning (DL) can aid doctors in detecting worsening patient states early, affording them time to react and prevent bad outcomes. While DL-based early warning models usually work well in the hospitals they were trained for, they tend to be less reliable when applied at new hospitals. This makes it difficult to deploy them at scale. Using carefully harmonised intensive care data from four data sources across Europe and the US (totalling 334,812 stays), we systematically assessed the reliability of DL models for three common adverse events: death, acute kidney injury (AKI), and sepsis. We tested whether using more than one data source and/or explicitly optimising for generalisability during training improves model performance at new hospitals. We found that models achieved high AUROC for mortality (0.838-0.869), AKI (0.823-0.866), and sepsis (0.749-0.824) at the training hospital. As expected, performance dropped at new hospitals, sometimes by as much as -0.200. Using more than one data source for training mitigated the performance drop, with multi-source models performing roughly on par with the best single-source model. This suggests that as data from more hospitals become available for training, model robustness is likely to increase, lower-bounding robustness with the performance of the most applicable data source in the training data. Dedicated methods promoting generalisability did not noticeably improve performance in our experiments.
Anonymization of labeled TOF-MRA images for brain vessel segmentation using generative adversarial networks
Sep 16, 2020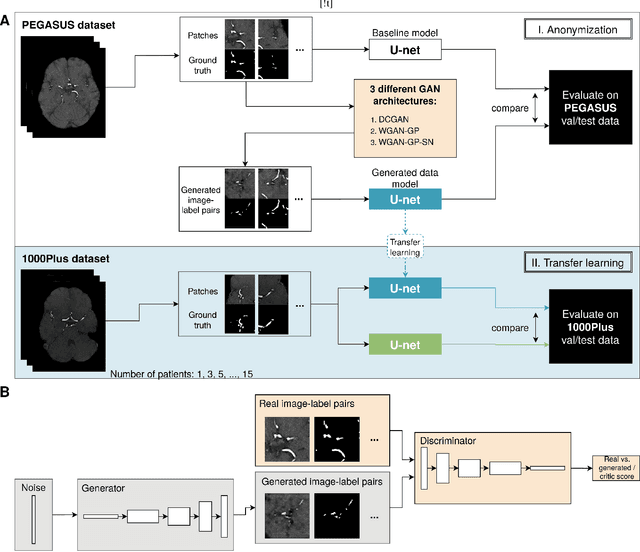
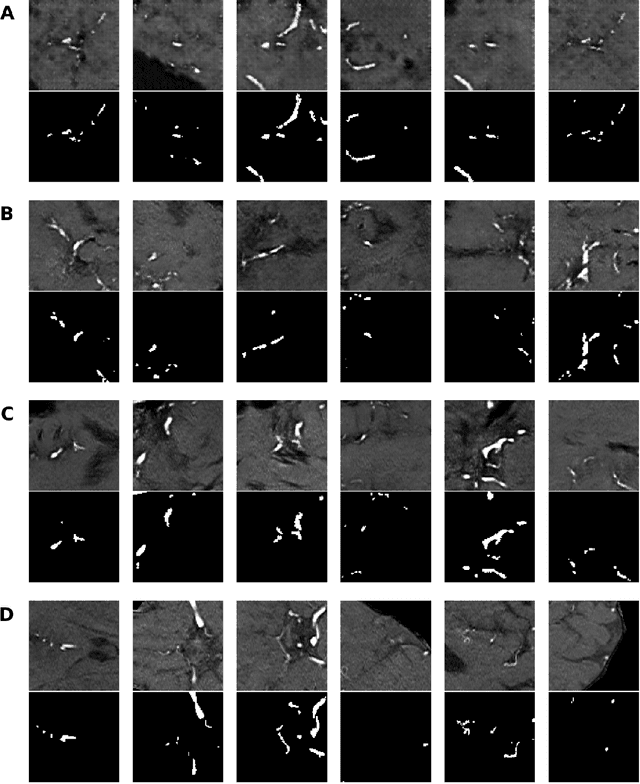
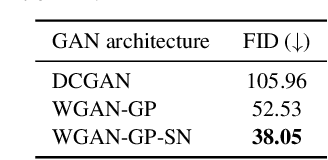

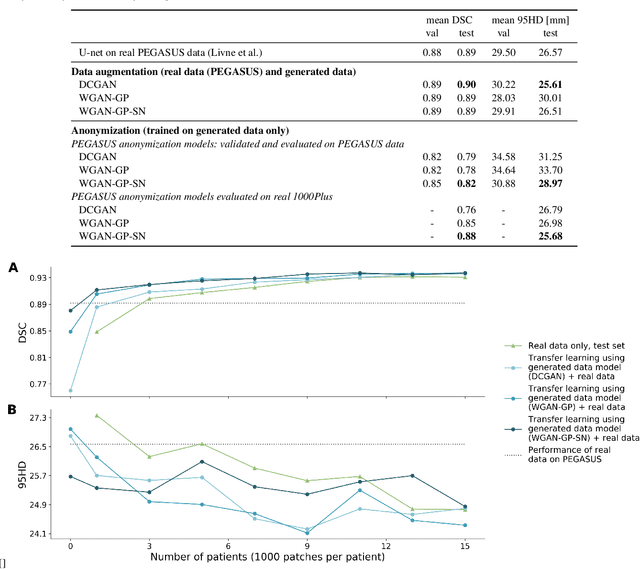
Abstract:Anonymization and data sharing are crucial for privacy protection and acquisition of large datasets for medical image analysis. This is a big challenge, especially for neuroimaging. Here, the brain's unique structure allows for re-identification and thus requires non-conventional anonymization. Generative adversarial networks (GANs) have the potential to provide anonymous images while preserving predictive properties. Analyzing brain vessel segmentation as a use case, we trained 3 GANs on time-of-flight (TOF) magnetic resonance angiography (MRA) patches for image-label generation: 1) Deep convolutional GAN, 2) Wasserstein-GAN with gradient penalty (WGAN-GP) and 3) WGAN-GP with spectral normalization (WGAN-GP-SN). The generated image-labels from each GAN were used to train a U-net for segmentation and tested on real data. Moreover, we applied our synthetic patches using transfer learning on a second dataset. For an increasing number of up to 15 patients we evaluated the model performance on real data with and without pre-training. The performance for all models was assessed by the Dice Similarity Coefficient (DSC) and the 95th percentile of the Hausdorff Distance (95HD). Comparing the 3 GANs, the U-net trained on synthetic data generated by the WGAN-GP-SN showed the highest performance to predict vessels (DSC/95HD 0.82/28.97) benchmarked by the U-net trained on real data (0.89/26.61). The transfer learning approach showed superior performance for the same GAN compared to no pre-training, especially for one patient only (0.91/25.68 vs. 0.85/27.36). In this work, synthetic image-label pairs retained generalizable information and showed good performance for vessel segmentation. Besides, we showed that synthetic patches can be used in a transfer learning approach with independent data. This paves the way to overcome the challenges of scarce data and anonymization in medical imaging.
On The Usage Of Average Hausdorff Distance For Segmentation Performance Assessment: Hidden Bias When Used For Ranking
Sep 13, 2020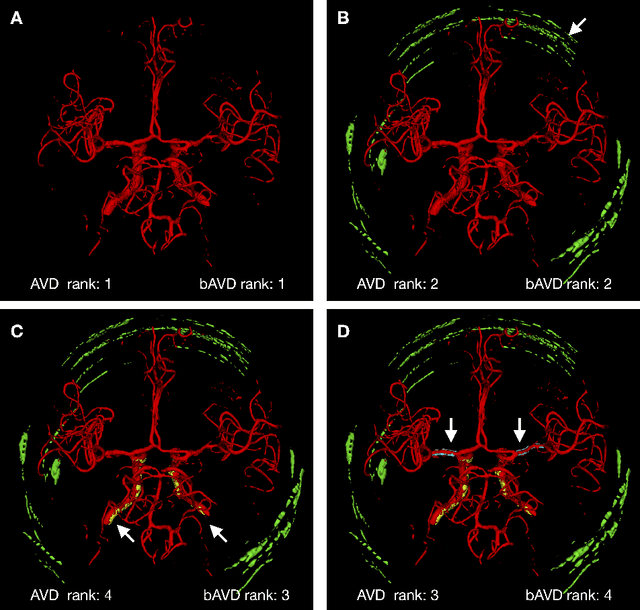
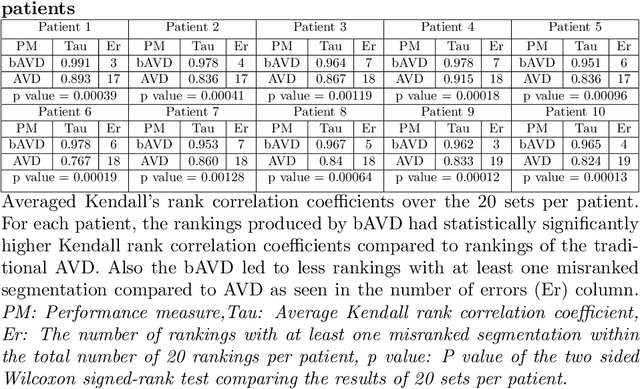
Abstract:Average Hausdorff Distance (AVD) is a widely used performance measure to calculate the distance between two point sets. In medical image segmentation, AVD is used to compare ground truth images with segmentation results allowing their ranking. We identified, however, a ranking bias of AVD making it less suitable for segmentation ranking. To mitigate this bias, we present a modified calculation of AVD that we have coined balanced AVD (bAVD). To simulate segmentations for ranking, we manually created non-overlapping segmentation errors common in cerebral vessel segmentation as our use-case. Adding the created errors consecutively and randomly to the ground truth, we created sets of simulated segmentations with increasing number of errors. Each set of simulated segmentations was ranked using AVD and bAVD. We calculated the Kendall-rank-correlation-coefficient between the segmentation ranking and the number of errors in each simulated segmentation. The rankings produced by bAVD had a significantly higher average correlation (0.969) than those of AVD (0.847). In 200 total rankings, bAVD misranked 52 and AVD misranked 179 segmentations. Our proposed evaluation measure, bAVD, alleviates AVDs ranking bias making it more suitable for rankings and quality assessment of segmentations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge