Chao Cui
GeomCLIP: Contrastive Geometry-Text Pre-training for Molecules
Nov 16, 2024
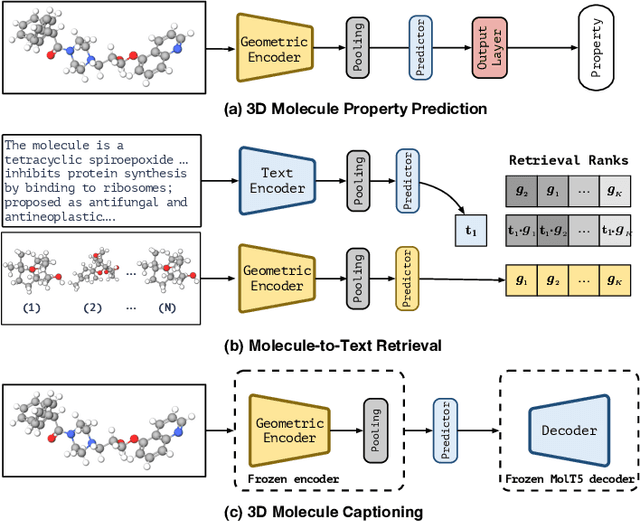
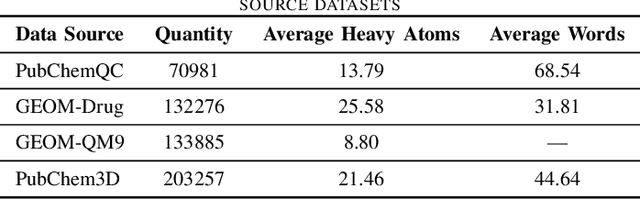
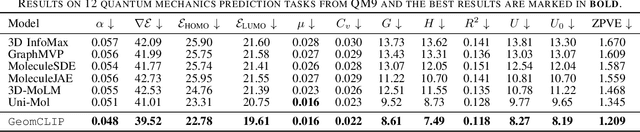
Abstract:Pretraining molecular representations is crucial for drug and material discovery. Recent methods focus on learning representations from geometric structures, effectively capturing 3D position information. Yet, they overlook the rich information in biomedical texts, which detail molecules' properties and substructures. With this in mind, we set up a data collection effort for 200K pairs of ground-state geometric structures and biomedical texts, resulting in a PubChem3D dataset. Based on this dataset, we propose the GeomCLIP framework to enhance for multi-modal representation learning from molecular structures and biomedical text. During pre-training, we design two types of tasks, i.e., multimodal representation alignment and unimodal denoising pretraining, to align the 3D geometric encoder with textual information and, at the same time, preserve its original representation power. Experimental results show the effectiveness of GeomCLIP in various tasks such as molecular property prediction, zero-shot text-molecule retrieval, and 3D molecule captioning. Our code and collected dataset are available at \url{https://github.com/xiaocui3737/GeomCLIP}
How to Leverage Demonstration Data in Alignment for Large Language Model? A Self-Imitation Learning Perspective
Oct 14, 2024Abstract:This paper introduces a novel generalized self-imitation learning ($\textbf{GSIL}$) framework, which effectively and efficiently aligns large language models with offline demonstration data. We develop $\textbf{GSIL}$ by deriving a surrogate objective of imitation learning with density ratio estimates, facilitating the use of self-generated data and optimizing the imitation learning objective with simple classification losses. $\textbf{GSIL}$ eliminates the need for complex adversarial training in standard imitation learning, achieving lightweight and efficient fine-tuning for large language models. In addition, $\textbf{GSIL}$ encompasses a family of offline losses parameterized by a general class of convex functions for density ratio estimation and enables a unified view for alignment with demonstration data. Extensive experiments show that $\textbf{GSIL}$ consistently and significantly outperforms baselines in many challenging benchmarks, such as coding (HuamnEval), mathematical reasoning (GSM8K) and instruction-following benchmark (MT-Bench).
MolBind: Multimodal Alignment of Language, Molecules, and Proteins
Mar 13, 2024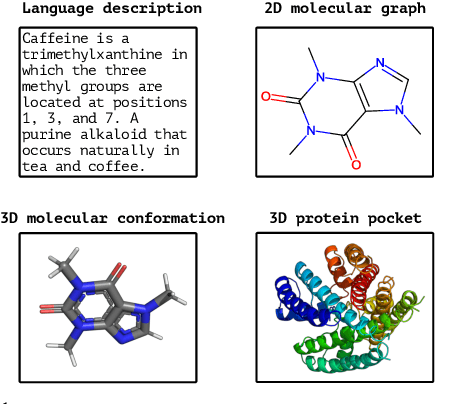
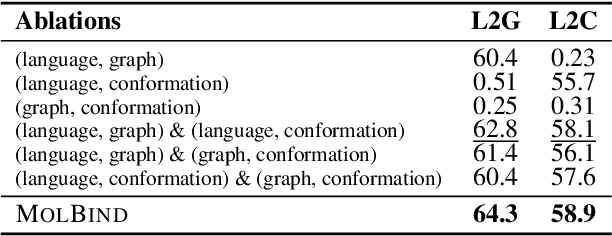
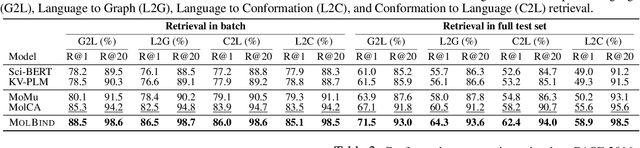
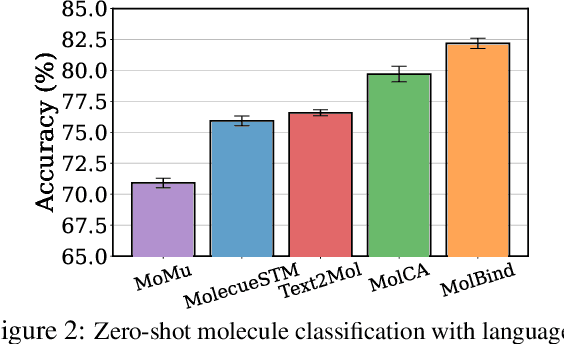

Abstract:Recent advancements in biology and chemistry have leveraged multi-modal learning, integrating molecules and their natural language descriptions to enhance drug discovery. However, current pre-training frameworks are limited to two modalities, and designing a unified network to process different modalities (e.g., natural language, 2D molecular graphs, 3D molecular conformations, and 3D proteins) remains challenging due to inherent gaps among them. In this work, we propose MolBind, a framework that trains encoders for multiple modalities through contrastive learning, mapping all modalities to a shared feature space for multi-modal semantic alignment. To facilitate effective pre-training of MolBind on multiple modalities, we also build and collect a high-quality dataset with four modalities, MolBind-M4, including graph-language, conformation-language, graph-conformation, and conformation-protein paired data. MolBind shows superior zero-shot learning performance across a wide range of tasks, demonstrating its strong capability of capturing the underlying semantics of multiple modalities.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge