Vasant G. Honavar
Skin-R1: Toward Trustworthy Clinical Reasoning for Dermatological Diagnosis
Nov 18, 2025Abstract:The emergence of vision-language models (VLMs) has opened new possibilities for clinical reasoning and has shown promising performance in dermatological diagnosis. However, their trustworthiness and clinical utility are often limited by three major factors: (1) Data heterogeneity, where diverse datasets lack consistent diagnostic labels and clinical concept annotations; (2) Absence of grounded diagnostic rationales, leading to a scarcity of reliable reasoning supervision; and (3) Limited scalability and generalization, as models trained on small, densely annotated datasets struggle to transfer nuanced reasoning to large, sparsely-annotated ones. To address these limitations, we propose SkinR1, a novel dermatological VLM that combines deep, textbook-based reasoning with the broad generalization capabilities of reinforcement learning (RL). SkinR1 systematically resolves the key challenges through a unified, end-to-end framework. First, we design a textbook-based reasoning generator that synthesizes high-fidelity, hierarchy-aware, and differential-diagnosis (DDx)-informed trajectories, providing reliable expert-level supervision. Second, we leverage the constructed trajectories for supervised fine-tuning (SFT) empowering the model with grounded reasoning ability. Third, we develop a novel RL paradigm that, by incorporating the hierarchical structure of diseases, effectively transfers these grounded reasoning patterns to large-scale, sparse data. Extensive experiments on multiple dermatology datasets demonstrate that SkinR1 achieves superior diagnostic accuracy. The ablation study demonstrates the importance of the reasoning foundation instilled by SFT.
GeomCLIP: Contrastive Geometry-Text Pre-training for Molecules
Nov 16, 2024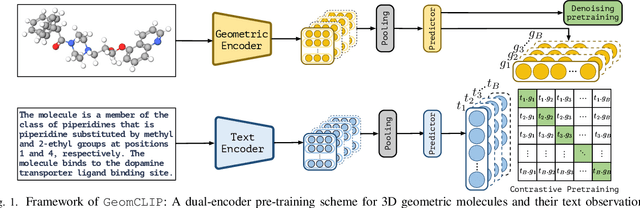
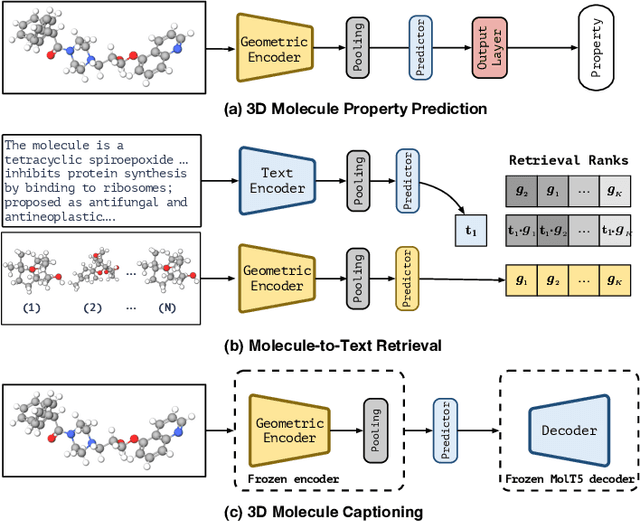
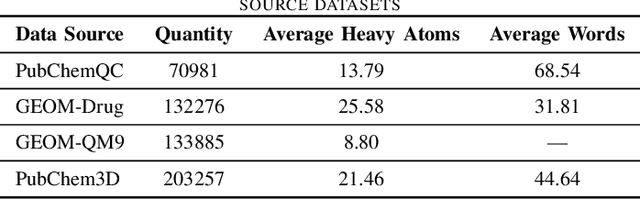
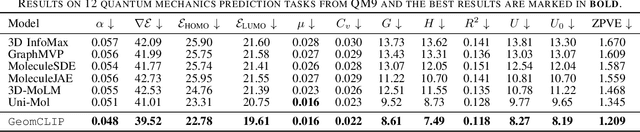
Abstract:Pretraining molecular representations is crucial for drug and material discovery. Recent methods focus on learning representations from geometric structures, effectively capturing 3D position information. Yet, they overlook the rich information in biomedical texts, which detail molecules' properties and substructures. With this in mind, we set up a data collection effort for 200K pairs of ground-state geometric structures and biomedical texts, resulting in a PubChem3D dataset. Based on this dataset, we propose the GeomCLIP framework to enhance for multi-modal representation learning from molecular structures and biomedical text. During pre-training, we design two types of tasks, i.e., multimodal representation alignment and unimodal denoising pretraining, to align the 3D geometric encoder with textual information and, at the same time, preserve its original representation power. Experimental results show the effectiveness of GeomCLIP in various tasks such as molecular property prediction, zero-shot text-molecule retrieval, and 3D molecule captioning. Our code and collected dataset are available at \url{https://github.com/xiaocui3737/GeomCLIP}
MolBind: Multimodal Alignment of Language, Molecules, and Proteins
Mar 13, 2024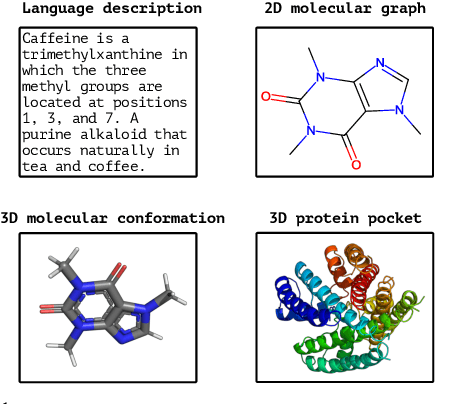
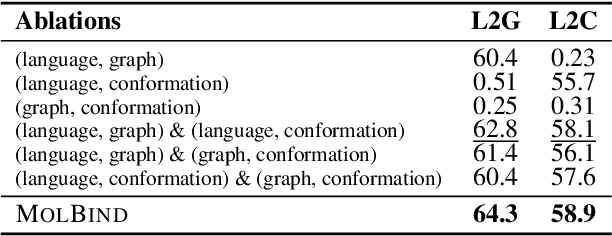
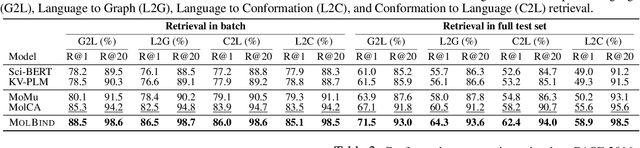
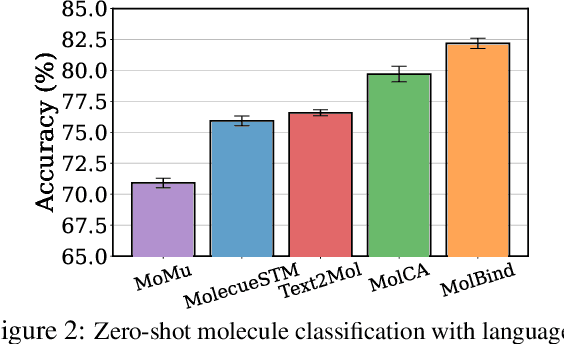

Abstract:Recent advancements in biology and chemistry have leveraged multi-modal learning, integrating molecules and their natural language descriptions to enhance drug discovery. However, current pre-training frameworks are limited to two modalities, and designing a unified network to process different modalities (e.g., natural language, 2D molecular graphs, 3D molecular conformations, and 3D proteins) remains challenging due to inherent gaps among them. In this work, we propose MolBind, a framework that trains encoders for multiple modalities through contrastive learning, mapping all modalities to a shared feature space for multi-modal semantic alignment. To facilitate effective pre-training of MolBind on multiple modalities, we also build and collect a high-quality dataset with four modalities, MolBind-M4, including graph-language, conformation-language, graph-conformation, and conformation-protein paired data. MolBind shows superior zero-shot learning performance across a wide range of tasks, demonstrating its strong capability of capturing the underlying semantics of multiple modalities.
Commuting Network Spillovers and COVID-19 Deaths Across US Counties
Oct 02, 2020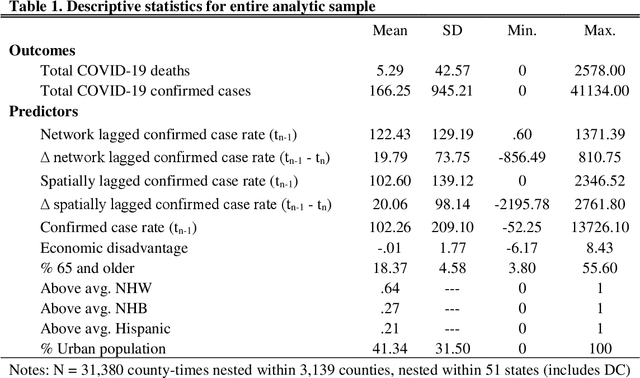
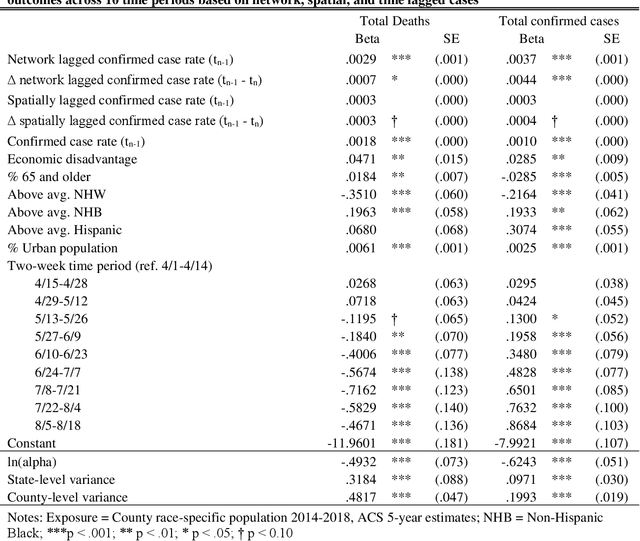
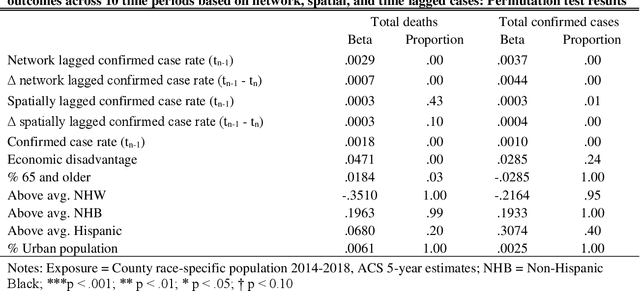
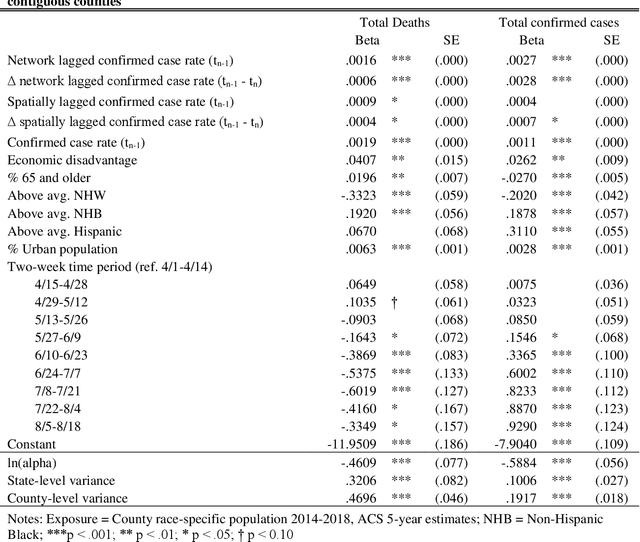
Abstract:This study explored how population mobility flows form commuting networks across US counties and influence the spread of COVID-19. We utilized 3-level mixed effects negative binomial regression models to estimate the impact of network COVID-19 exposure on county confirmed cases and deaths over time. We also conducted weighting-based analyses to estimate the causal effect of network exposure. Results showed that commuting networks matter for COVID-19 deaths and cases, net of spatial proximity, socioeconomic, and demographic factors. Different local racial and ethnic concentrations are also associated with unequal outcomes. These findings suggest that commuting is an important causal mechanism in the spread of COVID-19 and highlight the significance of interconnected of communities. The results suggest that local level mitigation and prevention efforts are more effective when complemented by similar efforts in the network of connected places. Implications for research on inequality in health and flexible work arrangements are discussed.
Accelerating Science: A Computing Research Agenda
Apr 06, 2016Abstract:The emergence of "big data" offers unprecedented opportunities for not only accelerating scientific advances but also enabling new modes of discovery. Scientific progress in many disciplines is increasingly enabled by our ability to examine natural phenomena through the computational lens, i.e., using algorithmic or information processing abstractions of the underlying processes; and our ability to acquire, share, integrate and analyze disparate types of data. However, there is a huge gap between our ability to acquire, store, and process data and our ability to make effective use of the data to advance discovery. Despite successful automation of routine aspects of data management and analytics, most elements of the scientific process currently require considerable human expertise and effort. Accelerating science to keep pace with the rate of data acquisition and data processing calls for the development of algorithmic or information processing abstractions, coupled with formal methods and tools for modeling and simulation of natural processes as well as major innovations in cognitive tools for scientists, i.e., computational tools that leverage and extend the reach of human intellect, and partner with humans on a broad range of tasks in scientific discovery (e.g., identifying, prioritizing formulating questions, designing, prioritizing and executing experiments designed to answer a chosen question, drawing inferences and evaluating the results, and formulating new questions, in a closed-loop fashion). This calls for concerted research agenda aimed at: Development, analysis, integration, sharing, and simulation of algorithmic or information processing abstractions of natural processes, coupled with formal methods and tools for their analyses and simulation; Innovations in cognitive tools that augment and extend human intellect and partner with humans in all aspects of science.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge