Barbara Lam
Impact of Large Language Model Assistance on Patients Reading Clinical Notes: A Mixed-Methods Study
Jan 17, 2024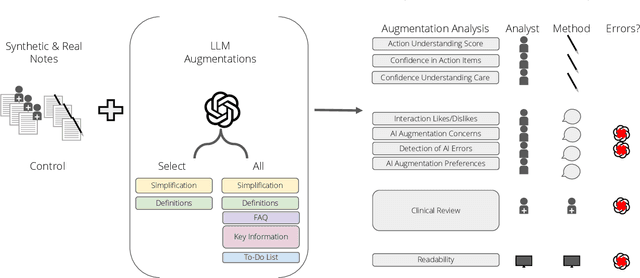
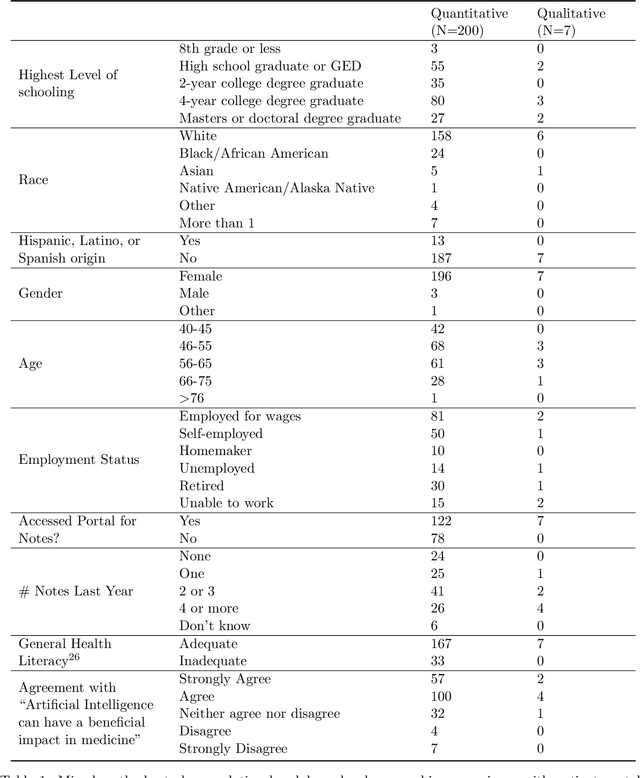
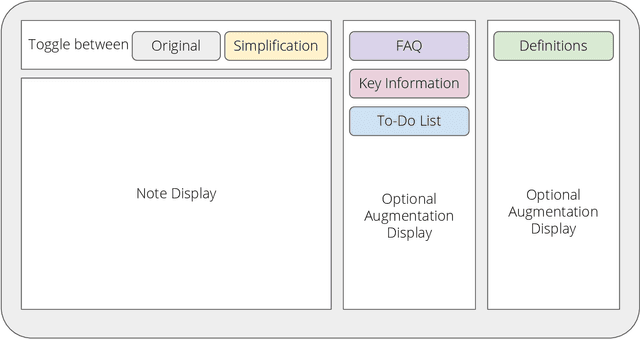
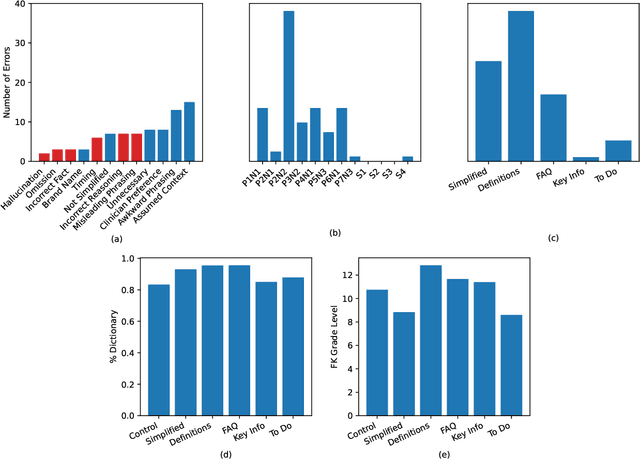
Abstract:Patients derive numerous benefits from reading their clinical notes, including an increased sense of control over their health and improved understanding of their care plan. However, complex medical concepts and jargon within clinical notes hinder patient comprehension and may lead to anxiety. We developed a patient-facing tool to make clinical notes more readable, leveraging large language models (LLMs) to simplify, extract information from, and add context to notes. We prompt engineered GPT-4 to perform these augmentation tasks on real clinical notes donated by breast cancer survivors and synthetic notes generated by a clinician, a total of 12 notes with 3868 words. In June 2023, 200 female-identifying US-based participants were randomly assigned three clinical notes with varying levels of augmentations using our tool. Participants answered questions about each note, evaluating their understanding of follow-up actions and self-reported confidence. We found that augmentations were associated with a significant increase in action understanding score (0.63 $\pm$ 0.04 for select augmentations, compared to 0.54 $\pm$ 0.02 for the control) with p=0.002. In-depth interviews of self-identifying breast cancer patients (N=7) were also conducted via video conferencing. Augmentations, especially definitions, elicited positive responses among the seven participants, with some concerns about relying on LLMs. Augmentations were evaluated for errors by clinicians, and we found misleading errors occur, with errors more common in real donated notes than synthetic notes, illustrating the importance of carefully written clinical notes. Augmentations improve some but not all readability metrics. This work demonstrates the potential of LLMs to improve patients' experience with clinical notes at a lower burden to clinicians. However, having a human in the loop is important to correct potential model errors.
Conceptualizing Machine Learning for Dynamic Information Retrieval of Electronic Health Record Notes
Aug 09, 2023Abstract:The large amount of time clinicians spend sifting through patient notes and documenting in electronic health records (EHRs) is a leading cause of clinician burnout. By proactively and dynamically retrieving relevant notes during the documentation process, we can reduce the effort required to find relevant patient history. In this work, we conceptualize the use of EHR audit logs for machine learning as a source of supervision of note relevance in a specific clinical context, at a particular point in time. Our evaluation focuses on the dynamic retrieval in the emergency department, a high acuity setting with unique patterns of information retrieval and note writing. We show that our methods can achieve an AUC of 0.963 for predicting which notes will be read in an individual note writing session. We additionally conduct a user study with several clinicians and find that our framework can help clinicians retrieve relevant information more efficiently. Demonstrating that our framework and methods can perform well in this demanding setting is a promising proof of concept that they will translate to other clinical settings and data modalities (e.g., labs, medications, imaging).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge