Ashwani Tanwar
FaceFilterSense: A Filter-Resistant Face Recognition and Facial Attribute Analysis Framework
Apr 12, 2024



Abstract:With the advent of social media, fun selfie filters have come into tremendous mainstream use affecting the functioning of facial biometric systems as well as image recognition systems. These filters vary from beautification filters and Augmented Reality (AR)-based filters to filters that modify facial landmarks. Hence, there is a need to assess the impact of such filters on the performance of existing face recognition systems. The limitation associated with existing solutions is that these solutions focus more on the beautification filters. However, the current AR-based filters and filters which distort facial key points are in vogue recently and make the faces highly unrecognizable even to the naked eye. Also, the filters considered are mostly obsolete with limited variations. To mitigate these limitations, we aim to perform a holistic impact analysis of the latest filters and propose an user recognition model with the filtered images. We have utilized a benchmark dataset for baseline images, and applied the latest filters over them to generate a beautified/filtered dataset. Next, we have introduced a model FaceFilterNet for beautified user recognition. In this framework, we also utilize our model to comment on various attributes of the person including age, gender, and ethnicity. In addition, we have also presented a filter-wise impact analysis on face recognition, age estimation, gender, and ethnicity prediction. The proposed method affirms the efficacy of our dataset with an accuracy of 87.25% and an optimal accuracy for facial attribute analysis.
A Scalable Workflow to Build Machine Learning Classifiers with Clinician-in-the-Loop to Identify Patients in Specific Diseases
May 18, 2022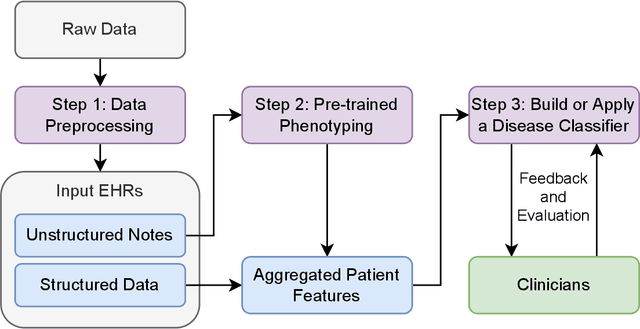

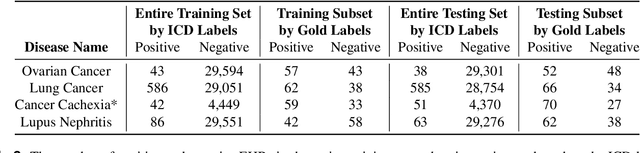
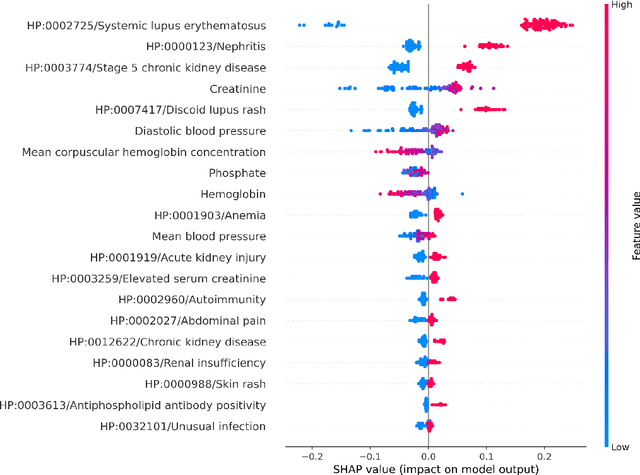
Abstract:Clinicians may rely on medical coding systems such as International Classification of Diseases (ICD) to identify patients with diseases from Electronic Health Records (EHRs). However, due to the lack of detail and specificity as well as a probability of miscoding, recent studies suggest the ICD codes often cannot characterise patients accurately for specific diseases in real clinical practice, and as a result, using them to find patients for studies or trials can result in high failure rates and missing out on uncoded patients. Manual inspection of all patients at scale is not feasible as it is highly costly and slow. This paper proposes a scalable workflow which leverages both structured data and unstructured textual notes from EHRs with techniques including NLP, AutoML and Clinician-in-the-Loop mechanism to build machine learning classifiers to identify patients at scale with given diseases, especially those who might currently be miscoded or missed by ICD codes. Case studies in the MIMIC-III dataset were conducted where the proposed workflow demonstrates a higher classification performance in terms of F1 scores compared to simply using ICD codes on gold testing subset to identify patients with Ovarian Cancer (0.901 vs 0.814), Lung Cancer (0.859 vs 0.828), Cancer Cachexia (0.862 vs 0.650), and Lupus Nephritis (0.959 vs 0.855). Also, the proposed workflow that leverages unstructured notes consistently outperforms the baseline that uses structured data only with an increase of F1 (Ovarian Cancer 0.901 vs 0.719, Lung Cancer 0.859 vs 0.787, Cancer Cachexia 0.862 vs 0.838 and Lupus Nephritis 0.959 vs 0.785). Experiments on the large testing set also demonstrate the proposed workflow can find more patients who are miscoded or missed by ICD codes. Moreover, interpretability studies are also conducted to clinically validate the top impact features of the classifiers.
Unsupervised Numerical Reasoning to Extract Phenotypes from Clinical Text by Leveraging External Knowledge
Apr 19, 2022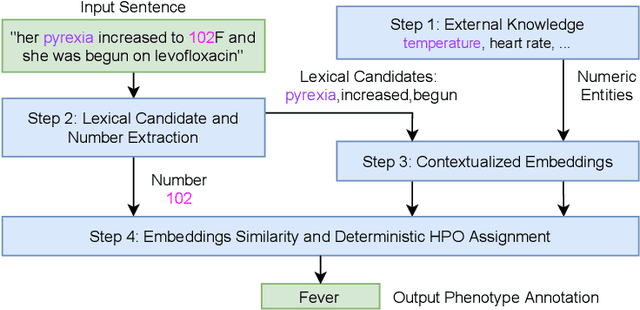
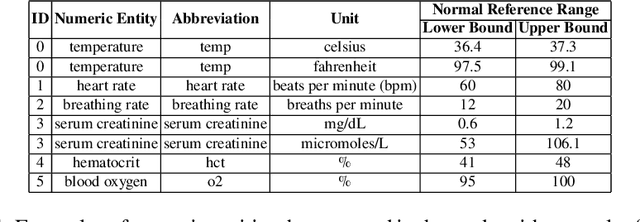
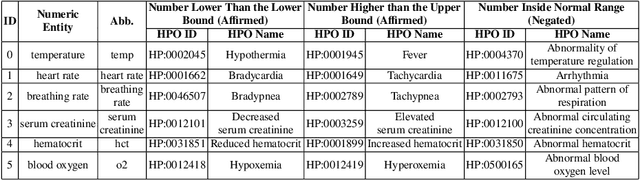
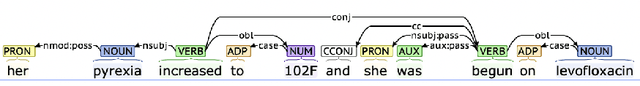
Abstract:Extracting phenotypes from clinical text has been shown to be useful for a variety of clinical use cases such as identifying patients with rare diseases. However, reasoning with numerical values remains challenging for phenotyping in clinical text, for example, temperature 102F representing Fever. Current state-of-the-art phenotyping models are able to detect general phenotypes, but perform poorly when they detect phenotypes requiring numerical reasoning. We present a novel unsupervised methodology leveraging external knowledge and contextualized word embeddings from ClinicalBERT for numerical reasoning in a variety of phenotypic contexts. Comparing against unsupervised benchmarks, it shows a substantial performance improvement with absolute gains on generalized Recall and F1 scores up to 79% and 71%, respectively. In the supervised setting, it also surpasses the performance of alternative approaches with absolute gains on generalized Recall and F1 scores up to 70% and 44%, respectively.
Self-Supervised Detection of Contextual Synonyms in a Multi-Class Setting: Phenotype Annotation Use Case
Sep 04, 2021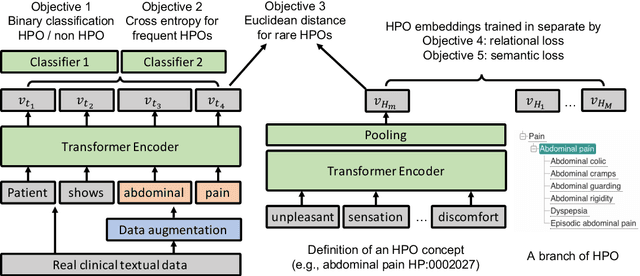
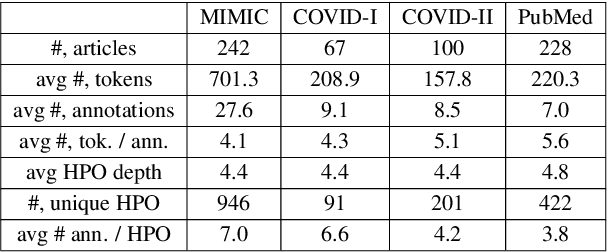
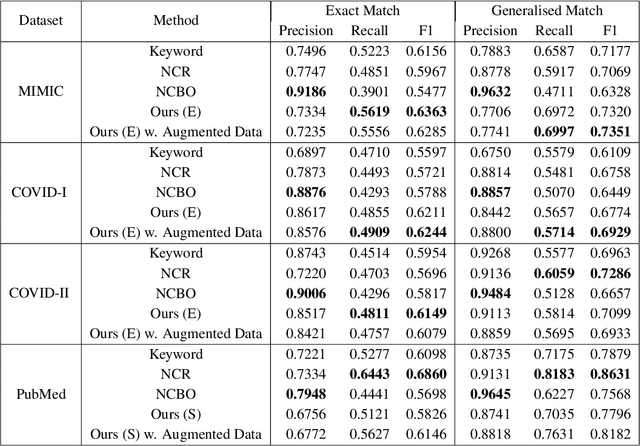
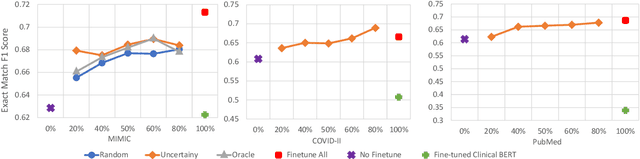
Abstract:Contextualised word embeddings is a powerful tool to detect contextual synonyms. However, most of the current state-of-the-art (SOTA) deep learning concept extraction methods remain supervised and underexploit the potential of the context. In this paper, we propose a self-supervised pre-training approach which is able to detect contextual synonyms of concepts being training on the data created by shallow matching. We apply our methodology in the sparse multi-class setting (over 15,000 concepts) to extract phenotype information from electronic health records. We further investigate data augmentation techniques to address the problem of the class sparsity. Our approach achieves a new SOTA for the unsupervised phenotype concept annotation on clinical text on F1 and Recall outperforming the previous SOTA with a gain of up to 4.5 and 4.0 absolute points, respectively. After fine-tuning with as little as 20\% of the labelled data, we also outperform BioBERT and ClinicalBERT. The extrinsic evaluation on three ICU benchmarks also shows the benefit of using the phenotypes annotated by our model as features.
Clinical Utility of the Automatic Phenotype Annotation in Unstructured Clinical Notes: ICU Use Cases
Jul 24, 2021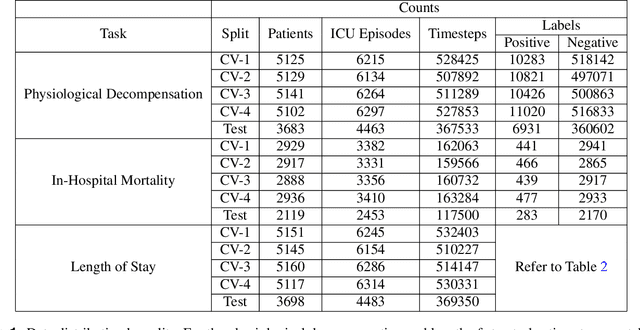
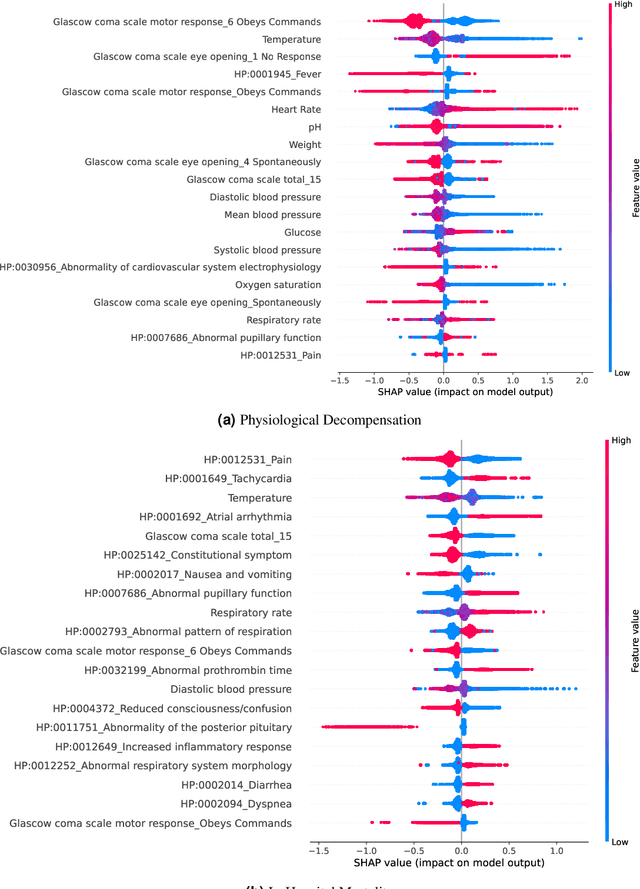
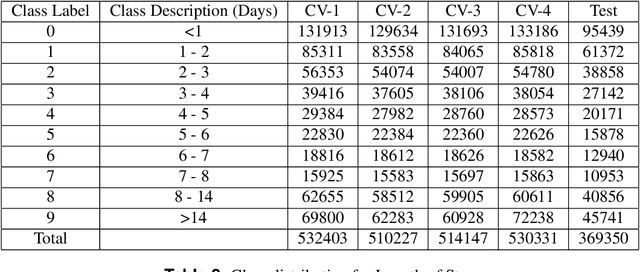
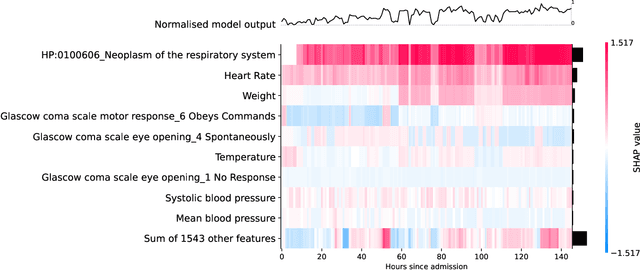
Abstract:Clinical notes contain information not present elsewhere, including drug response and symptoms, all of which are highly important when predicting key outcomes in acute care patients. We propose the automatic annotation of phenotypes from clinical notes as a method to capture essential information to predict outcomes in the Intensive Care Unit (ICU). This information is complementary to typically used vital signs and laboratory test results. We demonstrate and validate our approach conducting experiments on the prediction of in-hospital mortality, physiological decompensation and length of stay in the ICU setting for over 24,000 patients. The prediction models incorporating phenotypic information consistently outperform the baseline models leveraging only vital signs and laboratory test results. Moreover, we conduct a thorough interpretability study, showing that phenotypes provide valuable insights at the patient and cohort levels. Our approach illustrates the viability of using phenotypes to determine outcomes in the ICU.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge