Alison Marsden
SDF4CHD: Generative Modeling of Cardiac Anatomies with Congenital Heart Defects
Nov 08, 2023
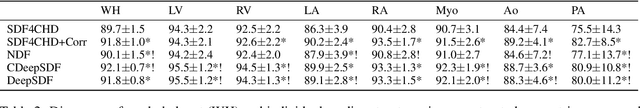
Abstract:Congenital heart disease (CHD) encompasses a spectrum of cardiovascular structural abnormalities, often requiring customized treatment plans for individual patients. Computational modeling and analysis of these unique cardiac anatomies can improve diagnosis and treatment planning and may ultimately lead to improved outcomes. Deep learning (DL) methods have demonstrated the potential to enable efficient treatment planning by automating cardiac segmentation and mesh construction for patients with normal cardiac anatomies. However, CHDs are often rare, making it challenging to acquire sufficiently large patient cohorts for training such DL models. Generative modeling of cardiac anatomies has the potential to fill this gap via the generation of virtual cohorts; however, prior approaches were largely designed for normal anatomies and cannot readily capture the significant topological variations seen in CHD patients. Therefore, we propose a type- and shape-disentangled generative approach suitable to capture the wide spectrum of cardiac anatomies observed in different CHD types and synthesize differently shaped cardiac anatomies that preserve the unique topology for specific CHD types. Our DL approach represents generic whole heart anatomies with CHD type-specific abnormalities implicitly using signed distance fields (SDF) based on CHD type diagnosis, which conveniently captures divergent anatomical variations across different types and represents meaningful intermediate CHD states. To capture the shape-specific variations, we then learn invertible deformations to morph the learned CHD type-specific anatomies and reconstruct patient-specific shapes. Our approach has the potential to augment the image-segmentation pairs for rarer CHD types for cardiac segmentation and generate cohorts of CHD cardiac meshes for computational simulation.
Geometric Uncertainty in Patient-Specific Cardiovascular Modeling with Convolutional Dropout Networks
Sep 16, 2020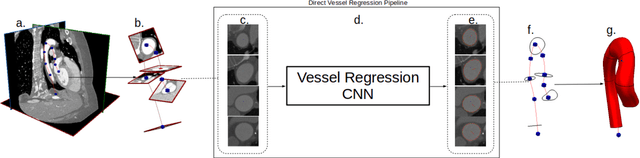


Abstract:We propose a novel approach to generate samples from the conditional distribution of patient-specific cardiovascular models given a clinically aquired image volume. A convolutional neural network architecture with dropout layers is first trained for vessel lumen segmentation using a regression approach, to enable Bayesian estimation of vessel lumen surfaces. This network is then integrated into a path-planning patient-specific modeling pipeline to generate families of cardiovascular models. We demonstrate our approach by quantifying the effect of geometric uncertainty on the hemodynamics for three patient-specific anatomies, an aorto-iliac bifurcation, an abdominal aortic aneurysm and a sub-model of the left coronary arteries. A key innovation introduced in the proposed approach is the ability to learn geometric uncertainty directly from training data. The results show how geometric uncertainty produces coefficients of variation comparable to or larger than other sources of uncertainty for wall shear stress and velocity magnitude, but has limited impact on pressure. Specifically, this is true for anatomies characterized by small vessel sizes, and for local vessel lesions seen infrequently during network training.
Dense Volume-to-Volume Vascular Boundary Detection
May 26, 2016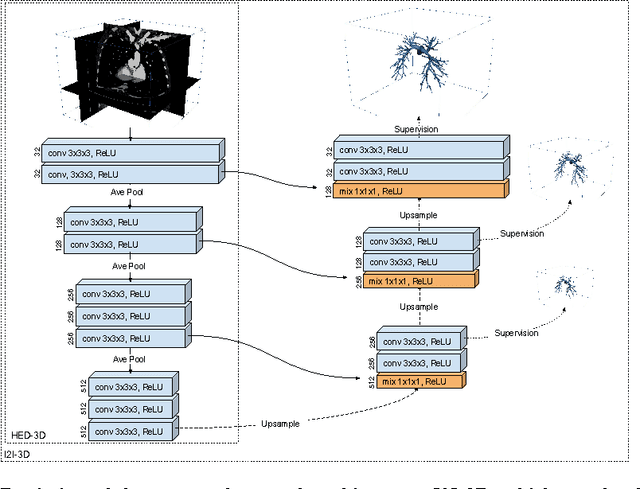
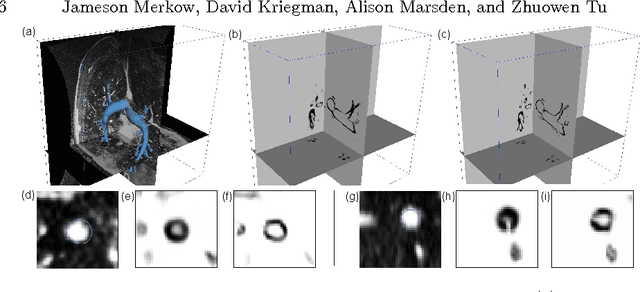

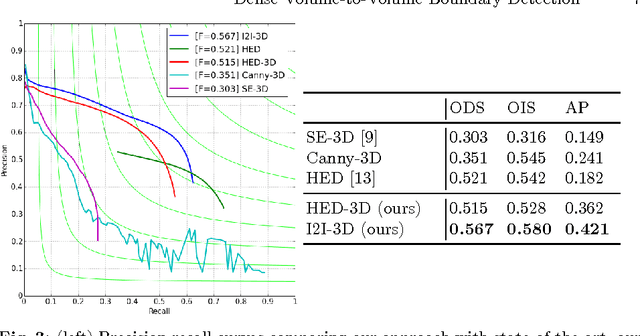
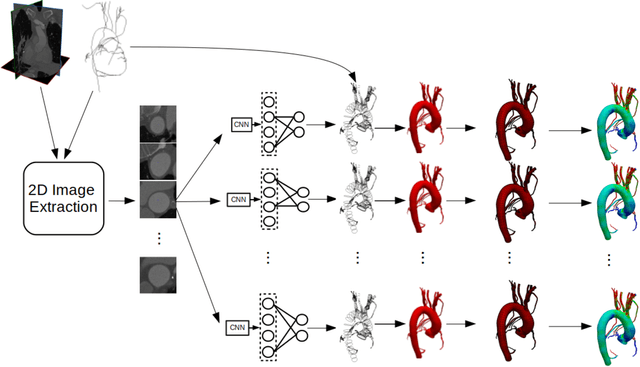
Abstract:In this work, we present a novel 3D-Convolutional Neural Network (CNN) architecture called I2I-3D that predicts boundary location in volumetric data. Our fine-to-fine, deeply supervised framework addresses three critical issues to 3D boundary detection: (1) efficient, holistic, end-to-end volumetric label training and prediction (2) precise voxel-level prediction to capture fine scale structures prevalent in medical data and (3) directed multi-scale, multi-level feature learning. We evaluate our approach on a dataset consisting of 93 medical image volumes with a wide variety of anatomical regions and vascular structures. In the process, we also introduce HED-3D, a 3D extension of the state-of-the-art 2D edge detector (HED). We show that our deep learning approach out-performs, the current state-of-the-art in 3D vascular boundary detection (structured forests 3D), by a large margin, as well as HED applied to slices, and HED-3D while successfully localizing fine structures. With our approach, boundary detection takes about one minute on a typical 512x512x512 volume.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge