Ali Abdollahzadeh
Scattering approach to diffusion quantifies axonal damage in brain injury
Jan 30, 2025Abstract:Early diagnosis and noninvasive monitoring of neurological disorders require sensitivity to elusive cellular-level alterations that occur much earlier than volumetric changes observable with the millimeter-resolution of medical imaging modalities. Morphological changes in axons, such as axonal varicosities or beadings, are observed in neurological disorders, as well as in development and aging. Here, we reveal the sensitivity of time-dependent diffusion MRI (dMRI) to axonal morphology at the micrometer scale. Scattering theory uncovers the two parameters that determine the diffusive dynamics of water in axons: the average reciprocal cross-section and the variance of long-range cross-sectional fluctuations. This theoretical development allowed us to predict dMRI metrics sensitive to axonal alterations across tens of thousands of axons in seconds rather than months of simulations in a rat model of traumatic brain injury. Our approach bridges the gap between micrometers and millimeters in resolution, offering quantitative, objective biomarkers applicable to a broad spectrum of neurological disorders.
gACSON software for automated segmentation and morphology analyses of myelinated axons in 3D electron microscopy
Dec 13, 2021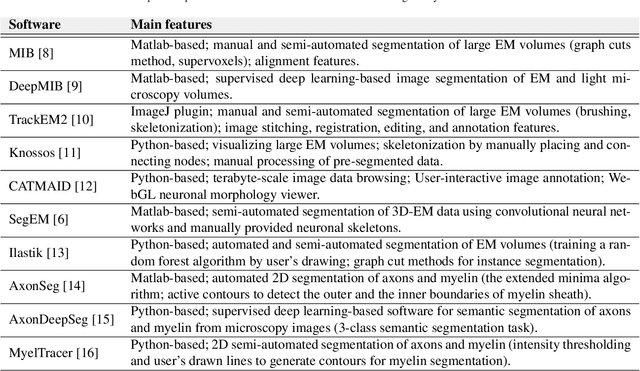
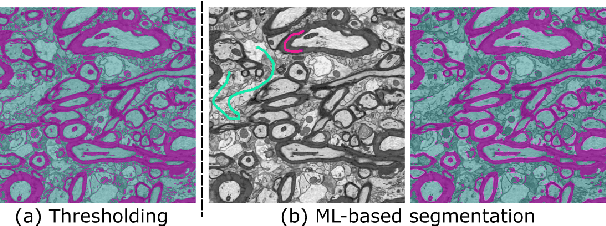
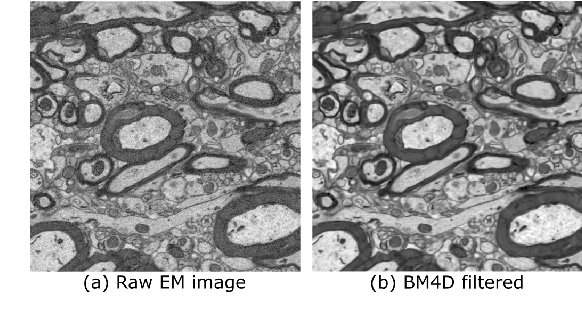


Abstract:Background and Objective: Advances in electron microscopy (EM) now allow three-dimensional (3D) imaging of hundreds of micrometers of tissue with nanometer-scale resolution, providing new opportunities to study the ultrastructure of the brain. In this work, we introduce a freely available gACSON software for visualization, segmentation, assessment, and morphology analysis of myelinated axons in 3D-EM volumes of brain tissue samples. Methods: The gACSON software is equipped with a graphical user interface (GUI). It automatically segments the intra-axonal space of myelinated axons and their corresponding myelin sheaths and allows manual segmentation, proofreading, and interactive correction of the segmented components. gACSON analyzes the morphology of myelinated axons, such as axonal diameter, axonal eccentricity, myelin thickness, or g-ratio. Results: We illustrate the use of gACSON by segmenting and analyzing myelinated axons in six 3D-EM volumes of rat somatosensory cortex after sham surgery or traumatic brain injury (TBI). Our results suggest that the equivalent diameter of myelinated axons in somatisensory cortex was decreased in TBI animals five months after the injury. Conclusions: Our results indicate that gACSON is a valuable tool for visualization, segmentation, assessment, and morphology analysis of myelinated axons in 3D-EM volumes. gACSON is freely available at https://github.com/AndreaBehan/g-ACSON under the MIT license.
Transfer Learning in Magnetic Resonance Brain Imaging: a Systematic Review
Feb 02, 2021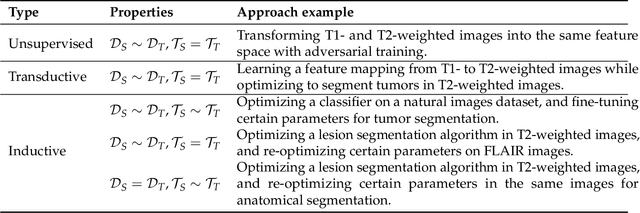
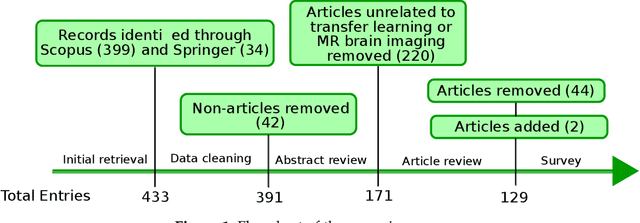

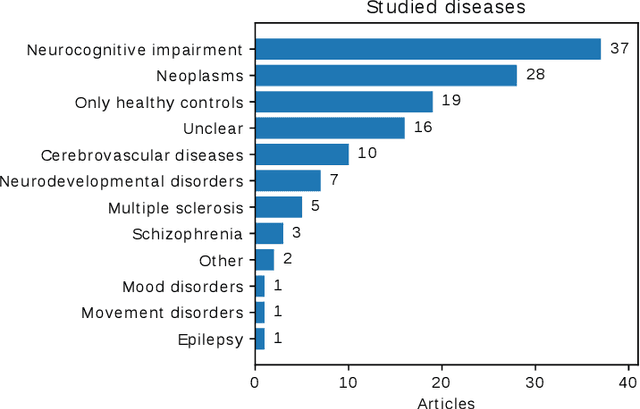
Abstract:Transfer learning refers to machine learning techniques that focus on acquiring knowledge from related tasks to improve generalization in the tasks of interest. In MRI, transfer learning is important for developing strategies that address the variation in MR images. Additionally, transfer learning is beneficial to re-utilize machine learning models that were trained to solve related tasks to the task of interest. Our goal is to identify research directions, gaps of knowledge, applications, and widely used strategies among the transfer learning approaches applied in MR brain imaging. We performed a systematic literature search for articles that applied transfer learning to MR brain imaging. We screened 433 studies and we categorized and extracted relevant information, including task type, application, and machine learning methods. Furthermore, we closely examined brain MRI-specific transfer learning approaches and other methods that tackled privacy, unseen target domains, and unlabeled data. We found 129 articles that applied transfer learning to brain MRI tasks. The most frequent applications were dementia related classification tasks and brain tumor segmentation. A majority of articles utilized transfer learning on convolutional neural networks (CNNs). Only few approaches were clearly brain MRI specific, considered privacy issues, unseen target domains or unlabeled data. We proposed a new categorization to group specific, widely-used approaches. There is an increasing interest in transfer learning within brain MRI. Public datasets have contributed to the popularity of Alzheimer's diagnostics/prognostics and tumor segmentation. Likewise, the availability of pretrained CNNs has promoted their utilization. Finally, the majority of the surveyed studies did not examine in detail the interpretation of their strategies after applying transfer learning, and did not compare to other approaches.
Cylindrical Shape Decomposition Algorithm for 3D Segmentation
Nov 01, 2019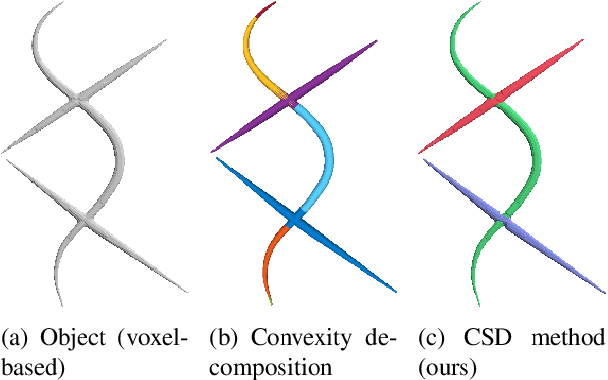
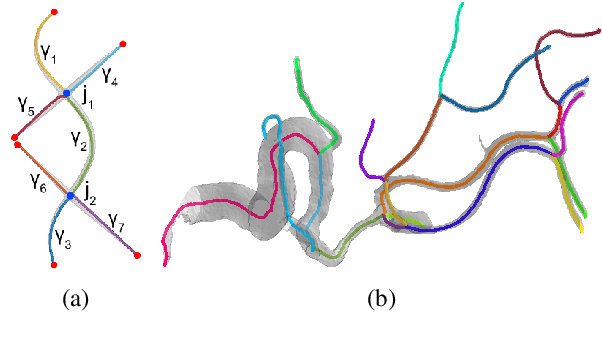
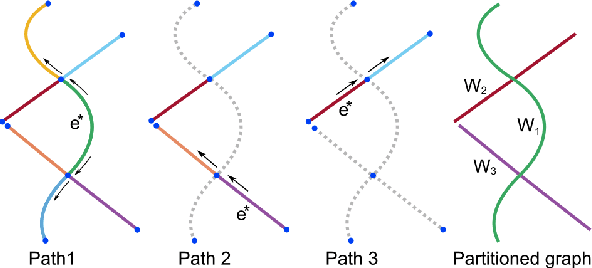

Abstract:Shape decomposition is a fundamental problem in geometry processing where an arbitrary object is regarded as an arrangement of simple primitives or semantic components. The application of 3D shape decomposition in the context of image segmentation, however, is not well-studied. In this paper, we develop a shape decomposition algorithm called cylindrical shape decomposition (CSD) to be applied for the segmentation of tubular structures in large-scale 3D images. CSD starts by partitioning the curve skeleton of a tubular object into maximal-length sub-skeletons, minimizing an orientation objective. Each sub-skeleton corresponds to a semantic component. To determine boundaries between the semantic components, CSD searches for critical points where the object cross-section substantially changes. CSD then cuts the object at critical points and assigns the same label to those object parts which are along the same sub-skeleton, defining a semantic tubular component. CSD further rectify/reconstructs these semantic components using generalized cylinders. We demonstrate the application of CSD in the segmentation of large-scale 3D electron microscopy image datasets of myelinated axons, the decomposition of vascular networks, and synthetic objects. We also compare CSD to other state-of-the-art decomposition techniques in these applications. These experiments indicate that CSD outperforms other decomposition techniques and achieves a promising performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge