Alexandra Badea
Skull stripping with purely synthetic data
May 12, 2025Abstract:While many skull stripping algorithms have been developed for multi-modal and multi-species cases, there is still a lack of a fundamentally generalizable approach. We present PUMBA(PUrely synthetic Multimodal/species invariant Brain extrAction), a strategy to train a model for brain extraction with no real brain images or labels. Our results show that even without any real images or anatomical priors, the model achieves comparable accuracy in multi-modal, multi-species and pathological cases. This work presents a new direction of research for any generalizable medical image segmentation task.
Learning sources of variability from high-dimensional observational studies
Jul 26, 2023Abstract:Causal inference studies whether the presence of a variable influences an observed outcome. As measured by quantities such as the "average treatment effect," this paradigm is employed across numerous biological fields, from vaccine and drug development to policy interventions. Unfortunately, the majority of these methods are often limited to univariate outcomes. Our work generalizes causal estimands to outcomes with any number of dimensions or any measurable space, and formulates traditional causal estimands for nominal variables as causal discrepancy tests. We propose a simple technique for adjusting universally consistent conditional independence tests and prove that these tests are universally consistent causal discrepancy tests. Numerical experiments illustrate that our method, Causal CDcorr, leads to improvements in both finite sample validity and power when compared to existing strategies. Our methods are all open source and available at github.com/ebridge2/cdcorr.
Connectome Smoothing via Low-rank Approximations
Jan 18, 2018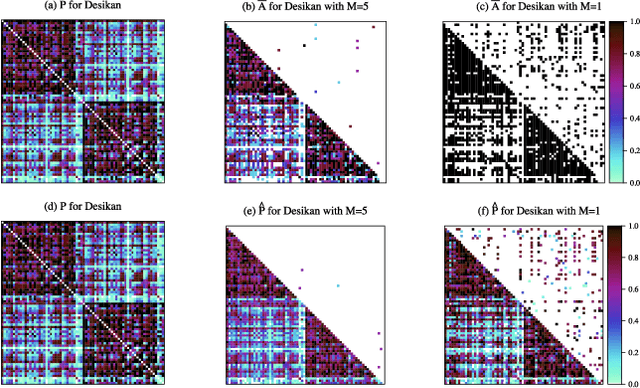
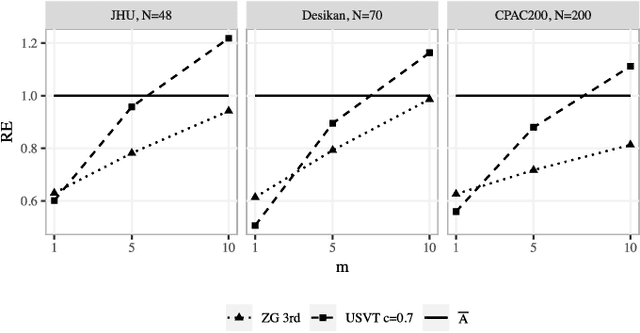
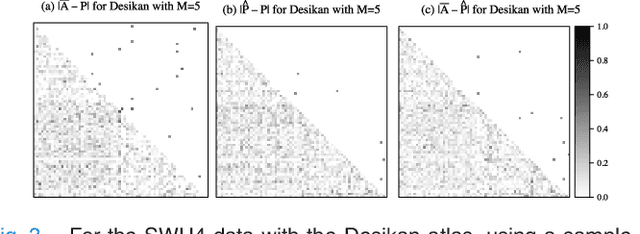
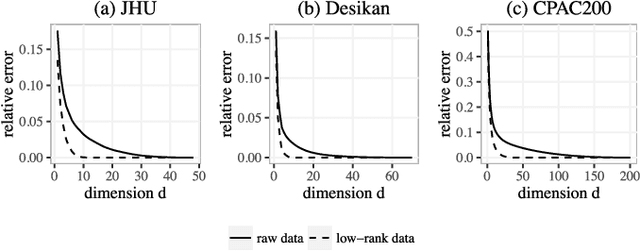
Abstract:In statistical connectomics, the quantitative study of brain networks, estimating the mean of a population of graphs based on a sample is a core problem. Often, this problem is especially difficult because the sample or cohort size is relatively small, sometimes even a single subject. While using the element-wise sample mean of the adjacency matrices is a common approach, this method does not exploit any underlying structural properties of the graphs. We propose using a low-rank method which incorporates tools for dimension selection and diagonal augmentation to smooth the estimates and improve performance over the naive methodology for small sample sizes. Theoretical results for the stochastic blockmodel show that this method offers major improvements when there are many vertices. Similarly, we demonstrate that the low-rank methods outperform the standard sample mean for a variety of independent edge distributions as well as human connectome data derived from magnetic resonance imaging, especially when sample sizes are small. Moreover, the low-rank methods yield "eigen-connectomes", which correlate with the lobe-structure of the human brain and superstructures of the mouse brain. These results indicate that low-rank methods are an important part of the tool box for researchers studying populations of graphs in general, and statistical connectomics in particular.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge