Alexander Weers
From Pixels to Histopathology: A Graph-Based Framework for Interpretable Whole Slide Image Analysis
Mar 14, 2025Abstract:The histopathological classification of whole-slide images (WSIs) is a fundamental task in digital pathology; yet it requires extensive time and expertise from specialists. While deep learning methods show promising results, they typically process WSIs by dividing them into artificial patches, which inherently prevents a network from learning from the entire image context, disregards natural tissue structures and compromises interpretability. Our method overcomes this limitation through a novel graph-based framework that constructs WSI graph representations. The WSI-graph efficiently captures essential histopathological information in a compact form. We build tissue representations (nodes) that follow biological boundaries rather than arbitrary patches all while providing interpretable features for explainability. Through adaptive graph coarsening guided by learned embeddings, we progressively merge regions while maintaining discriminative local features and enabling efficient global information exchange. In our method's final step, we solve the diagnostic task through a graph attention network. We empirically demonstrate strong performance on multiple challenging tasks such as cancer stage classification and survival prediction, while also identifying predictive factors using Integrated Gradients. Our implementation is publicly available at https://github.com/HistoGraph31/pix2pathology
Pitfalls of topology-aware image segmentation
Dec 19, 2024



Abstract:Topological correctness, i.e., the preservation of structural integrity and specific characteristics of shape, is a fundamental requirement for medical imaging tasks, such as neuron or vessel segmentation. Despite the recent surge in topology-aware methods addressing this challenge, their real-world applicability is hindered by flawed benchmarking practices. In this paper, we identify critical pitfalls in model evaluation that include inadequate connectivity choices, overlooked topological artifacts in ground truth annotations, and inappropriate use of evaluation metrics. Through detailed empirical analysis, we uncover these issues' profound impact on the evaluation and ranking of segmentation methods. Drawing from our findings, we propose a set of actionable recommendations to establish fair and robust evaluation standards for topology-aware medical image segmentation methods.
Topograph: An efficient Graph-Based Framework for Strictly Topology Preserving Image Segmentation
Nov 05, 2024



Abstract:Topological correctness plays a critical role in many image segmentation tasks, yet most networks are trained using pixel-wise loss functions, such as Dice, neglecting topological accuracy. Existing topology-aware methods often lack robust topological guarantees, are limited to specific use cases, or impose high computational costs. In this work, we propose a novel, graph-based framework for topologically accurate image segmentation that is both computationally efficient and generally applicable. Our method constructs a component graph that fully encodes the topological information of both the prediction and ground truth, allowing us to efficiently identify topologically critical regions and aggregate a loss based on local neighborhood information. Furthermore, we introduce a strict topological metric capturing the homotopy equivalence between the union and intersection of prediction-label pairs. We formally prove the topological guarantees of our approach and empirically validate its effectiveness on binary and multi-class datasets. Our loss demonstrates state-of-the-art performance with up to fivefold faster loss computation compared to persistent homology methods.
ICML Topological Deep Learning Challenge 2024: Beyond the Graph Domain
Sep 08, 2024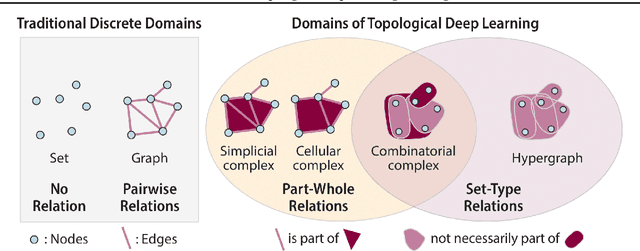
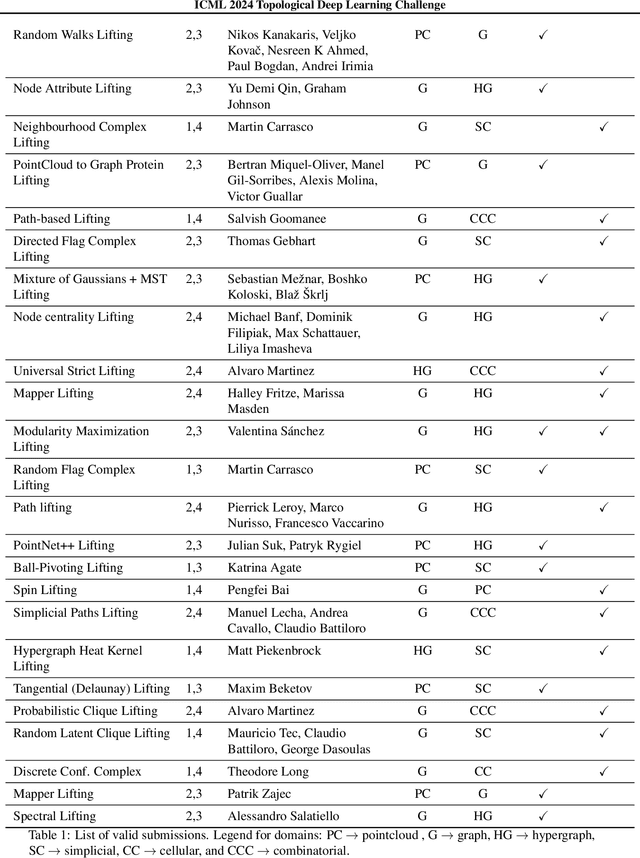
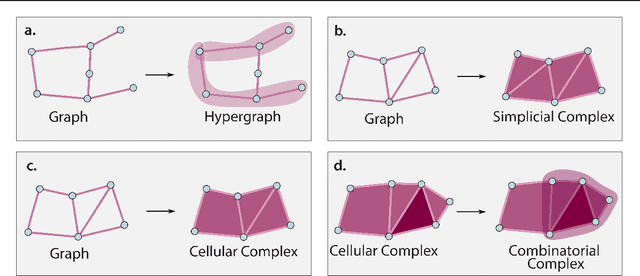
Abstract:This paper describes the 2nd edition of the ICML Topological Deep Learning Challenge that was hosted within the ICML 2024 ELLIS Workshop on Geometry-grounded Representation Learning and Generative Modeling (GRaM). The challenge focused on the problem of representing data in different discrete topological domains in order to bridge the gap between Topological Deep Learning (TDL) and other types of structured datasets (e.g. point clouds, graphs). Specifically, participants were asked to design and implement topological liftings, i.e. mappings between different data structures and topological domains --like hypergraphs, or simplicial/cell/combinatorial complexes. The challenge received 52 submissions satisfying all the requirements. This paper introduces the main scope of the challenge, and summarizes the main results and findings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge