Akira Kudo
Image Synthesis-based Late Stage Cancer Augmentation and Semi-Supervised Segmentation for MRI Rectal Cancer Staging
Dec 08, 2023Abstract:Rectal cancer is one of the most common diseases and a major cause of mortality. For deciding rectal cancer treatment plans, T-staging is important. However, evaluating the index from preoperative MRI images requires high radiologists' skill and experience. Therefore, the aim of this study is to segment the mesorectum, rectum, and rectal cancer region so that the system can predict T-stage from segmentation results. Generally, shortage of large and diverse dataset and high quality annotation are known to be the bottlenecks in computer aided diagnostics development. Regarding rectal cancer, advanced cancer images are very rare, and per-pixel annotation requires high radiologists' skill and time. Therefore, it is not feasible to collect comprehensive disease patterns in a training dataset. To tackle this, we propose two kinds of approaches of image synthesis-based late stage cancer augmentation and semi-supervised learning which is designed for T-stage prediction. In the image synthesis data augmentation approach, we generated advanced cancer images from labels. The real cancer labels were deformed to resemble advanced cancer labels by artificial cancer progress simulation. Next, we introduce a T-staging loss which enables us to train segmentation models from per-image T-stage labels. The loss works to keep inclusion/invasion relationships between rectum and cancer region consistent to the ground truth T-stage. The verification tests show that the proposed method obtains the best sensitivity (0.76) and specificity (0.80) in distinguishing between over T3 stage and underT2. In the ablation studies, our semi-supervised learning approach with the T-staging loss improved specificity by 0.13. Adding the image synthesis-based data augmentation improved the DICE score of invasion cancer area by 0.08 from baseline.
Virtual Thin Slice: 3D Conditional GAN-based Super-resolution for CT Slice Interval
Sep 02, 2019
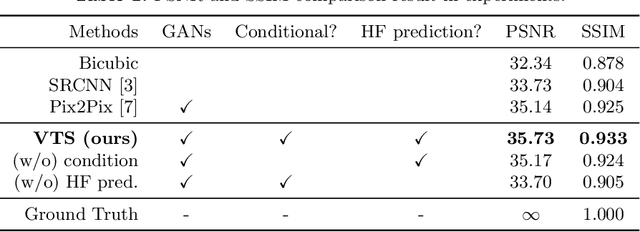

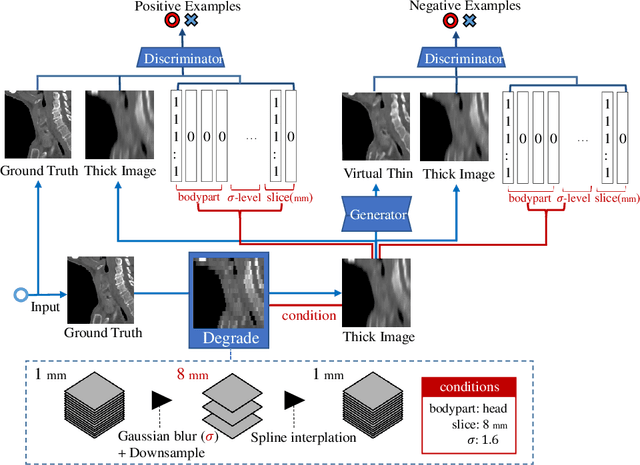
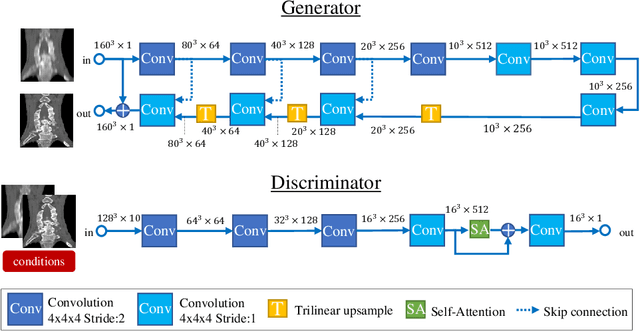
Abstract:Many CT slice images are stored with large slice intervals to reduce storage size in clinical practice. This leads to low resolution perpendicular to the slice images (i.e., z-axis), which is insufficient for 3D visualization or image analysis. In this paper, we present a novel architecture based on conditional Generative Adversarial Networks (cGANs) with the goal of generating high resolution images of main body parts including head, chest, abdomen and legs. However, GANs are known to have a difficulty with generating a diversity of patterns due to a phenomena known as mode collapse. To overcome the lack of generated pattern variety, we propose to condition the discriminator on the different body parts. Furthermore, our generator networks are extended to be three dimensional fully convolutional neural networks, allowing for the generation of high resolution images from arbitrary fields of view. In our verification tests, we show that the proposed method obtains the best scores by PSNR/SSIM metrics and Visual Turing Test, allowing for accurate reproduction of the principle anatomy in high resolution. We expect that the proposed method contribute to effective utilization of the existing vast amounts of thick CT images stored in hospitals.
Synthesizing Diverse Lung Nodules Wherever Massively: 3D Multi-Conditional GAN-based CT Image Augmentation for Object Detection
Jun 12, 2019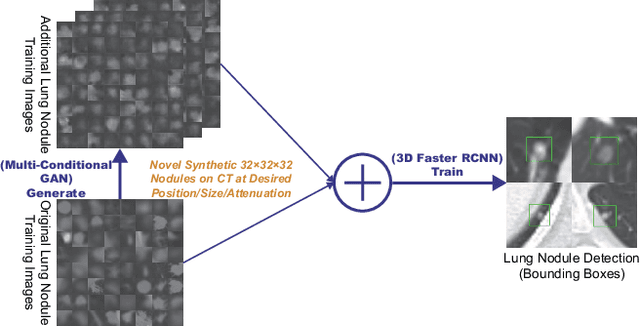
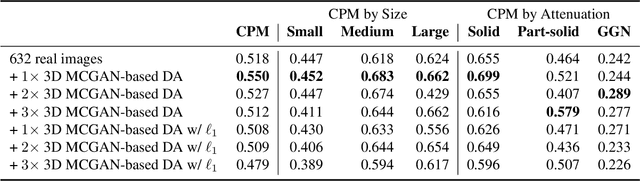
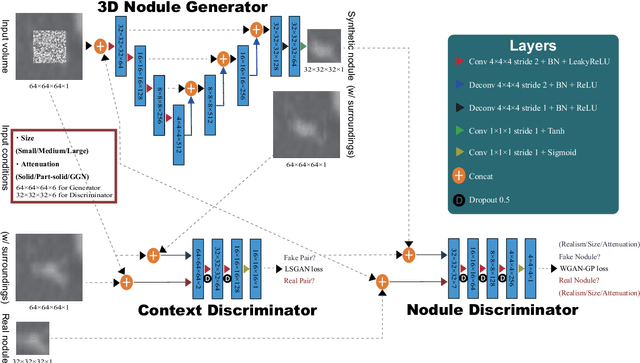
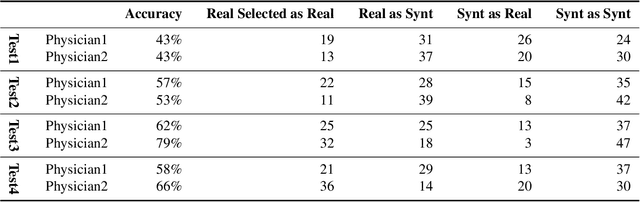
Abstract:Accurate computer-assisted diagnosis, relying on large-scale annotated pathological images, can alleviate the risk of overlooking the diagnosis. Unfortunately, in medical imaging, most available datasets are small/fragmented. To tackle this, as a Data Augmentation (DA) method, 3D conditional Generative Adversarial Networks (GANs) can synthesize desired realistic/diverse 3D images as additional training data. However, no 3D conditional GAN-based DA approach exists for general bounding box-based 3D object detection, while it can locate disease areas with physicians' minimum annotation cost, unlike rigorous 3D segmentation. Moreover, since lesions vary in position/size/attenuation, further GAN-based DA performance requires multiple conditions. Therefore, we propose 3D Multi-Conditional GAN (MCGAN) to generate realistic/diverse 32 x 32 x 32 nodules placed naturally on lung Computed Tomography images to boost sensitivity in 3D object detection. Our MCGAN adopts two discriminators for conditioning: the context discriminator learns to classify real vs synthetic nodule/surrounding pairs with noise box-centered surroundings; the nodule discriminator attempts to classify real vs synthetic nodules with size/attenuation conditions. The results show that 3D Convolutional Neural Network-based detection can achieve higher sensitivity under any nodule size/attenuation at fixed False Positive rates and overcome the medical data paucity with the MCGAN-generated realistic nodules---even expert physicians fail to distinguish them from the real ones in Visual Turing Test.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge