Ahmed Nebli
A Few-shot Learning Graph Multi-Trajectory Evolution Network for Forecasting Multimodal Baby Connectivity Development from a Baseline Timepoint
Oct 06, 2021
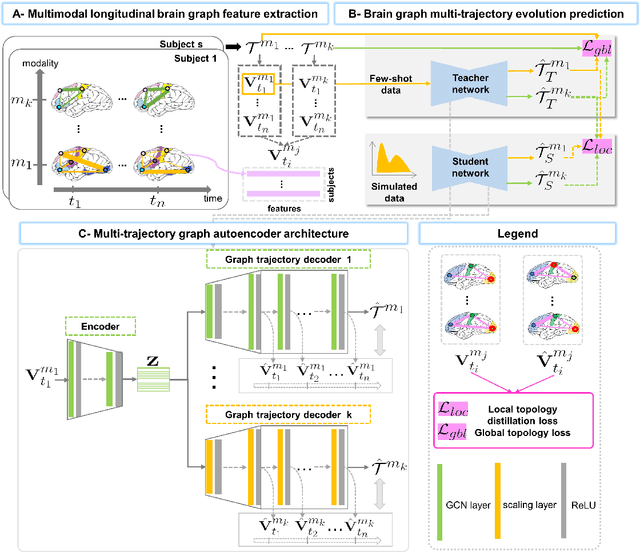
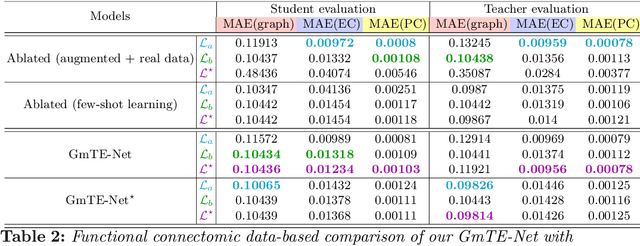
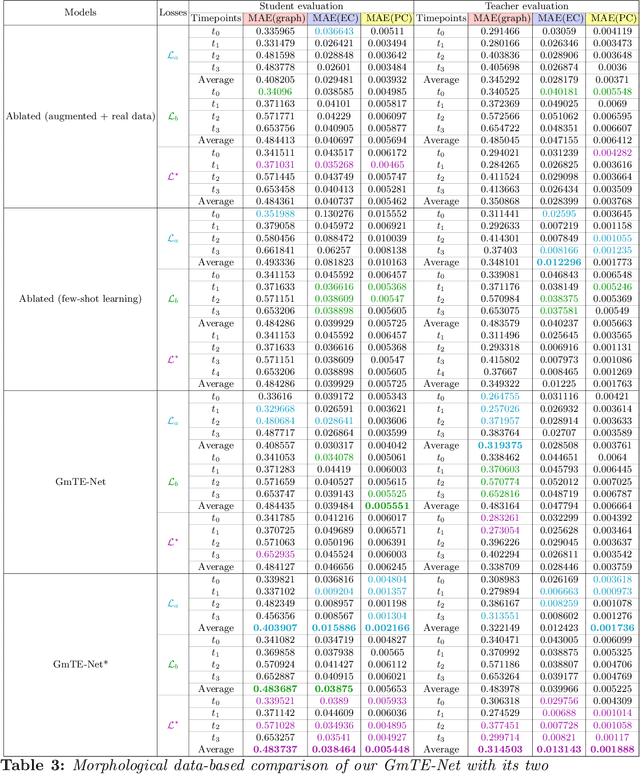
Abstract:Charting the baby connectome evolution trajectory during the first year after birth plays a vital role in understanding dynamic connectivity development of baby brains. Such analysis requires acquisition of longitudinal connectomic datasets. However, both neonatal and postnatal scans are rarely acquired due to various difficulties. A small body of works has focused on predicting baby brain evolution trajectory from a neonatal brain connectome derived from a single modality. Although promising, large training datasets are essential to boost model learning and to generalize to a multi-trajectory prediction from different modalities (i.e., functional and morphological connectomes). Here, we unprecedentedly explore the question: Can we design a few-shot learning-based framework for predicting brain graph trajectories across different modalities? To this aim, we propose a Graph Multi-Trajectory Evolution Network (GmTE-Net), which adopts a teacher-student paradigm where the teacher network learns on pure neonatal brain graphs and the student network learns on simulated brain graphs given a set of different timepoints. To the best of our knowledge, this is the first teacher-student architecture tailored for brain graph multi-trajectory growth prediction that is based on few-shot learning and generalized to graph neural networks (GNNs). To boost the performance of the student network, we introduce a local topology-aware distillation loss that forces the predicted graph topology of the student network to be consistent with the teacher network. Experimental results demonstrate substantial performance gains over benchmark methods. Hence, our GmTE-Net can be leveraged to predict atypical brain connectivity trajectory evolution across various modalities. Our code is available at https: //github.com/basiralab/GmTE-Net.
Non-isomorphic Inter-modality Graph Alignment and Synthesis for Holistic Brain Mapping
Jun 30, 2021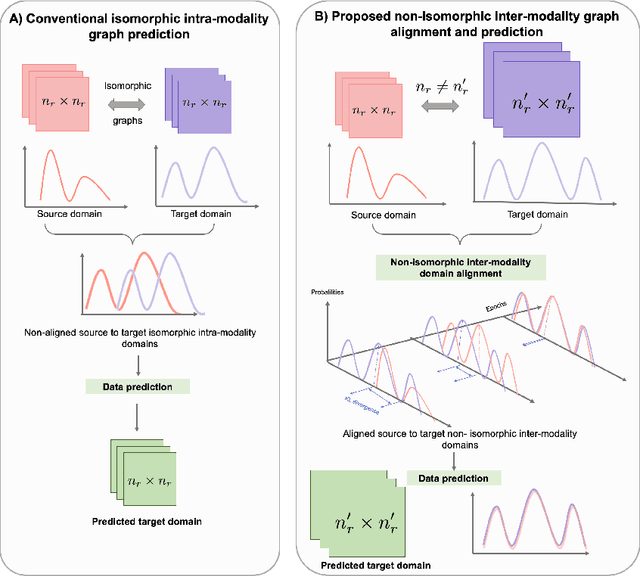
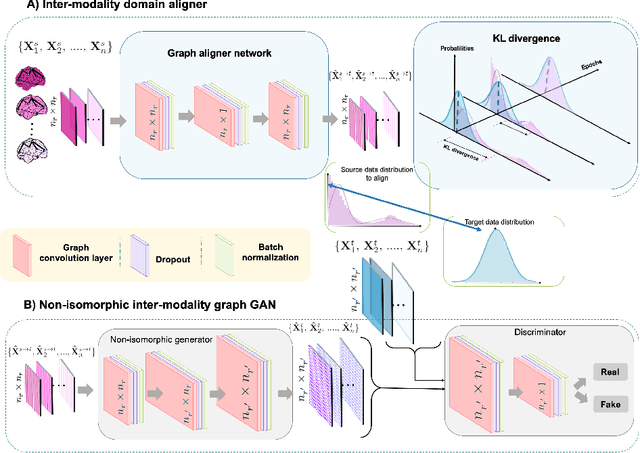
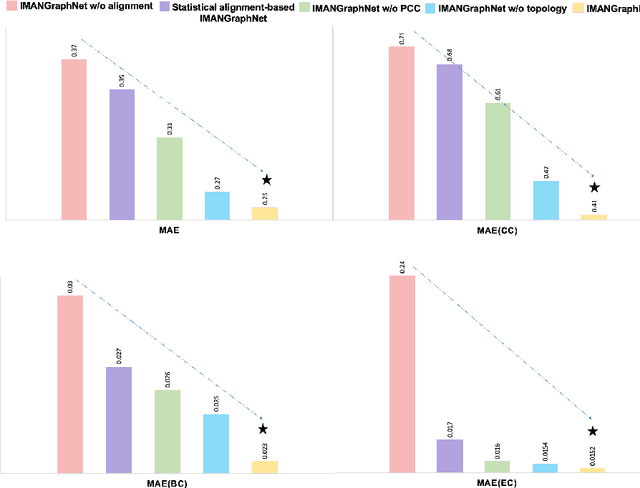
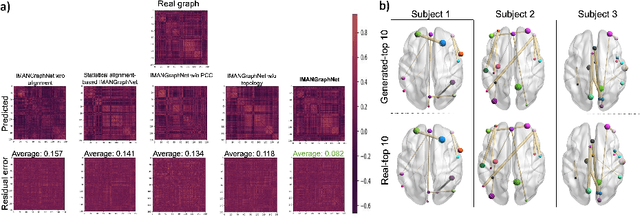
Abstract:Brain graph synthesis marked a new era for predicting a target brain graph from a source one without incurring the high acquisition cost and processing time of neuroimaging data. However, existing multi-modal graph synthesis frameworks have several limitations. First, they mainly focus on generating graphs from the same domain (intra-modality), overlooking the rich multimodal representations of brain connectivity (inter-modality). Second, they can only handle isomorphic graph generation tasks, limiting their generalizability to synthesizing target graphs with a different node size and topological structure from those of the source one. More importantly, both target and source domains might have different distributions, which causes a domain fracture between them (i.e., distribution misalignment). To address such challenges, we propose an inter-modality aligner of non-isomorphic graphs (IMANGraphNet) framework to infer a target graph modality based on a given modality. Our three core contributions lie in (i) predicting a target graph (e.g., functional) from a source graph (e.g., morphological) based on a novel graph generative adversarial network (gGAN); (ii) using non-isomorphic graphs for both source and target domains with a different number of nodes, edges and structure; and (iii) enforcing the predicted target distribution to match that of the ground truth graphs using a graph autoencoder to relax the designed loss oprimization. To handle the unstable behavior of gGAN, we design a new Ground Truth-Preserving (GT-P) loss function to guide the generator in learning the topological structure of ground truth brain graphs. Our comprehensive experiments on predicting functional from morphological graphs demonstrate the outperformance of IMANGraphNet in comparison with its variants. This can be further leveraged for integrative and holistic brain mapping in health and disease.
Deep EvoGraphNet Architecture For Time-Dependent Brain Graph Data Synthesis From a Single Timepoint
Sep 28, 2020
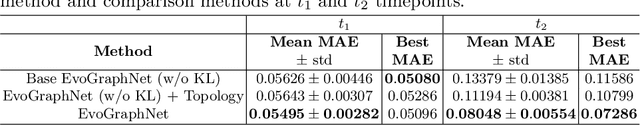
Abstract:Learning how to predict the brain connectome (i.e. graph) development and aging is of paramount importance for charting the future of within-disorder and cross-disorder landscape of brain dysconnectivity evolution. Indeed, predicting the longitudinal (i.e., time-dependent ) brain dysconnectivity as it emerges and evolves over time from a single timepoint can help design personalized treatments for disordered patients in a very early stage. Despite its significance, evolution models of the brain graph are largely overlooked in the literature. Here, we propose EvoGraphNet, the first end-to-end geometric deep learning-powered graph-generative adversarial network (gGAN) for predicting time-dependent brain graph evolution from a single timepoint. Our EvoGraphNet architecture cascades a set of time-dependent gGANs, where each gGAN communicates its predicted brain graphs at a particular timepoint to train the next gGAN in the cascade at follow-up timepoint. Therefore, we obtain each next predicted timepoint by setting the output of each generator as the input of its successor which enables us to predict a given number of timepoints using only one single timepoint in an end- to-end fashion. At each timepoint, to better align the distribution of the predicted brain graphs with that of the ground-truth graphs, we further integrate an auxiliary Kullback-Leibler divergence loss function. To capture time-dependency between two consecutive observations, we impose an l1 loss to minimize the sparse distance between two serialized brain graphs. A series of benchmarks against variants and ablated versions of our EvoGraphNet showed that we can achieve the lowest brain graph evolution prediction error using a single baseline timepoint. Our EvoGraphNet code is available at http://github.com/basiralab/EvoGraphNet.
Adversarial Brain Multiplex Prediction From a Single Network for High-Order Connectional Gender-Specific Brain Mapping
Sep 24, 2020
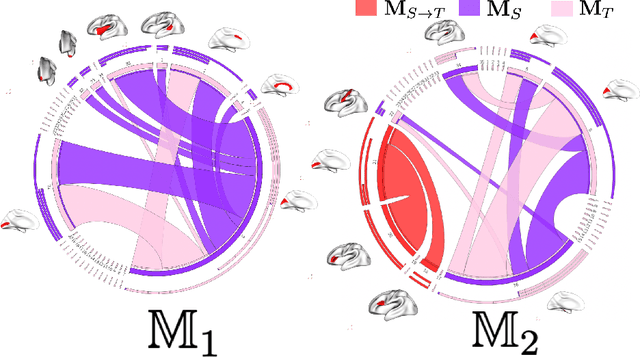
Abstract:Brain connectivity networks, derived from magnetic resonance imaging (MRI), non-invasively quantify the relationship in function, structure, and morphology between two brain regions of interest (ROIs) and give insights into gender-related connectional differences. However, to the best of our knowledge, studies on gender differences in brain connectivity were limited to investigating pairwise (i.e., low-order) relationship ROIs, overlooking the complex high-order interconnectedness of the brain as a network. To address this limitation, brain multiplexes have been introduced to model the relationship between at least two different brain networks. However, this inhibits their application to datasets with single brain networks such as functional networks. To fill this gap, we propose the first work on predicting brain multiplexes from a source network to investigate gender differences. Recently, generative adversarial networks (GANs) submerged the field of medical data synthesis. However, although conventional GANs work well on images, they cannot handle brain networks due to their non-Euclidean topological structure. Differently, in this paper, we tap into the nascent field of geometric-GANs (G-GAN) to design a deep multiplex prediction architecture comprising (i) a geometric source to target network translator mimicking a U-Net architecture with skip connections and (ii) a conditional discriminator which classifies predicted target intra-layers by conditioning on the multiplex source intra-layers. Such architecture simultaneously learns the latent source network representation and the deep non-linear mapping from the source to target multiplex intra-layers. Our experiments on a large dataset demonstrated that predicted multiplexes significantly boost gender classification accuracy compared with source networks and identifies both low and high-order gender-specific multiplex connections.
Foreseeing Brain Graph Evolution Over Time Using Deep Adversarial Network Normalizer
Sep 23, 2020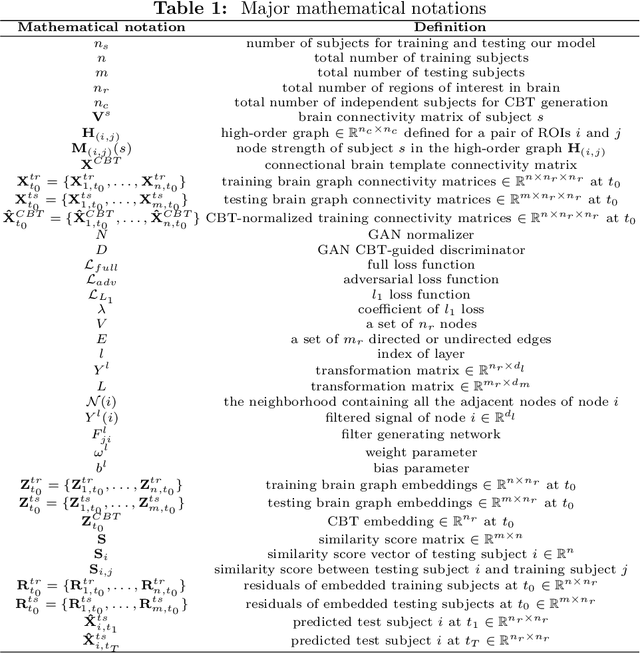

Abstract:Foreseeing the brain evolution as a complex highly inter-connected system, widely modeled as a graph, is crucial for mapping dynamic interactions between different anatomical regions of interest (ROIs) in health and disease. Interestingly, brain graph evolution models remain almost absent in the literature. Here we design an adversarial brain network normalizer for representing each brain network as a transformation of a fixed centered population-driven connectional template. Such graph normalization with respect to a fixed reference paves the way for reliably identifying the most similar training samples (i.e., brain graphs) to the testing sample at baseline timepoint. The testing evolution trajectory will be then spanned by the selected training graphs and their corresponding evolution trajectories. We base our prediction framework on geometric deep learning which naturally operates on graphs and nicely preserves their topological properties. Specifically, we propose the first graph-based Generative Adversarial Network (gGAN) that not only learns how to normalize brain graphs with respect to a fixed connectional brain template (CBT) (i.e., a brain template that selectively captures the most common features across a brain population) but also learns a high-order representation of the brain graphs also called embeddings. We use these embeddings to compute the similarity between training and testing subjects which allows us to pick the closest training subjects at baseline timepoint to predict the evolution of the testing brain graph over time. A series of benchmarks against several comparison methods showed that our proposed method achieved the lowest brain disease evolution prediction error using a single baseline timepoint. Our gGAN code is available at http://github.com/basiralab/gGAN.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge