Adrian Celaya
Learning Discontinuous Galerkin Solutions to Elliptic Problems via Small Linear Convolutional Neural Networks
Feb 12, 2025



Abstract:In recent years, there has been an increasing interest in using deep learning and neural networks to tackle scientific problems, particularly in solving partial differential equations (PDEs). However, many neural network-based methods, such as physics-informed neural networks, depend on automatic differentiation and the sampling of collocation points, which can result in a lack of interpretability and lower accuracy compared to traditional numerical methods. To address this issue, we propose two approaches for learning discontinuous Galerkin solutions to PDEs using small linear convolutional neural networks. Our first approach is supervised and depends on labeled data, while our second approach is unsupervised and does not rely on any training data. In both cases, our methods use substantially fewer parameters than similar numerics-based neural networks while also demonstrating comparable accuracy to the true and DG solutions for elliptic problems.
An Adaptive Collocation Point Strategy For Physics Informed Neural Networks via the QR Discrete Empirical Interpolation Method
Jan 16, 2025



Abstract:Physics-informed neural networks (PINNs) have gained significant attention for solving forward and inverse problems related to partial differential equations (PDEs). While advancements in loss functions and network architectures have improved PINN accuracy, the impact of collocation point sampling on their performance remains underexplored. Fixed sampling methods, such as uniform random sampling and equispaced grids, can fail to capture critical regions with high solution gradients, limiting their effectiveness for complex PDEs. Adaptive methods, inspired by adaptive mesh refinement from traditional numerical methods, address this by dynamically updating collocation points during training but may overlook residual dynamics between updates, potentially losing valuable information. To overcome this limitation, we propose an adaptive collocation point selection strategy utilizing the QR Discrete Empirical Interpolation Method (QR-DEIM), a reduced-order modeling technique for efficiently approximating nonlinear functions. Our results on benchmark PDEs, including the wave, Allen-Cahn, and Burgers' equations, demonstrate that our QR-DEIM-based approach improves PINN accuracy compared to existing methods, offering a promising direction for adaptive collocation point strategies.
MIST: A Simple and Scalable End-To-End 3D Medical Imaging Segmentation Framework
Jul 31, 2024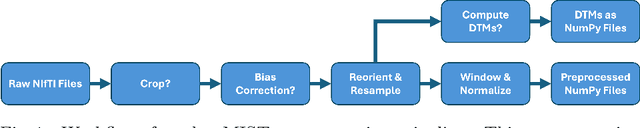
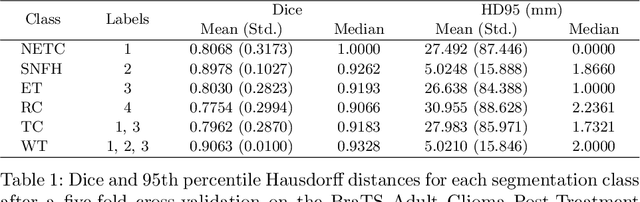
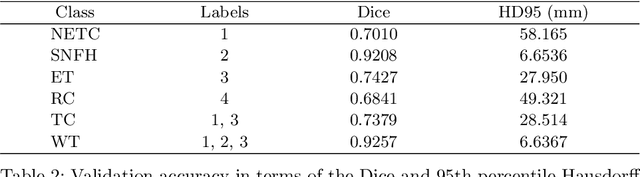
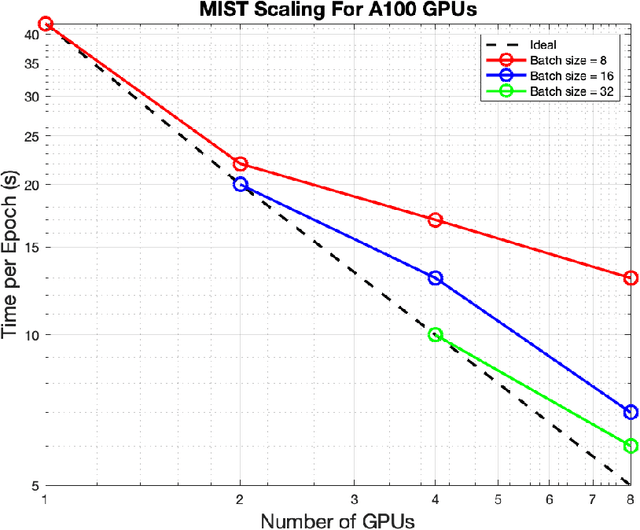
Abstract:Medical imaging segmentation is a highly active area of research, with deep learning-based methods achieving state-of-the-art results in several benchmarks. However, the lack of standardized tools for training, testing, and evaluating new methods makes the comparison of methods difficult. To address this, we introduce the Medical Imaging Segmentation Toolkit (MIST), a simple, modular, and end-to-end medical imaging segmentation framework designed to facilitate consistent training, testing, and evaluation of deep learning-based medical imaging segmentation methods. MIST standardizes data analysis, preprocessing, and evaluation pipelines, accommodating multiple architectures and loss functions. This standardization ensures reproducible and fair comparisons across different methods. We detail MIST's data format requirements, pipelines, and auxiliary features and demonstrate its efficacy using the BraTS Adult Glioma Post-Treatment Challenge dataset. Our results highlight MIST's ability to produce accurate segmentation masks and its scalability across multiple GPUs, showcasing its potential as a powerful tool for future medical imaging research and development.
Solutions to Elliptic and Parabolic Problems via Finite Difference Based Unsupervised Small Linear Convolutional Neural Networks
Nov 01, 2023



Abstract:In recent years, there has been a growing interest in leveraging deep learning and neural networks to address scientific problems, particularly in solving partial differential equations (PDEs). However, current neural network-based PDE solvers often rely on extensive training data or labeled input-output pairs, making them prone to challenges in generalizing to out-of-distribution examples. To mitigate the generalization gap encountered by conventional neural network-based methods in estimating PDE solutions, we formulate a fully unsupervised approach, requiring no training data, to estimate finite difference solutions for PDEs directly via small convolutional neural networks. Our proposed algorithms demonstrate a comparable accuracy to the true solution for several selected elliptic and parabolic problems compared to the finite difference method.
FMG-Net and W-Net: Multigrid Inspired Deep Learning Architectures For Medical Imaging Segmentation
Apr 05, 2023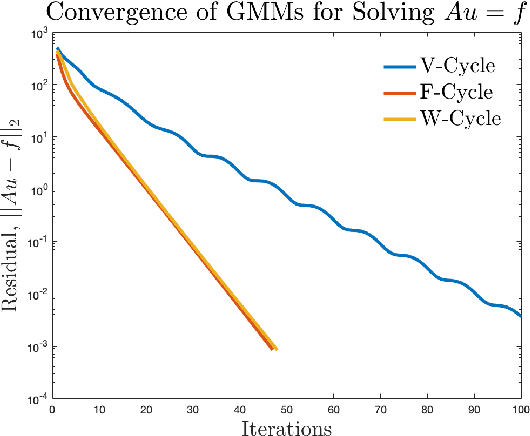
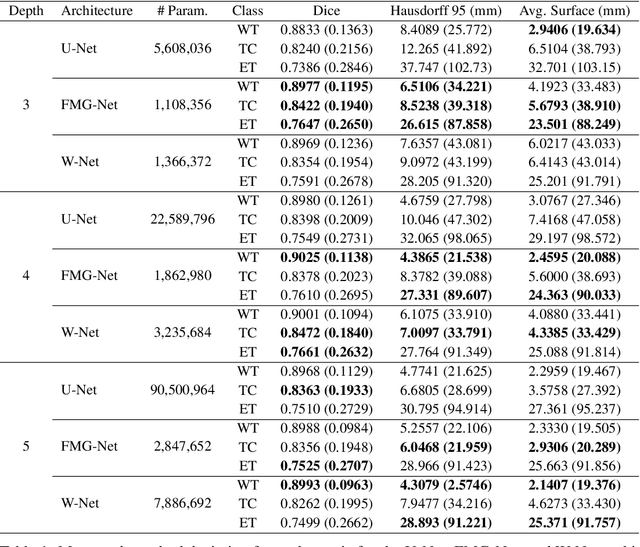
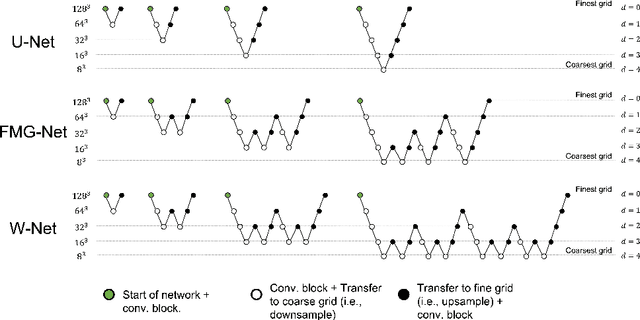
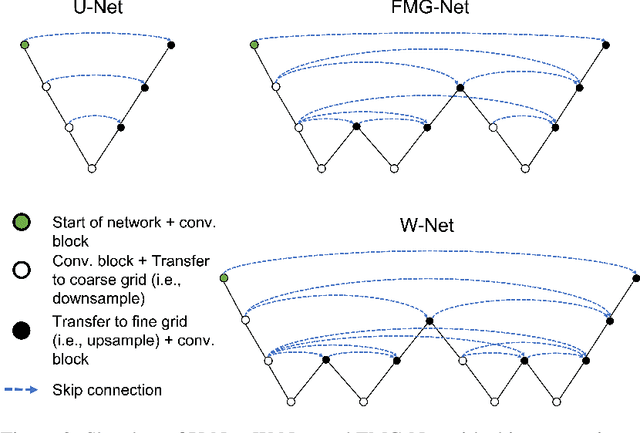
Abstract:Accurate medical imaging segmentation is critical for precise and effective medical interventions. However, despite the success of convolutional neural networks (CNNs) in medical image segmentation, they still face challenges in handling fine-scale features and variations in image scales. These challenges are particularly evident in complex and challenging segmentation tasks, such as the BraTS multi-label brain tumor segmentation challenge. In this task, accurately segmenting the various tumor sub-components, which vary significantly in size and shape, remains a significant challenge, with even state-of-the-art methods producing substantial errors. Therefore, we propose two architectures, FMG-Net and W-Net, that incorporate the principles of geometric multigrid methods for solving linear systems of equations into CNNs to address these challenges. Our experiments on the BraTS 2020 dataset demonstrate that both FMG-Net and W-Net outperform the widely used U-Net architecture regarding tumor subcomponent segmentation accuracy and training efficiency. These findings highlight the potential of incorporating the principles of multigrid methods into CNNs to improve the accuracy and efficiency of medical imaging segmentation.
A Weighted Normalized Boundary Loss for Reducing the Hausdorff Distance in Medical Imaging Segmentation
Feb 08, 2023Abstract:Within medical imaging segmentation, the Dice coefficient and Hausdorff-based metrics are standard measures of success for deep learning models. However, modern loss functions for medical image segmentation often only consider the Dice coefficient or similar region-based metrics during training. As a result, segmentation architectures trained over such loss functions run the risk of achieving high accuracy for the Dice coefficient but low accuracy for Hausdorff-based metrics. Low accuracy on Hausdorff-based metrics can be problematic for applications such as tumor segmentation, where such benchmarks are crucial. For example, high Dice scores accompanied by significant Hausdorff errors could indicate that the predictions fail to detect small tumors. We propose the Weighted Normalized Boundary Loss, a novel loss function to minimize Hausdorff-based metrics with more desirable numerical properties than current methods and with weighting terms for class imbalance. Our loss function outperforms other losses when tested on the BraTS dataset using a standard 3D U-Net and the state-of-the-art nnUNet architectures. These results suggest we can improve segmentation accuracy with our novel loss function.
Inversion of Time-Lapse Surface Gravity Data for Detection of 3D CO$_2$ Plumes via Deep Learning
Sep 06, 2022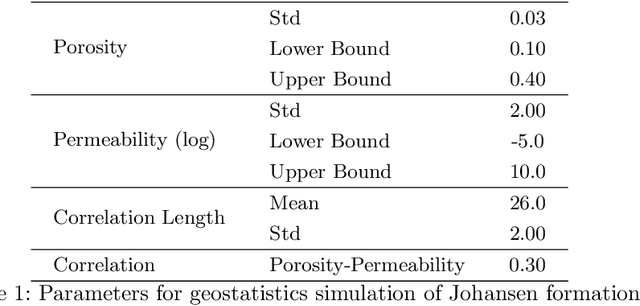
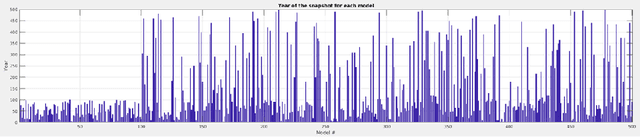
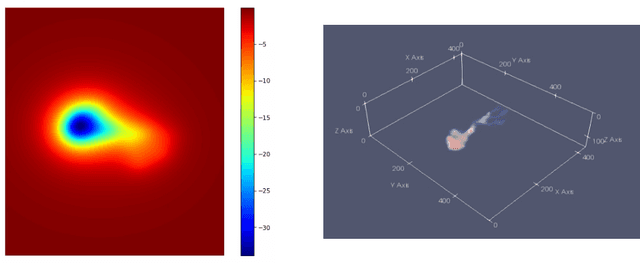
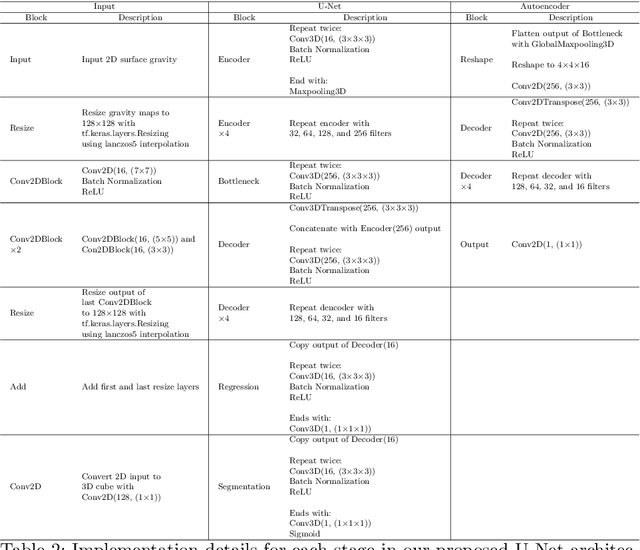
Abstract:We introduce three algorithms that invert simulated gravity data to 3D subsurface rock/flow properties. The first algorithm is a data-driven, deep learning-based approach, the second mixes a deep learning approach with physical modeling into a single workflow, and the third considers the time dependence of surface gravity monitoring. The target application of these proposed algorithms is the prediction of subsurface CO$_2$ plumes as a complementary tool for monitoring CO$_2$ sequestration deployments. Each proposed algorithm outperforms traditional inversion methods and produces high-resolution, 3D subsurface reconstructions in near real-time. Our proposed methods achieve Dice scores of up to 0.8 for predicted plume geometry and near perfect data misfit in terms of $\mu$Gals. These results indicate that combining 4D surface gravity monitoring with deep learning techniques represents a low-cost, rapid, and non-intrusive method for monitoring CO$_2$ storage sites.
Correlation between image quality metrics of magnetic resonance images and the neural network segmentation accuracy
Nov 01, 2021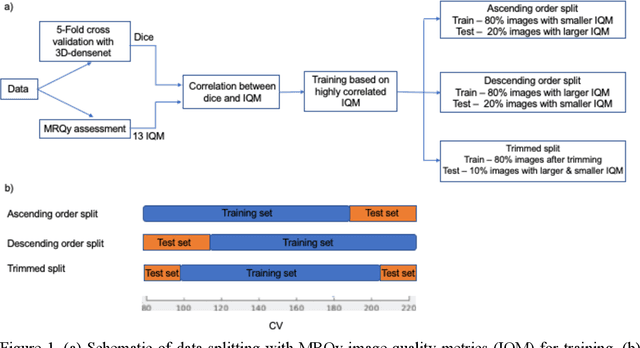
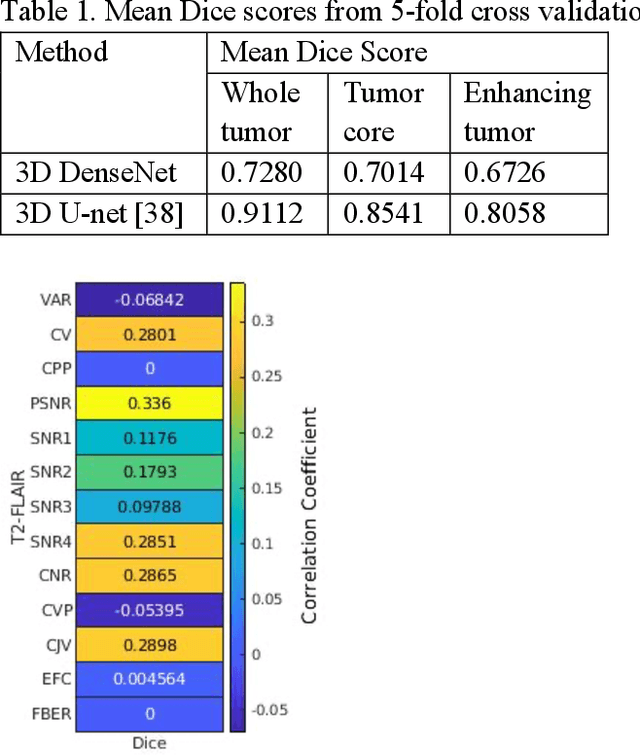
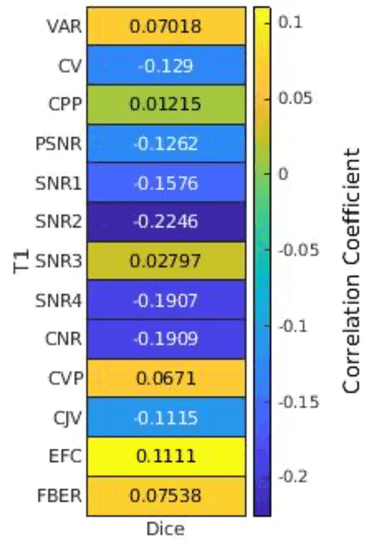
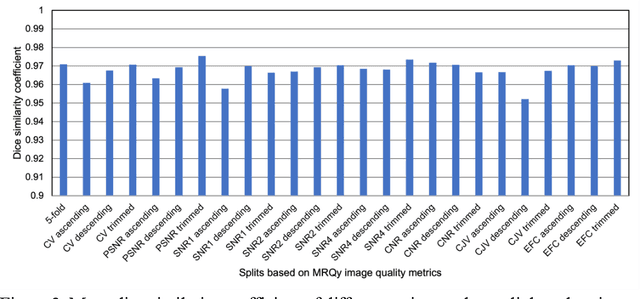
Abstract:Deep neural networks with multilevel connections process input data in complex ways to learn the information.A networks learning efficiency depends not only on the complex neural network architecture but also on the input training images.Medical image segmentation with deep neural networks for skull stripping or tumor segmentation from magnetic resonance images enables learning both global and local features of the images.Though medical images are collected in a controlled environment,there may be artifacts or equipment based variance that cause inherent bias in the input set.In this study, we investigated the correlation between the image quality metrics of MR images with the neural network segmentation accuracy.For that we have used the 3D DenseNet architecture and let the network trained on the same input but applying different methodologies to select the training data set based on the IQM values.The difference in the segmentation accuracy between models based on the random training inputs with IQM based training inputs shed light on the role of image quality metrics on segmentation accuracy.By running the image quality metrics to choose the training inputs,further we may tune the learning efficiency of the network and the segmentation accuracy.
PocketNet: A Smaller Neural Network for 3D Medical Image Segmentation
Apr 21, 2021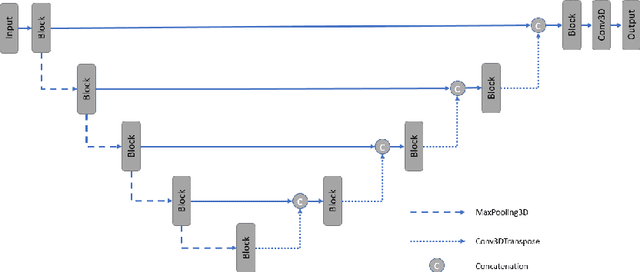
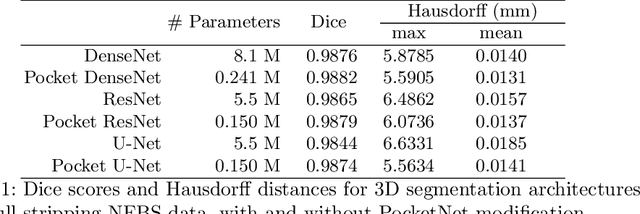
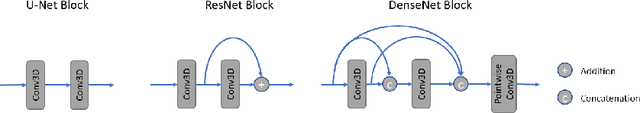
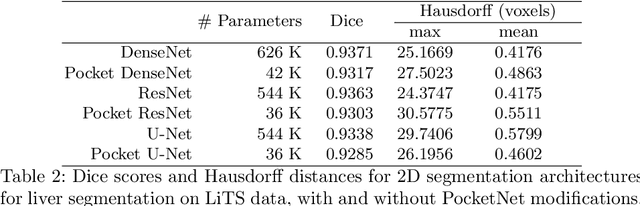
Abstract:Overparameterized deep learning networks have shown impressive performance in the area of automatic medical image segmentation. However, they achieve this performance at an enormous cost in memory, runtime, and energy. A large source of overparameterization in modern neural networks results from doubling the number of feature maps with each downsampling layer. This rapid growth in the number of parameters results in network architectures that require a significant amount of computing resources, making them less accessible and difficult to use. By keeping the number of feature maps constant throughout the network, we derive a new CNN architecture called PocketNet that achieves comparable segmentation results to conventional CNNs while using less than 3% of the number of parameters.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge