Weakly Supervised Multi-Organ Multi-Disease Classification of Body CT Scans
Paper and Code
Aug 03, 2020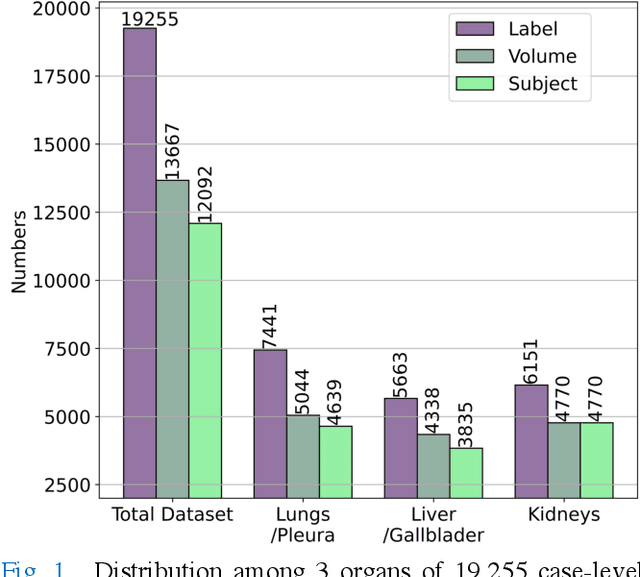
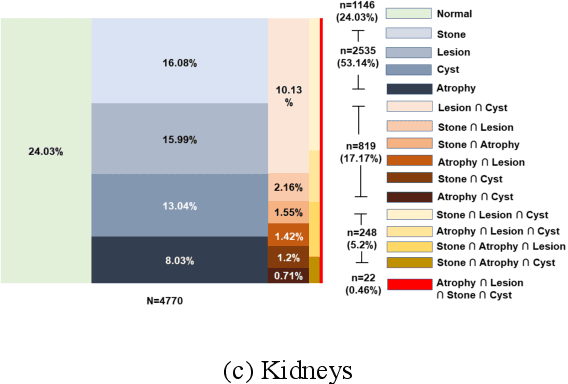
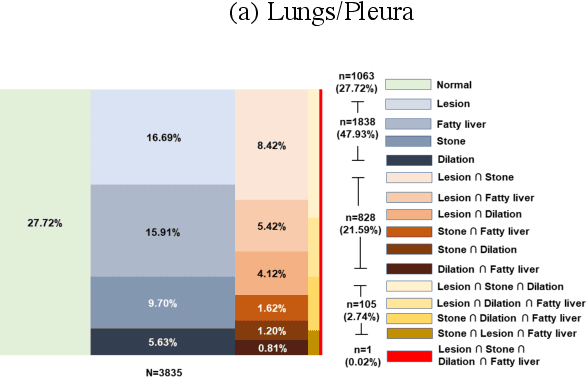
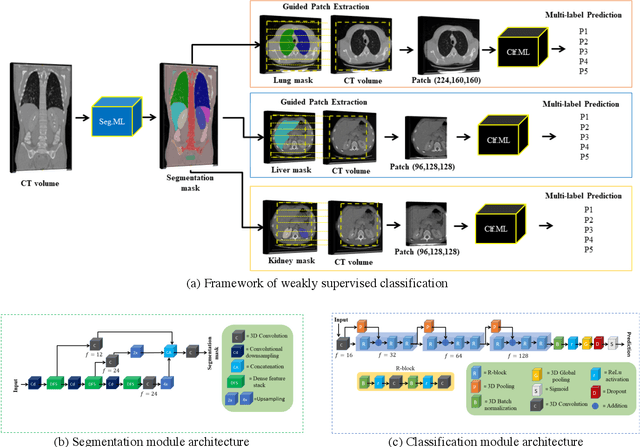
We designed a multi-organ, multi-label disease classification algorithm for computed tomography (CT) scans using case-level labels from radiology text reports. A rule-based algorithm extracted 19,255 disease labels from reports of 13,667 body CT scans from 12,092 subjects. A 3D DenseVNet was trained to segment 3 organ systems: lungs/pleura, liver/gallbladder, and kidneys. From patches guided by segmentations, a 3D convolutional neural network provided multi-label disease classification for normality versus four common diseases per organ. The process was tested on 2,158 CT volumes with 2,875 manually obtained labels. Manual validation of the rulebased labels confirmed 91 to 99% accuracy. Results were characterized using the receiver operating characteristic area under the curve (AUC). Classification AUCs for lungs/pleura labels were as follows: atelectasis 0.77 (95% confidence intervals 0.74 to 0.81), nodule 0.65 (0.61 to 0.69), emphysema 0.89 (0.86 to 0.92), effusion 0.97 (0.96 to 0.98), and normal 0.89 (0.87 to 0.91). For liver/gallbladder, AUCs were: stone 0.62 (0.56 to 0.67), lesion 0.73 (0.69 to 0.77), dilation 0.87 (0.84 to 0.90), fatty 0.89 (0.86 to 0.92), and normal 0.82 (0.78 to 0.85). For kidneys, AUCs were: stone 0.83 (0.79 to 0.87), atrophy 0.92 (0.89 to 0.94), lesion 0.68 (0.64 to 0.72), cyst 0.70 (0.66 to 0.73), and normal 0.79 (0.75 to 0.83). In conclusion, by using automated extraction of disease labels from radiology reports, we created a weakly supervised, multi-organ, multi-disease classifier that can be easily adapted to efficiently leverage massive amounts of unannotated data associated with medical images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge