Towards Cross-Domain Single Blood Cell Image Classification via Large-Scale LoRA-based Segment Anything Model
Paper and Code
Aug 13, 2024
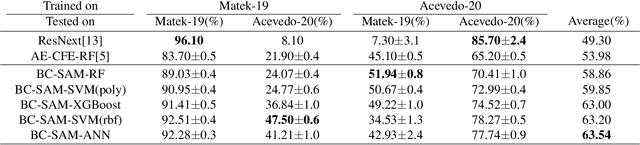
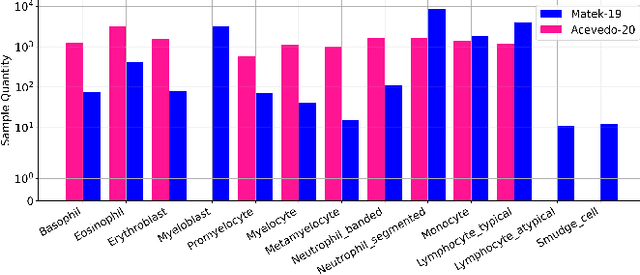
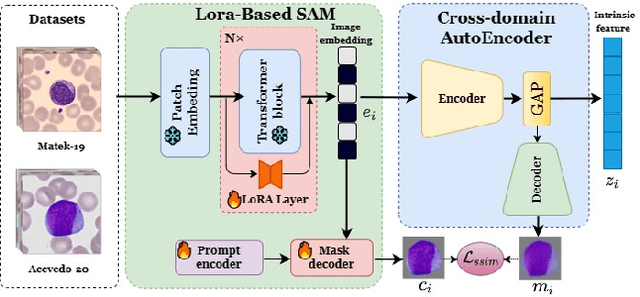
Accurate classification of blood cells plays a vital role in hematological analysis as it aids physicians in diagnosing various medical conditions. In this study, we present a novel approach for classifying blood cell images known as BC-SAM. BC-SAM leverages the large-scale foundation model of Segment Anything Model (SAM) and incorporates a fine-tuning technique using LoRA, allowing it to extract general image embeddings from blood cell images. To enhance the applicability of BC-SAM across different blood cell image datasets, we introduce an unsupervised cross-domain autoencoder that focuses on learning intrinsic features while suppressing artifacts in the images. To assess the performance of BC-SAM, we employ four widely used machine learning classifiers (Random Forest, Support Vector Machine, Artificial Neural Network, and XGBoost) to construct blood cell classification models and compare them against existing state-of-the-art methods. Experimental results conducted on two publicly available blood cell datasets (Matek-19 and Acevedo-20) demonstrate that our proposed BC-SAM achieves a new state-of-the-art result, surpassing the baseline methods with a significant improvement. The source code of this paper is available at https://github.com/AnoK3111/BC-SAM.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge