SMT-DTA: Improving Drug-Target Affinity Prediction with Semi-supervised Multi-task Training
Paper and Code
Jun 22, 2022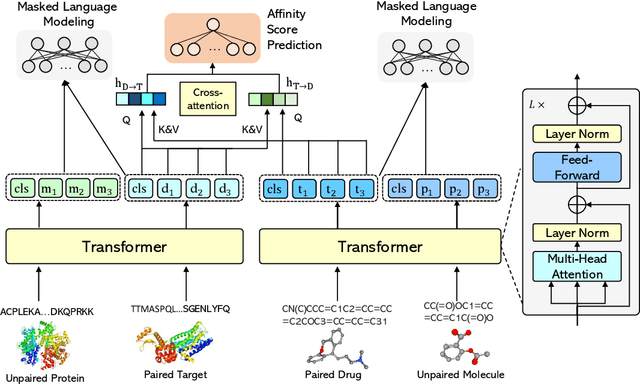
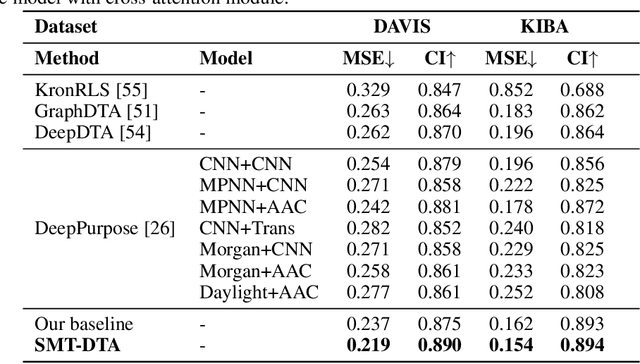


Drug-Target Affinity (DTA) prediction is an essential task for drug discovery and pharmaceutical research. Accurate predictions of DTA can greatly benefit the design of new drug. As wet experiments are costly and time consuming, the supervised data for DTA prediction is extremely limited. This seriously hinders the application of deep learning based methods, which require a large scale of supervised data. To address this challenge and improve the DTA prediction accuracy, we propose a framework with several simple yet effective strategies in this work: (1) a multi-task training strategy, which takes the DTA prediction and the masked language modeling (MLM) task on the paired drug-target dataset; (2) a semi-supervised training method to empower the drug and target representation learning by leveraging large-scale unpaired molecules and proteins in training, which differs from previous pre-training and fine-tuning methods that only utilize molecules or proteins in pre-training; and (3) a cross-attention module to enhance the interaction between drug and target representation. Extensive experiments are conducted on three real-world benchmark datasets: BindingDB, DAVIS and KIBA. The results show that our framework significantly outperforms existing methods and achieves state-of-the-art performances, e.g., $0.712$ RMSE on BindingDB IC$_{50}$ measurement with more than $5\%$ improvement than previous best work. In addition, case studies on specific drug-target binding activities, drug feature visualizations, and real-world applications demonstrate the great potential of our work. The code and data are released at https://github.com/QizhiPei/SMT-DTA
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge