Fabric Image Representation Encoding Networks for Large-scale 3D Medical Image Analysis
Paper and Code
Jun 30, 2020
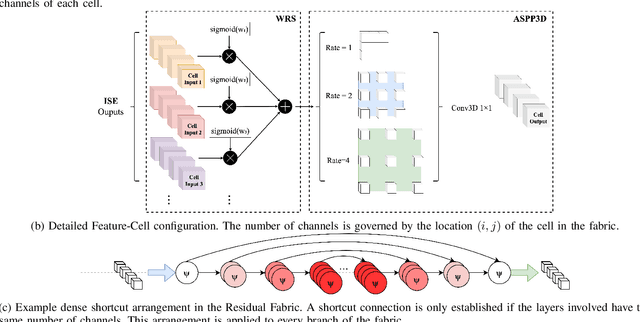
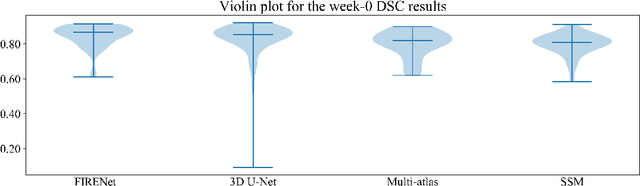
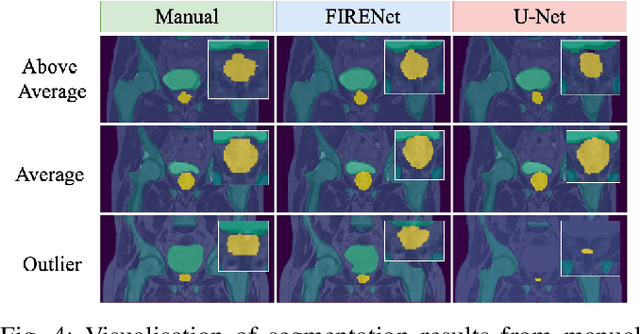
Deep neural networks are parameterised by weights that encode feature representations, whose performance is dictated through generalisation by using large-scale feature-rich datasets. The lack of large-scale labelled 3D medical imaging datasets restrict constructing such generalised networks. In this work, a novel 3D segmentation network, Fabric Image Representation Networks (FIRENet), is proposed to extract and encode generalisable feature representations from multiple medical image datasets in a large-scale manner. FIRENet learns image specific feature representations by way of 3D fabric network architecture that contains exponential number of sub-architectures to handle various protocols and coverage of anatomical regions and structures. The fabric network uses Atrous Spatial Pyramid Pooling (ASPP) extended to 3D to extract local and image-level features at a fine selection of scales. The fabric is constructed with weighted edges allowing the learnt features to dynamically adapt to the training data at an architecture level. Conditional padding modules, which are integrated into the network to reinsert voxels discarded by feature pooling, allow the network to inherently process different-size images at their original resolutions. FIRENet was trained for feature learning via automated semantic segmentation of pelvic structures and obtained a state-of-the-art median DSC score of 0.867. FIRENet was also simultaneously trained on MR (Magnatic Resonance) images acquired from 3D examinations of musculoskeletal elements in the (hip, knee, shoulder) joints and a public OAI knee dataset to perform automated segmentation of bone across anatomy. Transfer learning was used to show that the features learnt through the pelvic segmentation helped achieve improved mean DSC scores of 0.962, 0.963, 0.945 and 0.986 for automated segmentation of bone across datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge