Xiyao Fu
Contrastive Brain Network Learning via Hierarchical Signed Graph Pooling Model
Jul 14, 2022



Abstract:Recently brain networks have been widely adopted to study brain dynamics, brain development and brain diseases. Graph representation learning techniques on brain functional networks can facilitate the discovery of novel biomarkers for clinical phenotypes and neurodegenerative diseases. However, current graph learning techniques have several issues on brain network mining. Firstly, most current graph learning models are designed for unsigned graph, which hinders the analysis of many signed network data (e.g., brain functional networks). Meanwhile, the insufficiency of brain network data limits the model performance on clinical phenotypes predictions. Moreover, few of current graph learning model is interpretable, which may not be capable to provide biological insights for model outcomes. Here, we propose an interpretable hierarchical signed graph representation learning model to extract graph-level representations from brain functional networks, which can be used for different prediction tasks. In order to further improve the model performance, we also propose a new strategy to augment functional brain network data for contrastive learning. We evaluate this framework on different classification and regression tasks using the data from HCP and OASIS. Our results from extensive experiments demonstrate the superiority of the proposed model compared to several state-of-the-art techniques. Additionally, we use graph saliency maps, derived from these prediction tasks, to demonstrate detection and interpretation of phenotypic biomarkers.
Functional2Structural: Cross-Modality Brain Networks Representation Learning
May 06, 2022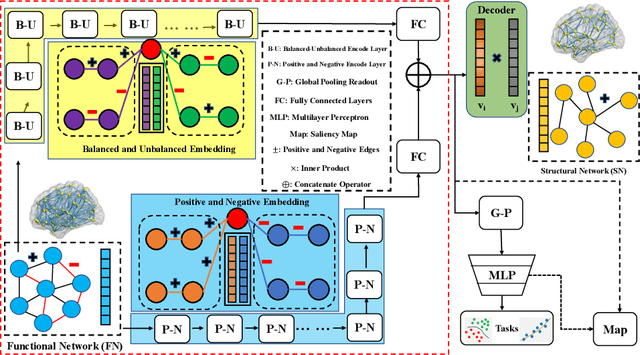
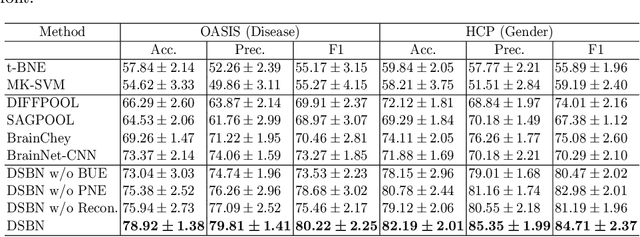
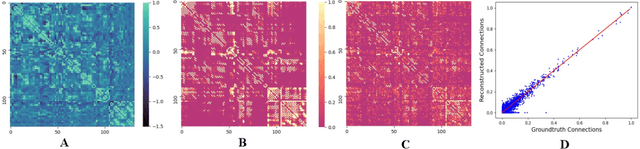
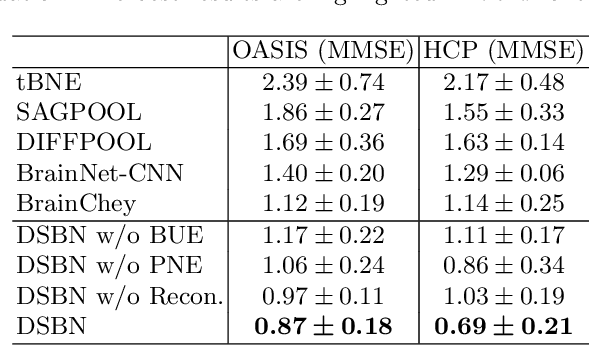
Abstract:MRI-based modeling of brain networks has been widely used to understand functional and structural interactions and connections among brain regions, and factors that affect them, such as brain development and disease. Graph mining on brain networks may facilitate the discovery of novel biomarkers for clinical phenotypes and neurodegenerative diseases. Since brain networks derived from functional and structural MRI describe the brain topology from different perspectives, exploring a representation that combines these cross-modality brain networks is non-trivial. Most current studies aim to extract a fused representation of the two types of brain network by projecting the structural network to the functional counterpart. Since the functional network is dynamic and the structural network is static, mapping a static object to a dynamic object is suboptimal. However, mapping in the opposite direction is not feasible due to the non-negativity requirement of current graph learning techniques. Here, we propose a novel graph learning framework, known as Deep Signed Brain Networks (DSBN), with a signed graph encoder that, from an opposite perspective, learns the cross-modality representations by projecting the functional network to the structural counterpart. We validate our framework on clinical phenotype and neurodegenerative disease prediction tasks using two independent, publicly available datasets (HCP and OASIS). The experimental results clearly demonstrate the advantages of our model compared to several state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge