Stamatios N. Sotiropoulos
MO-CTranS: A unified multi-organ segmentation model learning from multiple heterogeneously labelled datasets
Mar 28, 2025


Abstract:Multi-organ segmentation holds paramount significance in many clinical tasks. In practice, compared to large fully annotated datasets, multiple small datasets are often more accessible and organs are not labelled consistently. Normally, an individual model is trained for each of these datasets, which is not an effective way of using data for model learning. It remains challenging to train a single model that can robustly learn from several partially labelled datasets due to label conflict and data imbalance problems. We propose MO-CTranS: a single model that can overcome such problems. MO-CTranS contains a CNN-based encoder and a Transformer-based decoder, which are connected in a multi-resolution manner. Task-specific tokens are introduced in the decoder to help differentiate label discrepancies. Our method was evaluated and compared to several baseline models and state-of-the-art (SOTA) solutions on abdominal MRI datasets that were acquired in different views (i.e. axial and coronal) and annotated for different organs (i.e. liver, kidney, spleen). Our method achieved better performance (most were statistically significant) than the compared methods. Github link: https://github.com/naisops/MO-CTranS.
Uncertainty Quantification in Deep Learning for Safer Neuroimage Enhancement
Jul 31, 2019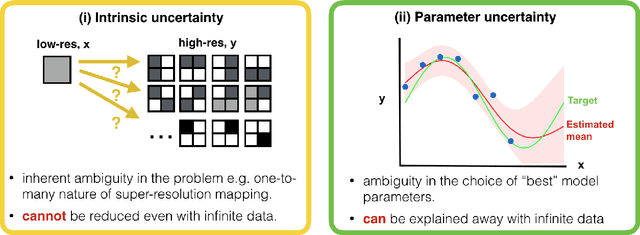
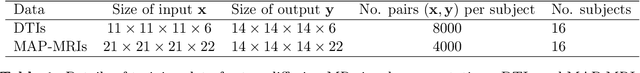
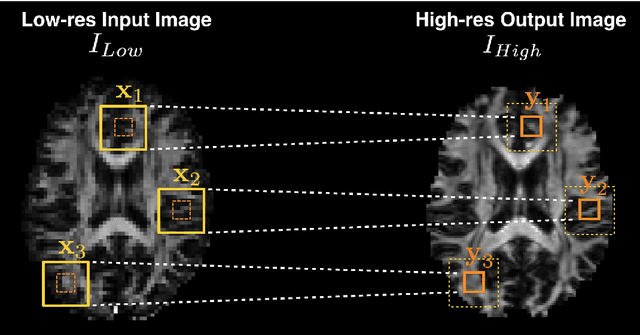
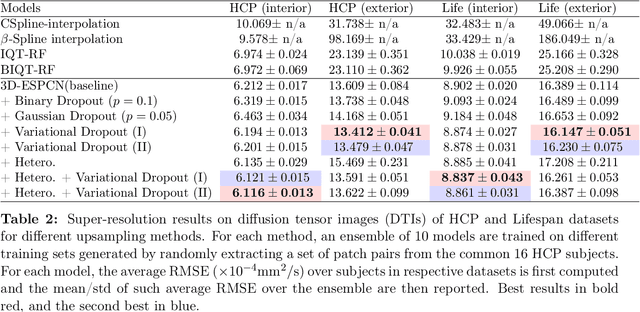
Abstract:Deep learning (DL) has shown great potential in medical image enhancement problems, such as super-resolution or image synthesis. However, to date, little consideration has been given to uncertainty quantification over the output image. Here we introduce methods to characterise different components of uncertainty in such problems and demonstrate the ideas using diffusion MRI super-resolution. Specifically, we propose to account for $intrinsic$ uncertainty through a heteroscedastic noise model and for $parameter$ uncertainty through approximate Bayesian inference, and integrate the two to quantify $predictive$ uncertainty over the output image. Moreover, we introduce a method to propagate the predictive uncertainty on a multi-channelled image to derived scalar parameters, and separately quantify the effects of intrinsic and parameter uncertainty therein. The methods are evaluated for super-resolution of two different signal representations of diffusion MR images---DTIs and Mean Apparent Propagator MRI---and their derived quantities such as MD and FA, on multiple datasets of both healthy and pathological human brains. Results highlight three key benefits of uncertainty modelling for improving the safety of DL-based image enhancement systems. Firstly, incorporating uncertainty improves the predictive performance even when test data departs from training data. Secondly, the predictive uncertainty highly correlates with errors, and is therefore capable of detecting predictive "failures". Results demonstrate that such an uncertainty measure enables subject-specific and voxel-wise risk assessment of the output images. Thirdly, we show that the method for decomposing predictive uncertainty into its independent sources provides high-level "explanations" for the performance by quantifying how much uncertainty arises from the inherent difficulty of the task or the limited training examples.
Bayesian Image Quality Transfer with CNNs: Exploring Uncertainty in dMRI Super-Resolution
May 30, 2017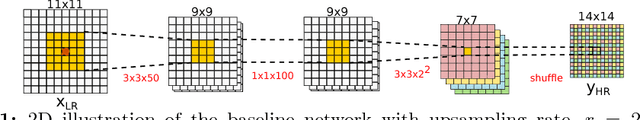
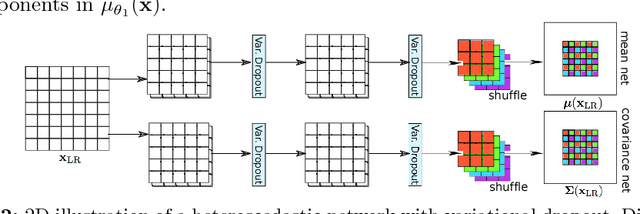
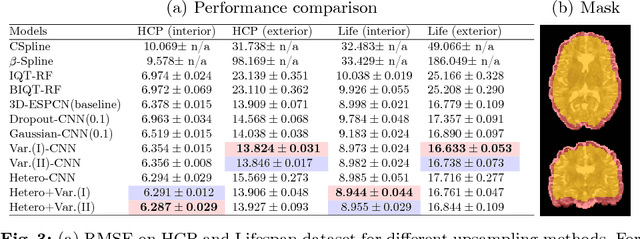
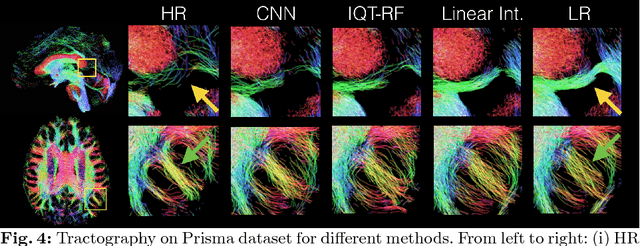
Abstract:In this work, we investigate the value of uncertainty modeling in 3D super-resolution with convolutional neural networks (CNNs). Deep learning has shown success in a plethora of medical image transformation problems, such as super-resolution (SR) and image synthesis. However, the highly ill-posed nature of such problems results in inevitable ambiguity in the learning of networks. We propose to account for intrinsic uncertainty through a per-patch heteroscedastic noise model and for parameter uncertainty through approximate Bayesian inference in the form of variational dropout. We show that the combined benefits of both lead to the state-of-the-art performance SR of diffusion MR brain images in terms of errors compared to ground truth. We further show that the reduced error scores produce tangible benefits in downstream tractography. In addition, the probabilistic nature of the methods naturally confers a mechanism to quantify uncertainty over the super-resolved output. We demonstrate through experiments on both healthy and pathological brains the potential utility of such an uncertainty measure in the risk assessment of the super-resolved images for subsequent clinical use.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge