Simon Langer
Pattern Recognition Lab, Friedrich-Alexander-Universität Erlangen-Nürnberg, Germany
Self-Supervised Radiograph Anatomical Region Classification -- How Clean Is Your Real-World Data?
Dec 20, 2024Abstract:Modern deep learning-based clinical imaging workflows rely on accurate labels of the examined anatomical region. Knowing the anatomical region is required to select applicable downstream models and to effectively generate cohorts of high quality data for future medical and machine learning research efforts. However, this information may not be available in externally sourced data or generally contain data entry errors. To address this problem, we show the effectiveness of self-supervised methods such as SimCLR and BYOL as well as supervised contrastive deep learning methods in assigning one of 14 anatomical region classes in our in-house dataset of 48,434 skeletal radiographs. We achieve a strong linear evaluation accuracy of 96.6% with a single model and 97.7% using an ensemble approach. Furthermore, only a few labeled instances (1% of the training set) suffice to achieve an accuracy of 92.2%, enabling usage in low-label and thus low-resource scenarios. Our model can be used to correct data entry mistakes: a follow-up analysis of the test set errors of our best-performing single model by an expert radiologist identified 35% incorrect labels and 11% out-of-domain images. When accounted for, the radiograph anatomical region labelling performance increased -- without and with an ensemble, respectively -- to a theoretical accuracy of 98.0% and 98.8%.
SEMPAI: a Self-Enhancing Multi-Photon Artificial Intelligence for prior-informed assessment of muscle function and pathology
Oct 28, 2022Abstract:Deep learning (DL) shows notable success in biomedical studies. However, most DL algorithms work as a black box, exclude biomedical experts, and need extensive data. We introduce the Self-Enhancing Multi-Photon Artificial Intelligence (SEMPAI), that integrates hypothesis-driven priors in a data-driven DL approach for research on multiphoton microscopy (MPM) of muscle fibers. SEMPAI utilizes meta-learning to optimize prior integration, data representation, and neural network architecture simultaneously. This allows hypothesis testing and provides interpretable feedback about the origin of biological information in MPM images. SEMPAI performs joint learning of several tasks to enable prediction for small datasets. The method is applied on an extensive multi-study dataset resulting in the largest joint analysis of pathologies and function for single muscle fibers. SEMPAI outperforms state-of-the-art biomarkers in six of seven predictive tasks, including those with scarce data. SEMPAI's DL models with integrated priors are superior to those without priors and to prior-only machine learning approaches.
DeepTechnome: Mitigating Unknown Bias in Deep Learning Based Assessment of CT Images
May 26, 2022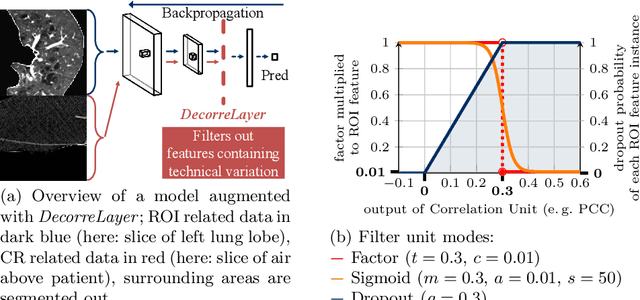
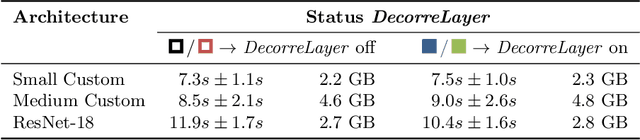
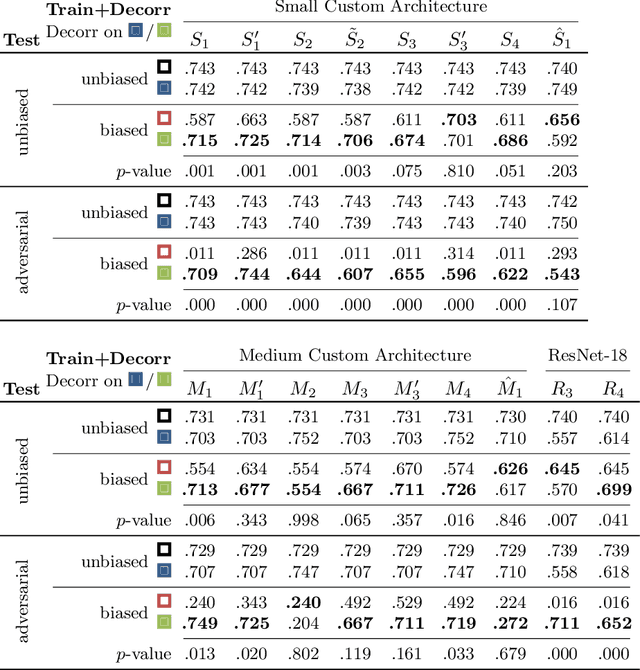


Abstract:Reliably detecting diseases using relevant biological information is crucial for real-world applicability of deep learning techniques in medical imaging. We debias deep learning models during training against unknown bias - without preprocessing/filtering the input beforehand or assuming specific knowledge about its distribution or precise nature in the dataset. We use control regions as surrogates that carry information regarding the bias, employ the classifier model to extract features, and suppress biased intermediate features with our custom, modular DecorreLayer. We evaluate our method on a dataset of 952 lung computed tomography scans by introducing simulated biases w.r.t. reconstruction kernel and noise level and propose including an adversarial test set in evaluations of bias reduction techniques. In a moderately sized model architecture, applying the proposed method to learn from data exhibiting a strong bias, it near-perfectly recovers the classification performance observed when training with corresponding unbiased data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge