Shuwei Li
Aggregating Diverse Cue Experts for AI-Generated Image Detection
Jan 13, 2026Abstract:The rapid emergence of image synthesis models poses challenges to the generalization of AI-generated image detectors. However, existing methods often rely on model-specific features, leading to overfitting and poor generalization. In this paper, we introduce the Multi-Cue Aggregation Network (MCAN), a novel framework that integrates different yet complementary cues in a unified network. MCAN employs a mixture-of-encoders adapter to dynamically process these cues, enabling more adaptive and robust feature representation. Our cues include the input image itself, which represents the overall content, and high-frequency components that emphasize edge details. Additionally, we introduce a Chromatic Inconsistency (CI) cue, which normalizes intensity values and captures noise information introduced during the image acquisition process in real images, making these noise patterns more distinguishable from those in AI-generated content. Unlike prior methods, MCAN's novelty lies in its unified multi-cue aggregation framework, which integrates spatial, frequency-domain, and chromaticity-based information for enhanced representation learning. These cues are intrinsically more indicative of real images, enhancing cross-model generalization. Extensive experiments on the GenImage, Chameleon, and UniversalFakeDetect benchmark validate the state-of-the-art performance of MCAN. In the GenImage dataset, MCAN outperforms the best state-of-the-art method by up to 7.4% in average ACC across eight different image generators.
Multimodal Modeling of CRISPR-Cas12 Activity Using Foundation Models and Chromatin Accessibility Data
Jun 12, 2025Abstract:Predicting guide RNA (gRNA) activity is critical for effective CRISPR-Cas12 genome editing but remains challenging due to limited data, variation across protospacer adjacent motifs (PAMs-short sequence requirements for Cas binding), and reliance on large-scale training. We investigate whether pre-trained biological foundation model originally trained on transcriptomic data can improve gRNA activity estimation even without domain-specific pre-training. Using embeddings from existing RNA foundation model as input to lightweight regressor, we show substantial gains over traditional baselines. We also integrate chromatin accessibility data to capture regulatory context, improving performance further. Our results highlight the effectiveness of pre-trained foundation models and chromatin accessibility data for gRNA activity prediction.
Interpretable Deep Regression Models with Interval-Censored Failure Time Data
Mar 25, 2025Abstract:Deep neural networks (DNNs) have become powerful tools for modeling complex data structures through sequentially integrating simple functions in each hidden layer. In survival analysis, recent advances of DNNs primarily focus on enhancing model capabilities, especially in exploring nonlinear covariate effects under right censoring. However, deep learning methods for interval-censored data, where the unobservable failure time is only known to lie in an interval, remain underexplored and limited to specific data type or model. This work proposes a general regression framework for interval-censored data with a broad class of partially linear transformation models, where key covariate effects are modeled parametrically while nonlinear effects of nuisance multi-modal covariates are approximated via DNNs, balancing interpretability and flexibility. We employ sieve maximum likelihood estimation by leveraging monotone splines to approximate the cumulative baseline hazard function. To ensure reliable and tractable estimation, we develop an EM algorithm incorporating stochastic gradient descent. We establish the asymptotic properties of parameter estimators and show that the DNN estimator achieves minimax-optimal convergence. Extensive simulations demonstrate superior estimation and prediction accuracy over state-of-the-art methods. Applying our method to the Alzheimer's Disease Neuroimaging Initiative dataset yields novel insights and improved predictive performance compared to traditional approaches.
InfoSEM: A Deep Generative Model with Informative Priors for Gene Regulatory Network Inference
Mar 06, 2025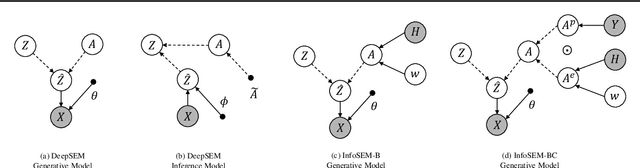
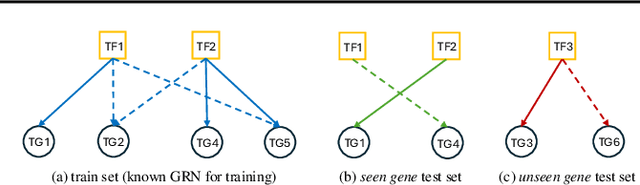
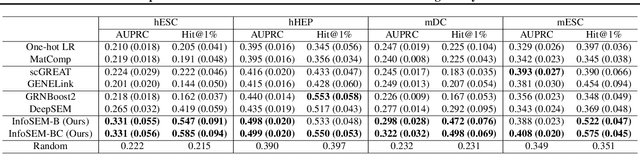
Abstract:Inferring Gene Regulatory Networks (GRNs) from gene expression data is crucial for understanding biological processes. While supervised models are reported to achieve high performance for this task, they rely on costly ground truth (GT) labels and risk learning gene-specific biases, such as class imbalances of GT interactions, rather than true regulatory mechanisms. To address these issues, we introduce InfoSEM, an unsupervised generative model that leverages textual gene embeddings as informative priors, improving GRN inference without GT labels. InfoSEM can also integrate GT labels as an additional prior when available, avoiding biases and further enhancing performance. Additionally, we propose a biologically motivated benchmarking framework that better reflects real-world applications such as biomarker discovery and reveals learned biases of existing supervised methods. InfoSEM outperforms existing models by 38.5% across four datasets using textual embeddings prior and further boosts performance by 11.1% when integrating labeled data as priors.
TransCC: Transformer-based Multiple Illuminant Color Constancy Using Multitask Learning
Nov 16, 2022



Abstract:Multi-illuminant color constancy is a challenging problem with only a few existing methods. For example, one prior work used a small set of predefined white balance settings and spatially blended among them, limiting the solution to predefined illuminations. Another method proposed a generative adversarial network and an angular loss, yet the performance is suboptimal due to the lack of regularization for multi-illumination colors. This paper introduces a transformer-based multi-task learning method to estimate single and multiple light colors from a single input image. To help our deep learning model have better cues of the light colors, achromatic-pixel detection, and edge detection are used as auxiliary tasks in our multi-task learning setting. By exploiting extracted content features from the input image as tokens, illuminant color correlations between pixels are learned by leveraging contextual information in our transformer. Our transformer approach is further assisted via a contrastive loss defined between the input, output, and ground truth. We demonstrate that our proposed model achieves 40.7% improvement compared to a state-of-the-art multi-illuminant color constancy method on a multi-illuminant dataset (LSMI). Moreover, our model maintains a robust performance on the single illuminant dataset (NUS-8) and provides 22.3% improvement on the state-of-the-art single color constancy method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge