Samuel Tesch
Hierarchical Annotation for Building A Suite of Clinical Natural Language Processing Tasks: Progress Note Understanding
Apr 06, 2022
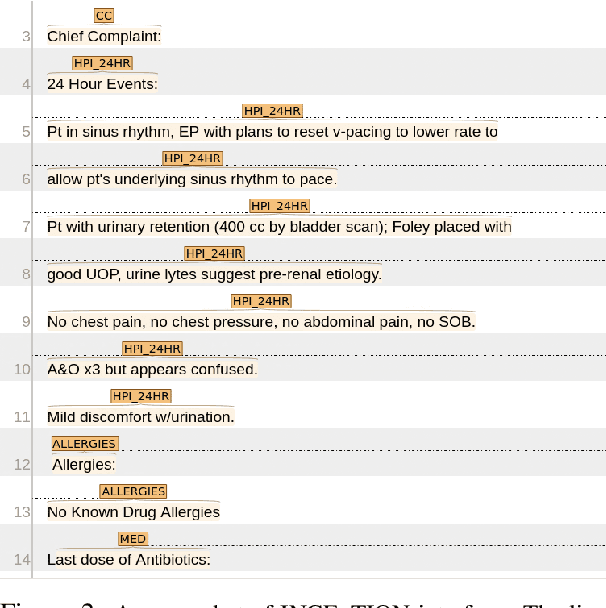
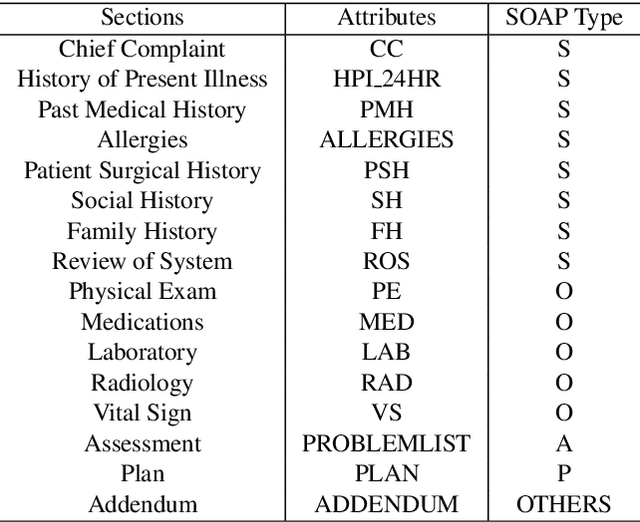

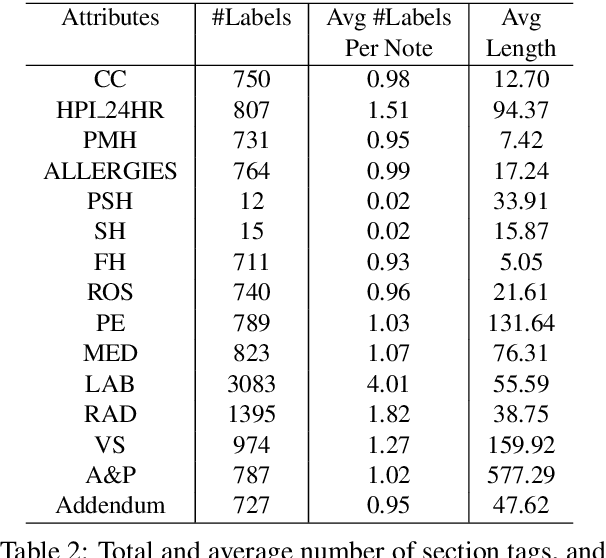
Abstract:Applying methods in natural language processing on electronic health records (EHR) data is a growing field. Existing corpus and annotation focus on modeling textual features and relation prediction. However, there is a paucity of annotated corpus built to model clinical diagnostic thinking, a process involving text understanding, domain knowledge abstraction and reasoning. This work introduces a hierarchical annotation schema with three stages to address clinical text understanding, clinical reasoning, and summarization. We created an annotated corpus based on an extensive collection of publicly available daily progress notes, a type of EHR documentation that is collected in time series in a problem-oriented format. The conventional format for a progress note follows a Subjective, Objective, Assessment and Plan heading (SOAP). We also define a new suite of tasks, Progress Note Understanding, with three tasks utilizing the three annotation stages. The novel suite of tasks was designed to train and evaluate future NLP models for clinical text understanding, clinical knowledge representation, inference, and summarization.
A Scoping Review of Publicly Available Language Tasks in Clinical Natural Language Processing
Dec 07, 2021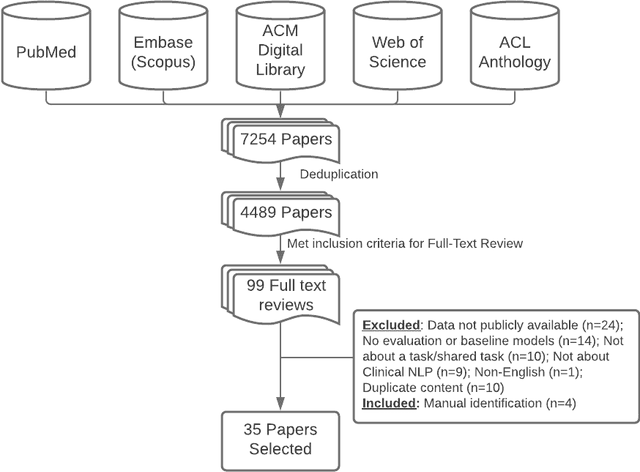
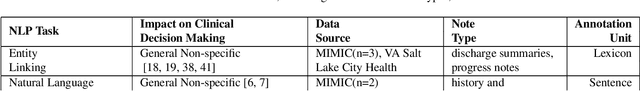
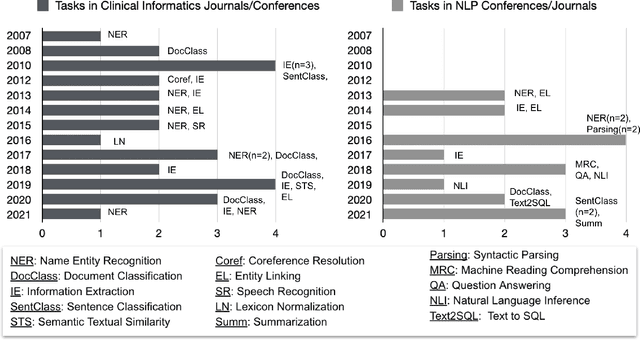

Abstract:Objective: to provide a scoping review of papers on clinical natural language processing (NLP) tasks that use publicly available electronic health record data from a cohort of patients. Materials and Methods: We searched six databases, including biomedical research and computer science literature database. A round of title/abstract screening and full-text screening were conducted by two reviewers. Our method followed the Preferred Reporting Items for Systematic Reviews and Meta-Analysis (PRISMA) guidelines. Results: A total of 35 papers with 47 clinical NLP tasks met inclusion criteria between 2007 and 2021. We categorized the tasks by the type of NLP problems, including name entity recognition, summarization, and other NLP tasks. Some tasks were introduced with a topic of clinical decision support applications, such as substance abuse, phenotyping, cohort selection for clinical trial. We summarized the tasks by publication and dataset information. Discussion: The breadth of clinical NLP tasks keeps growing as the field of NLP evolves with advancements in language systems. However, gaps exist in divergent interests between general domain NLP community and clinical informatics community, and in generalizability of the data sources. We also identified issues in data selection and preparation including the lack of time-sensitive data, and invalidity of problem size and evaluation. Conclusions: The existing clinical NLP tasks cover a wide range of topics and the field will continue to grow and attract more attention from both general domain NLP and clinical informatics community. We encourage future work to incorporate multi-disciplinary collaboration, reporting transparency, and standardization in data preparation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge