Richard Barcus
Nonhuman Primate Brain Tissue Segmentation Using a Transfer Learning Approach
Apr 01, 2025Abstract:Non-human primates (NHPs) serve as critical models for understanding human brain function and neurological disorders due to their close evolutionary relationship with humans. Accurate brain tissue segmentation in NHPs is critical for understanding neurological disorders, but challenging due to the scarcity of annotated NHP brain MRI datasets, the small size of the NHP brain, the limited resolution of available imaging data and the anatomical differences between human and NHP brains. To address these challenges, we propose a novel approach utilizing STU-Net with transfer learning to leverage knowledge transferred from human brain MRI data to enhance segmentation accuracy in the NHP brain MRI, particularly when training data is limited. The combination of STU-Net and transfer learning effectively delineates complex tissue boundaries and captures fine anatomical details specific to NHP brains. Notably, our method demonstrated improvement in segmenting small subcortical structures such as putamen and thalamus that are challenging to resolve with limited spatial resolution and tissue contrast, and achieved DSC of over 0.88, IoU over 0.8 and HD95 under 7. This study introduces a robust method for multi-class brain tissue segmentation in NHPs, potentially accelerating research in evolutionary neuroscience and preclinical studies of neurological disorders relevant to human health.
A transformer-based deep learning approach for classifying brain metastases into primary organ sites using clinical whole brain MRI images
Oct 07, 2021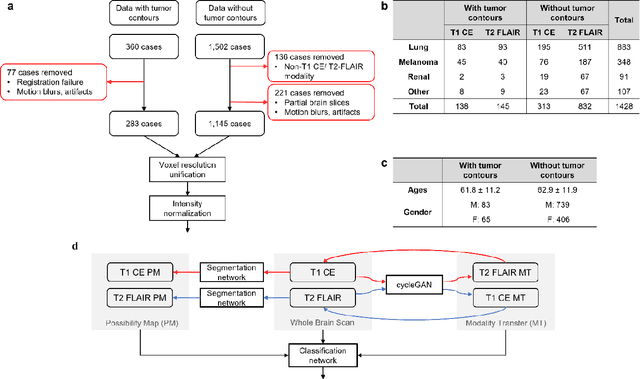
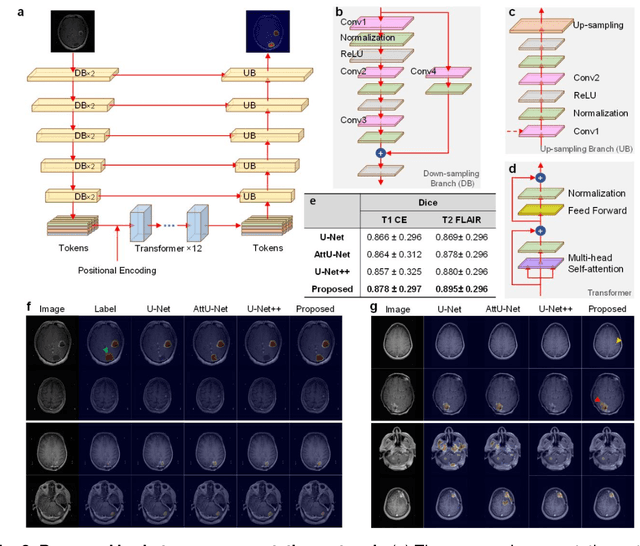
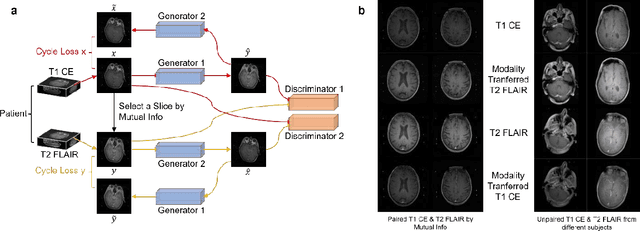
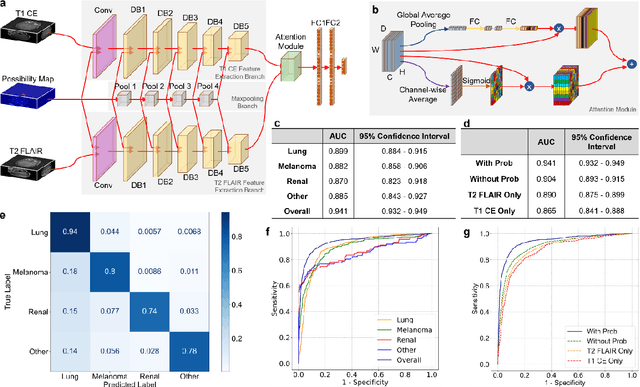
Abstract:The treatment decisions for brain metastatic disease are driven by knowledge of the primary organ site cancer histology, often requiring invasive biopsy. This study aims to develop a novel deep learning approach for accurate and rapid non-invasive identification of brain metastatic tumor histology with conventional whole-brain MRI. The use of clinical whole-brain data and the end-to-end pipeline obviate external human intervention. This IRB-approved single-site retrospective study was comprised of patients (n=1,293) referred for MRI treatment-planning and gamma knife radiosurgery from July 2000 to May 2019. Contrast-enhanced T1-weighted contrast enhanced and T2-weighted-Fluid-Attenuated Inversion Recovery brain MRI exams (n=1,428) were minimally preprocessed (voxel resolution unification and signal-intensity rescaling/normalization), requiring only seconds per an MRI scan, and input into the proposed deep learning workflow for tumor segmentation, modality transfer, and primary site classification associated with brain metastatic disease in one of four classes (lung, melanoma, renal, and other). Ten-fold cross-validation generated the overall AUC of 0.941, lung class AUC of 0.899, melanoma class AUC of 0.882, renal class AUC of 0.870, and other class AUC of 0.885. It is convincingly established that whole-brain imaging features would be sufficiently discriminative to allow accurate diagnosis of the primary organ site of malignancy. Our end-to-end deep learning-based radiomic method has a great translational potential for classifying metastatic tumor types using whole-brain MRI images, without additional human intervention. Further refinement may offer invaluable tools to expedite primary organ site cancer identification for treatment of brain metastatic disease and improvement of patient outcomes and survival.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge