Peter Gemmar
Shape-aware Surface Reconstruction from Sparse 3D Point-Clouds
Feb 15, 2017
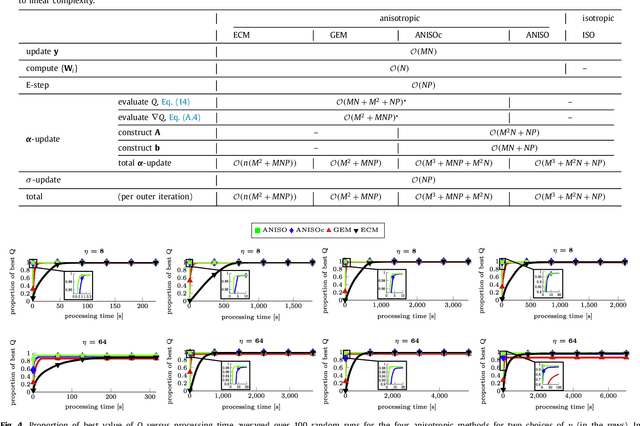
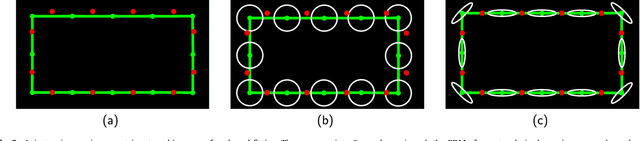
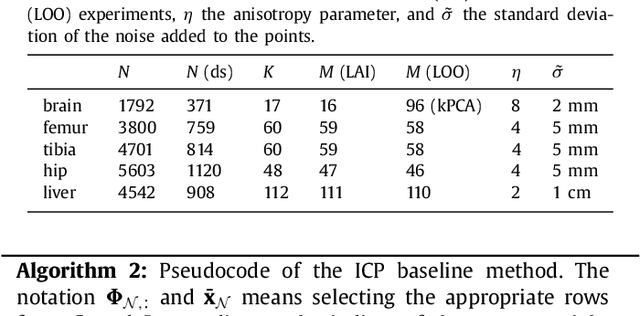
Abstract:The reconstruction of an object's shape or surface from a set of 3D points plays an important role in medical image analysis, e.g. in anatomy reconstruction from tomographic measurements or in the process of aligning intra-operative navigation and preoperative planning data. In such scenarios, one usually has to deal with sparse data, which significantly aggravates the problem of reconstruction. However, medical applications often provide contextual information about the 3D point data that allow to incorporate prior knowledge about the shape that is to be reconstructed. To this end, we propose the use of a statistical shape model (SSM) as a prior for surface reconstruction. The SSM is represented by a point distribution model (PDM), which is associated with a surface mesh. Using the shape distribution that is modelled by the PDM, we formulate the problem of surface reconstruction from a probabilistic perspective based on a Gaussian Mixture Model (GMM). In order to do so, the given points are interpreted as samples of the GMM. By using mixture components with anisotropic covariances that are "oriented" according to the surface normals at the PDM points, a surface-based fitting is accomplished. Estimating the parameters of the GMM in a maximum a posteriori manner yields the reconstruction of the surface from the given data points. We compare our method to the extensively used Iterative Closest Points method on several different anatomical datasets/SSMs (brain, femur, tibia, hip, liver) and demonstrate superior accuracy and robustness on sparse data.
Linear Shape Deformation Models with Local Support Using Graph-based Structured Matrix Factorisation
May 11, 2016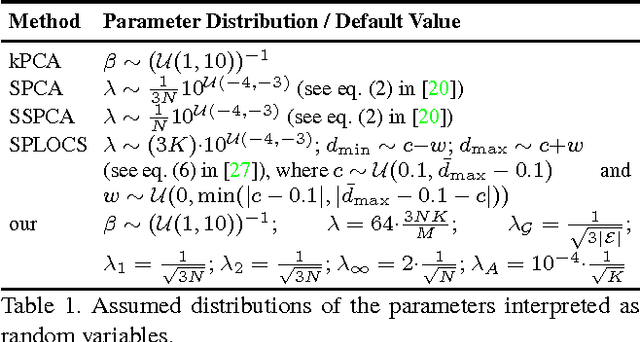
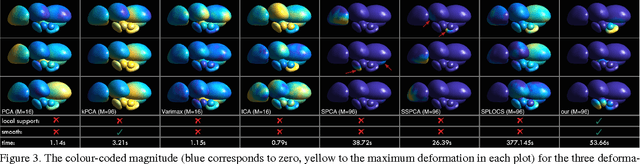
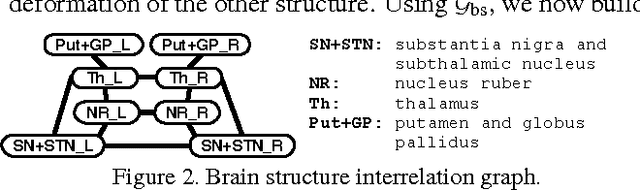

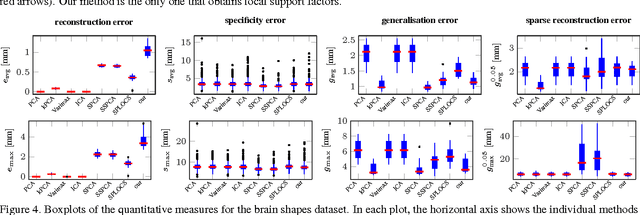
Abstract:Representing 3D shape deformations by linear models in high-dimensional space has many applications in computer vision and medical imaging, such as shape-based interpolation or segmentation. Commonly, using Principal Components Analysis a low-dimensional (affine) subspace of the high-dimensional shape space is determined. However, the resulting factors (the most dominant eigenvectors of the covariance matrix) have global support, i.e. changing the coefficient of a single factor deforms the entire shape. In this paper, a method to obtain deformation factors with local support is presented. The benefits of such models include better flexibility and interpretability as well as the possibility of interactively deforming shapes locally. For that, based on a well-grounded theoretical motivation, we formulate a matrix factorisation problem employing sparsity and graph-based regularisation terms. We demonstrate that for brain shapes our method outperforms the state of the art in local support models with respect to generalisation ability and sparse shape reconstruction, whereas for human body shapes our method gives more realistic deformations.
A Solution for Multi-Alignment by Transformation Synchronisation
Apr 14, 2015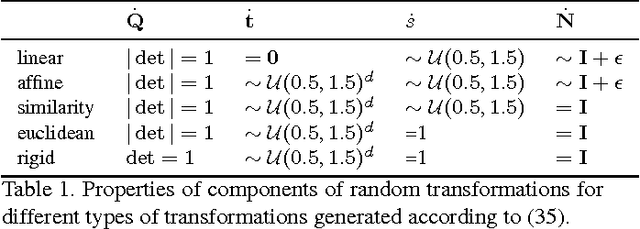
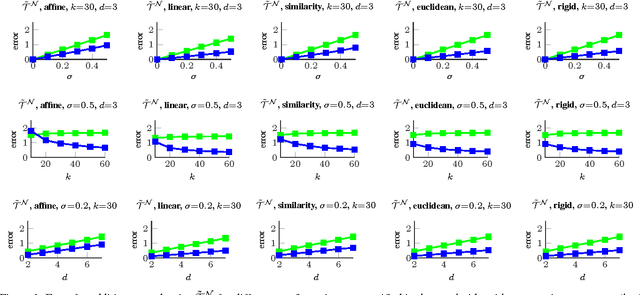


Abstract:The alignment of a set of objects by means of transformations plays an important role in computer vision. Whilst the case for only two objects can be solved globally, when multiple objects are considered usually iterative methods are used. In practice the iterative methods perform well if the relative transformations between any pair of objects are free of noise. However, if only noisy relative transformations are available (e.g. due to missing data or wrong correspondences) the iterative methods may fail. Based on the observation that the underlying noise-free transformations can be retrieved from the null space of a matrix that can directly be obtained from pairwise alignments, this paper presents a novel method for the synchronisation of pairwise transformations such that they are transitively consistent. Simulations demonstrate that for noisy transformations, a large proportion of missing data and even for wrong correspondence assignments the method delivers encouraging results.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge