Paul Xing
Polytechnique Montréal
Towards Transcranial 3D Ultrasound Localization Microscopy of the Nonhuman Primate Brain
Apr 04, 2024



Abstract:Hemodynamic changes occur in stroke and neurodegenerative diseases. Developing imaging techniques allowing the in vivo visualization and quantification of cerebral blood flow would help better understand the underlying mechanism of those cerebrovascular diseases. 3D ultrasound localization microscopy (ULM) is a novel technology that can map the microvasculature of the brain at large depth and has been mainly used until now in rodents. Here, we demonstrated the feasibility of 3D ULM of the nonhuman primate (NHP) brain with a single 256-channels programmable ultrasound scanner. We achieved a highly resolved vascular map of the macaque brain at large depth in presence of craniotomy and durectomy using an 8-MHz multiplexed matrix probe. We were able to distinguish vessels as small as 26.9 {\mu}m. We also demonstrated that transcranial imaging of the macaque brain at similar depth was feasible using a 3-MHz probe and achieved a resolution of 60.4 {\mu}m. This work paves the way to clinical application of 3D ULM.
Pruning Sparse Tensor Neural Networks Enables Deep Learning for 3D Ultrasound Localization Microscopy
Feb 14, 2024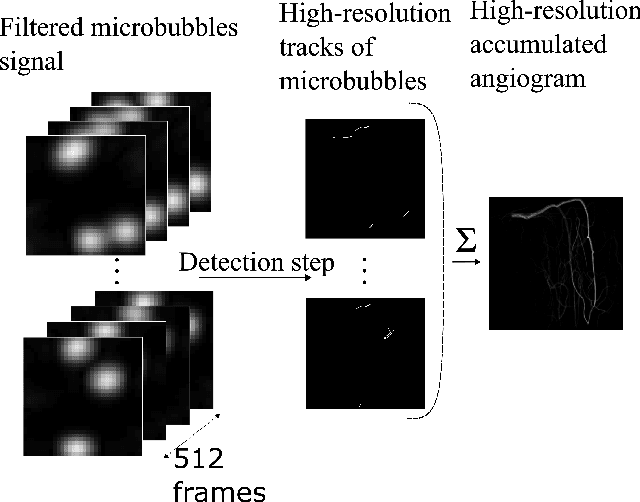
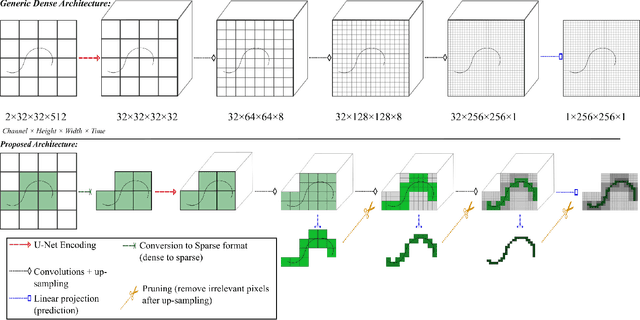
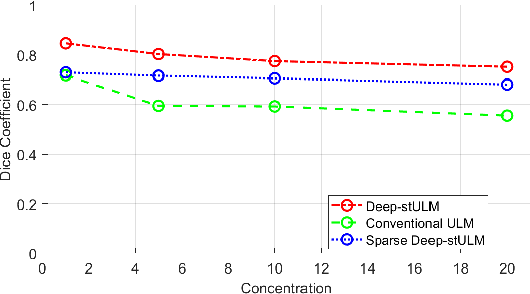
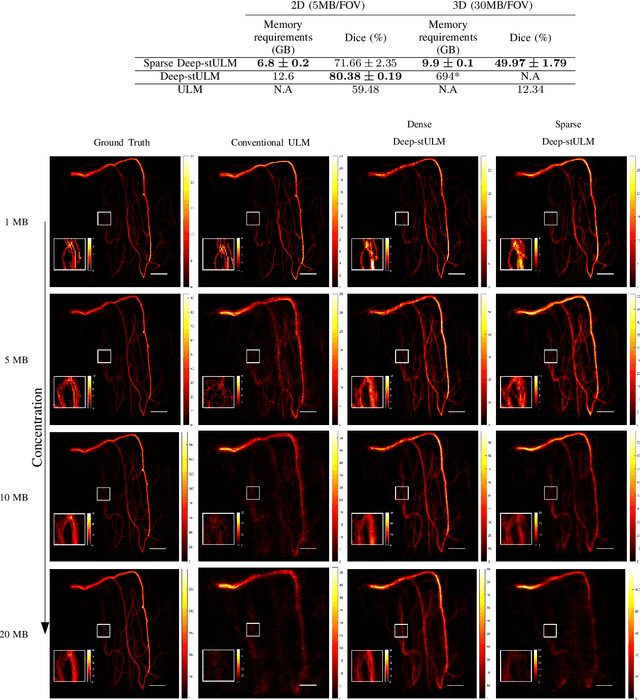
Abstract:Ultrasound Localization Microscopy (ULM) is a non-invasive technique that allows for the imaging of micro-vessels in vivo, at depth and with a resolution on the order of ten microns. ULM is based on the sub-resolution localization of individual microbubbles injected in the bloodstream. Mapping the whole angioarchitecture requires the accumulation of microbubbles trajectories from thousands of frames, typically acquired over a few minutes. ULM acquisition times can be reduced by increasing the microbubble concentration, but requires more advanced algorithms to detect them individually. Several deep learning approaches have been proposed for this task, but they remain limited to 2D imaging, in part due to the associated large memory requirements. Herein, we propose to use sparse tensor neural networks to reduce memory usage in 2D and to improve the scaling of the memory requirement for the extension of deep learning architecture to 3D. We study several approaches to efficiently convert ultrasound data into a sparse format and study the impact of the associated loss of information. When applied in 2D, the sparse formulation reduces the memory requirements by a factor 2 at the cost of a small reduction of performance when compared against dense networks. In 3D, the proposed approach reduces memory requirements by two order of magnitude while largely outperforming conventional ULM in high concentration settings. We show that Sparse Tensor Neural Networks in 3D ULM allow for the same benefits as dense deep learning based method in 2D ULM i.e. the use of higher concentration in silico and reduced acquisition time.
Inverse Problem Based on a Sparse Representation of Contrast-enhanced Ultrasound Data for in vivo Transcranial Imaging
Jan 18, 2024Abstract:Transcranial ultrasound imaging is currently limited by attenuation and aberration induced by the skull. First used in contrast-enhanced ultrasound (CEUS), highly echoic microbubbles allowed for the development of novel imaging modalities such as ultrasound localization microscopy (ULM). Herein, we develop an inverse problem approach to aberration correction (IPAC) that leverages the sparsity of microbubble signals. We propose to use the \textit{a priori} knowledge of the medium based upon microbubble localization and wave propagation to build a forward model to link the measured signals directly to the aberration function. A standard least-squares inversion is then used to retrieve the aberration function. We first validated IPAC on simulated data of a vascular network using plane wave as well as divergent wave emissions. We then evaluated the reproducibility of IPAC \textit{in vivo} in 5 mouse brains. We showed that aberration correction improved the contrast of CEUS images by 4.6 dB. For ULM images, IPAC yielded sharper vessels, reduced vessel duplications, and improved the resolution from 21.1 $\mu$m to 18.3 $\mu$m. Aberration correction also improved hemodynamic quantification for velocity magnitude and flow direction.
A Tracking prior to Localization workflow for Ultrasound Localization Microscopy
Aug 04, 2023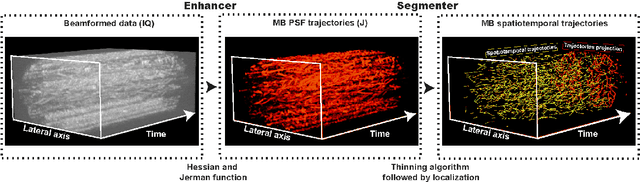
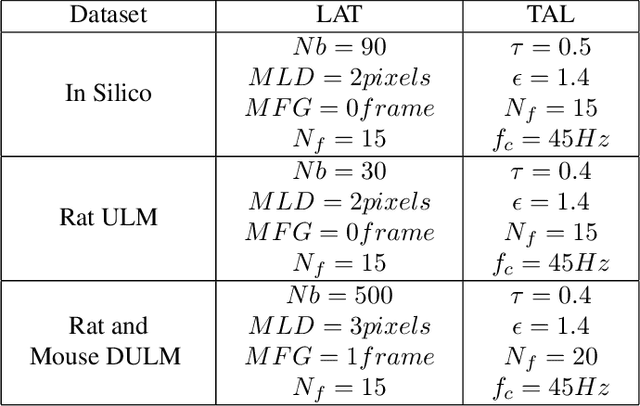
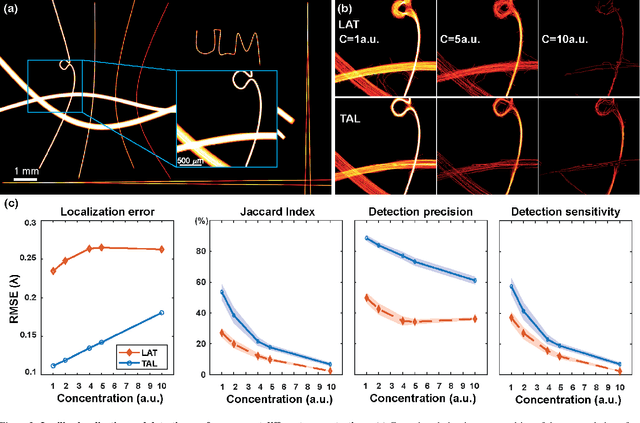
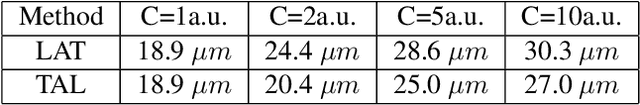
Abstract:Ultrasound Localization Microscopy (ULM) has proven effective in resolving microvascular structures and local mean velocities at sub-diffraction-limited scales, offering high-resolution imaging capabilities. Dynamic ULM (DULM) enables the creation of angiography or velocity movies throughout cardiac cycles. Currently, these techniques rely on a Localization-and-Tracking (LAT) workflow consisting in detecting microbubbles (MB) in the frames before pairing them to generate tracks. While conventional LAT methods perform well at low concentrations, they suffer from longer acquisition times and degraded localization and tracking accuracy at higher concentrations, leading to biased angiogram reconstruction and velocity estimation. In this study, we propose a novel approach to address these challenges by reversing the current workflow. The proposed method, Tracking-and-Localization (TAL), relies on first tracking the MB and then performing localization. Through comprehensive benchmarking using both in silico and in vivo experiments and employing various metrics to quantify ULM angiography and velocity maps, we demonstrate that the TAL method consistently outperforms the reference LAT workflow. Moreover, when applied to DULM, TAL successfully extracts velocity variations along the cardiac cycle with improved repeatability. The findings of this work highlight the effectiveness of the TAL approach in overcoming the limitations of conventional LAT methods, providing enhanced ULM angiography and velocity imaging.
Phase Aberration Correction for in vivo Ultrasound Localization Microscopy Using a Spatiotemporal Complex-Valued Neural Network
Sep 23, 2022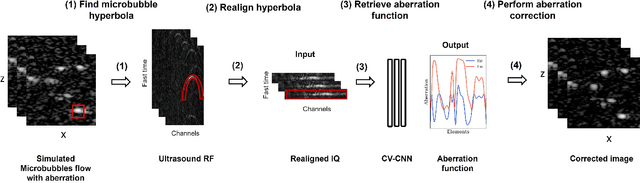
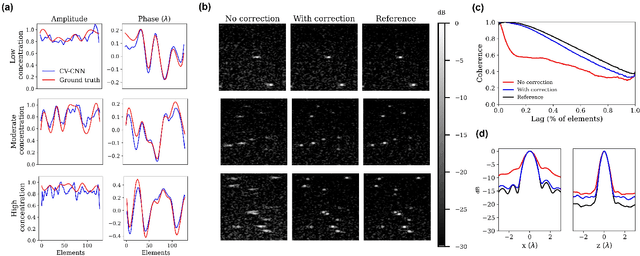
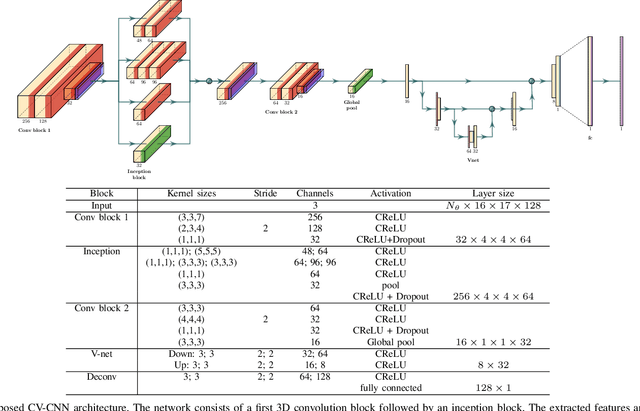
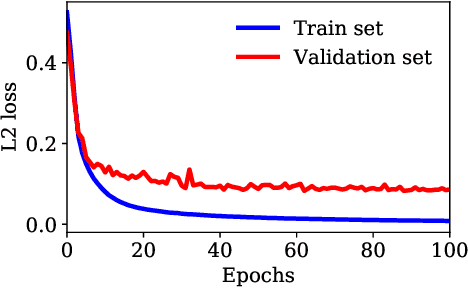

Abstract:Ultrasound Localization Microscopy (ULM) can map microvessels at a resolution of a few micrometers ({\mu}m). Transcranial ULM remains challenging in presence of aberrations caused by the skull, which lead to localization errors. Herein, we propose a deep learning approach based on recently introduced complex-valued convolutional neural networks (CV-CNNs) to retrieve the aberration function, which can then be used to form enhanced images using standard delay-and-sum beamforming. Complex-valued convolutional networks were selected as they can apply time delays through multiplication with in-phase quadrature input data. Predicting the aberration function rather than corrected images also confers enhanced explainability to the network. In addition, 3D spatiotemporal convolutions were used for the network to leverage entire microbubble tracks. For training and validation, we used an anatomically and hemodynamically realistic mouse brain microvascular network model to simulate the flow of microbubbles in presence of aberration. We then confirmed the capability of our network to generalize to transcranial in vivo data in the mouse brain (n=2). Qualitatively, vascular reconstructions using a pixel-wise predicted aberration function included additional and sharper vessels. The spatial resolution was evaluated by using the Fourier ring correlation (FRC). After correction, we measured a resolution of 16.7 {\mu}m in vivo, representing an improvement of up to 27.5 %. This work leads to different applications for complex-valued convolutions in biomedical imaging and strategies to perform transcranial ULM.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge