Chloé Bourquin
A Deep Learning Framework for Spatiotemporal Ultrasound Localization Microscopy
Oct 12, 2023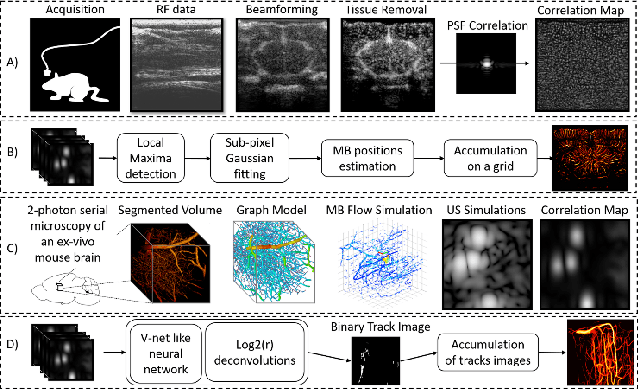
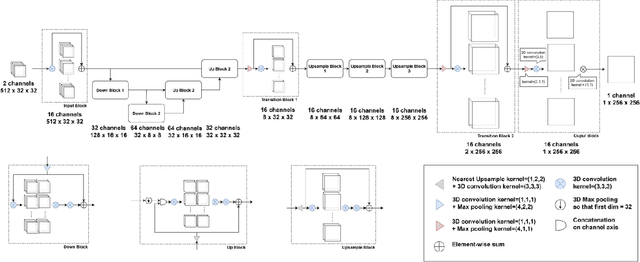
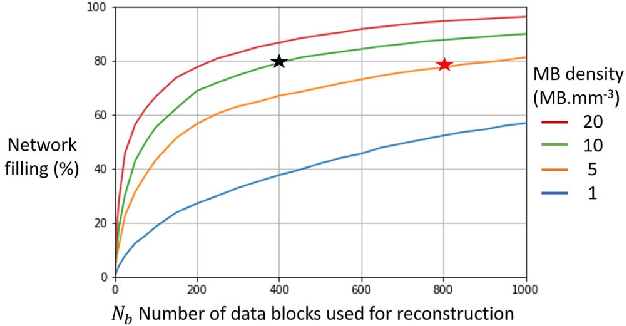
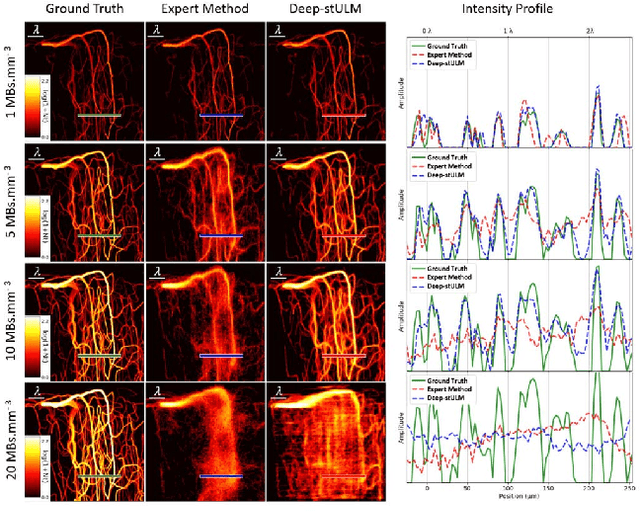
Abstract:Ultrasound Localization Microscopy can resolve the microvascular bed down to a few micrometers. To achieve such performance microbubble contrast agents must perfuse the entire microvascular network. Microbubbles are then located individually and tracked over time to sample individual vessels, typically over hundreds of thousands of images. To overcome the fundamental limit of diffraction and achieve a dense reconstruction of the network, low microbubble concentrations must be used, which lead to acquisitions lasting several minutes. Conventional processing pipelines are currently unable to deal with interference from multiple nearby microbubbles, further reducing achievable concentrations. This work overcomes this problem by proposing a Deep Learning approach to recover dense vascular networks from ultrasound acquisitions with high microbubble concentrations. A realistic mouse brain microvascular network, segmented from 2-photon microscopy, was used to train a three-dimensional convolutional neural network based on a V-net architecture. Ultrasound data sets from multiple microbubbles flowing through the microvascular network were simulated and used as ground truth to train the 3D CNN to track microbubbles. The 3D-CNN approach was validated in silico using a subset of the data and in vivo on a rat brain acquisition. In silico, the CNN reconstructed vascular networks with higher precision (81%) than a conventional ULM framework (70%). In vivo, the CNN could resolve micro vessels as small as 10 $\mu$m with an increase in resolution when compared against a conventional approach.
* Copyright 2021 IEEE. Personal use of this material is permitted. Permission from IEEE must be obtained for all other uses, in any current or future media, including reprinting/republishing this material for advertising or promotional purposes, creating new collective works, for resale or redistribution to servers or lists, or reuse of any copyrighted component of this work in other works
A Tracking prior to Localization workflow for Ultrasound Localization Microscopy
Aug 04, 2023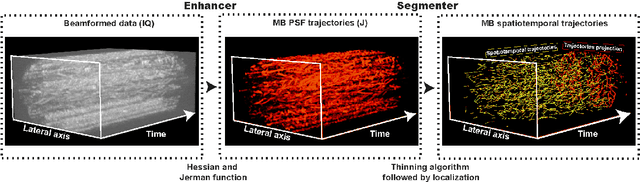
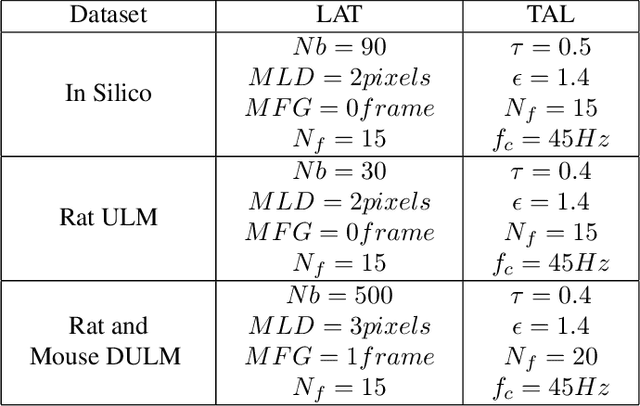
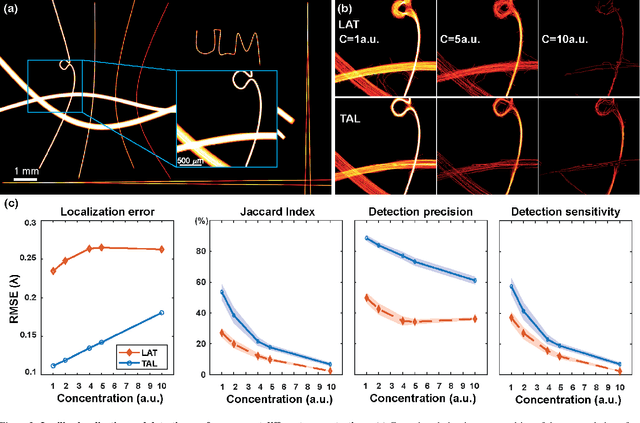
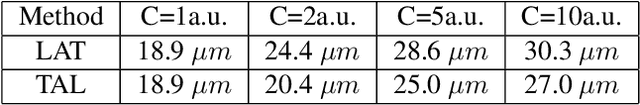
Abstract:Ultrasound Localization Microscopy (ULM) has proven effective in resolving microvascular structures and local mean velocities at sub-diffraction-limited scales, offering high-resolution imaging capabilities. Dynamic ULM (DULM) enables the creation of angiography or velocity movies throughout cardiac cycles. Currently, these techniques rely on a Localization-and-Tracking (LAT) workflow consisting in detecting microbubbles (MB) in the frames before pairing them to generate tracks. While conventional LAT methods perform well at low concentrations, they suffer from longer acquisition times and degraded localization and tracking accuracy at higher concentrations, leading to biased angiogram reconstruction and velocity estimation. In this study, we propose a novel approach to address these challenges by reversing the current workflow. The proposed method, Tracking-and-Localization (TAL), relies on first tracking the MB and then performing localization. Through comprehensive benchmarking using both in silico and in vivo experiments and employing various metrics to quantify ULM angiography and velocity maps, we demonstrate that the TAL method consistently outperforms the reference LAT workflow. Moreover, when applied to DULM, TAL successfully extracts velocity variations along the cardiac cycle with improved repeatability. The findings of this work highlight the effectiveness of the TAL approach in overcoming the limitations of conventional LAT methods, providing enhanced ULM angiography and velocity imaging.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge